"Image": models, code, and papers
Fast light-field 3D microscopy with out-of-distribution detection and adaptation through Conditional Normalizing Flows
Jun 14, 2023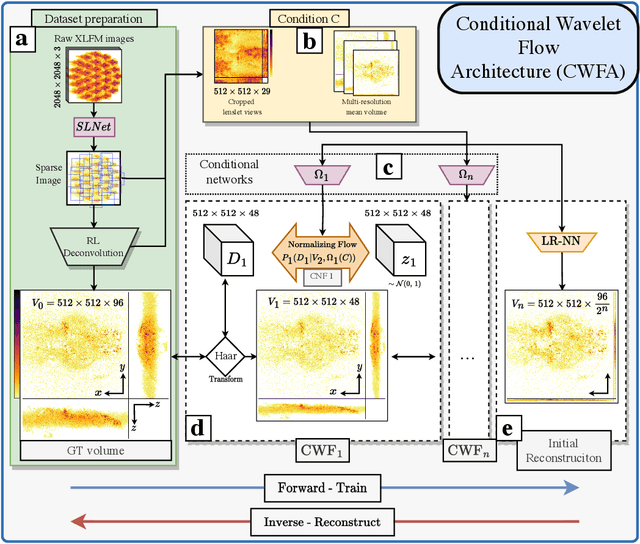
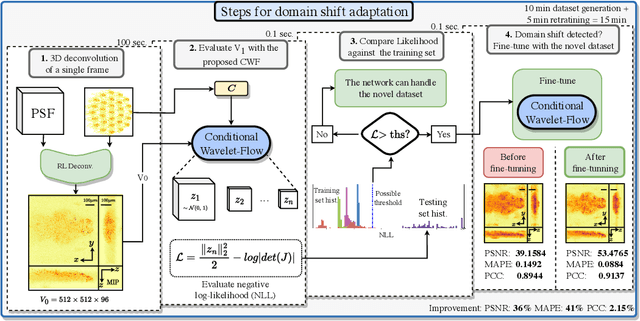
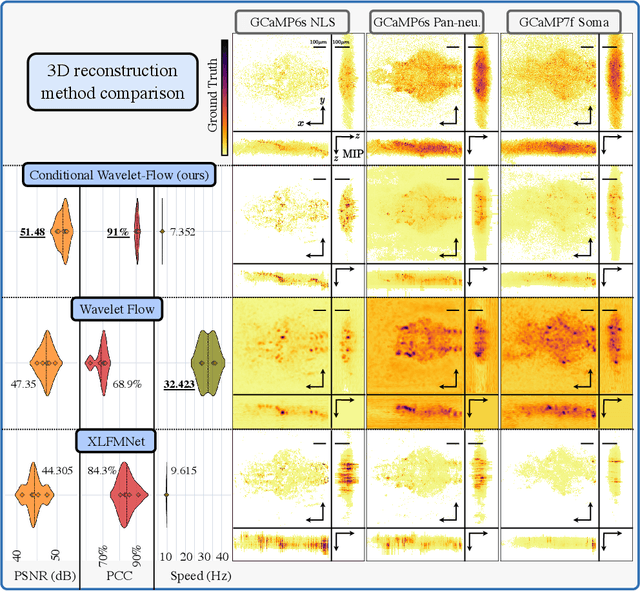
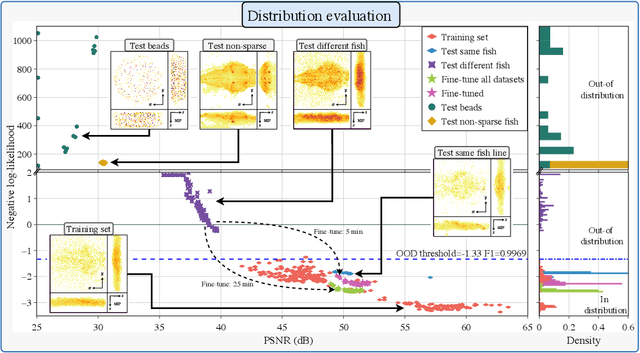
Real-time 3D fluorescence microscopy is crucial for the spatiotemporal analysis of live organisms, such as neural activity monitoring. The eXtended field-of-view light field microscope (XLFM), also known as Fourier light field microscope, is a straightforward, single snapshot solution to achieve this. The XLFM acquires spatial-angular information in a single camera exposure. In a subsequent step, a 3D volume can be algorithmically reconstructed, making it exceptionally well-suited for real-time 3D acquisition and potential analysis. Unfortunately, traditional reconstruction methods (like deconvolution) require lengthy processing times (0.0220 Hz), hampering the speed advantages of the XLFM. Neural network architectures can overcome the speed constraints at the expense of lacking certainty metrics, which renders them untrustworthy for the biomedical realm. This work proposes a novel architecture to perform fast 3D reconstructions of live immobilized zebrafish neural activity based on a conditional normalizing flow. It reconstructs volumes at 8 Hz spanning 512x512x96 voxels, and it can be trained in under two hours due to the small dataset requirements (10 image-volume pairs). Furthermore, normalizing flows allow for exact Likelihood computation, enabling distribution monitoring, followed by out-of-distribution detection and retraining of the system when a novel sample is detected. We evaluate the proposed method on a cross-validation approach involving multiple in-distribution samples (genetically identical zebrafish) and various out-of-distribution ones.
Semi-supervised Cell Recognition under Point Supervision
Jun 14, 2023Cell recognition is a fundamental task in digital histopathology image analysis. Point-based cell recognition (PCR) methods normally require a vast number of annotations, which is extremely costly, time-consuming and labor-intensive. Semi-supervised learning (SSL) can provide a shortcut to make full use of cell information in gigapixel whole slide images without exhaustive labeling. However, research into semi-supervised point-based cell recognition (SSPCR) remains largely overlooked. Previous SSPCR works are all built on density map-based PCR models, which suffer from unsatisfactory accuracy, slow inference speed and high sensitivity to hyper-parameters. To address these issues, end-to-end PCR models are proposed recently. In this paper, we develop a SSPCR framework suitable for the end-to-end PCR models for the first time. Overall, we use the current models to generate pseudo labels for unlabeled images, which are in turn utilized to supervise the models training. Besides, we introduce a co-teaching strategy to overcome the confirmation bias problem that generally exists in self-training. A distribution alignment technique is also incorporated to produce high-quality, unbiased pseudo labels for unlabeled data. Experimental results on four histopathology datasets concerning different types of staining styles show the effectiveness and versatility of the proposed framework. Code is available at \textcolor{magenta}{\url{https://github.com/windygooo/SSPCR}
Noise Stability Optimization for Flat Minima with Optimal Convergence Rates
Jun 14, 2023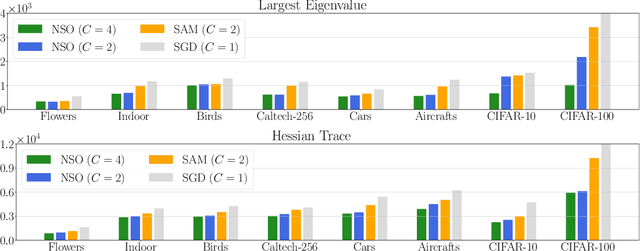
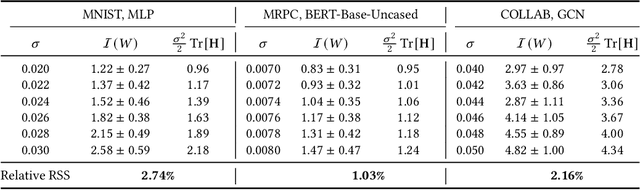
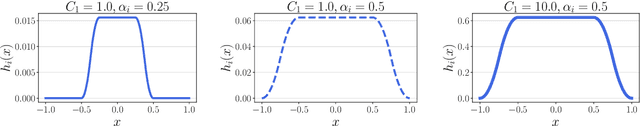
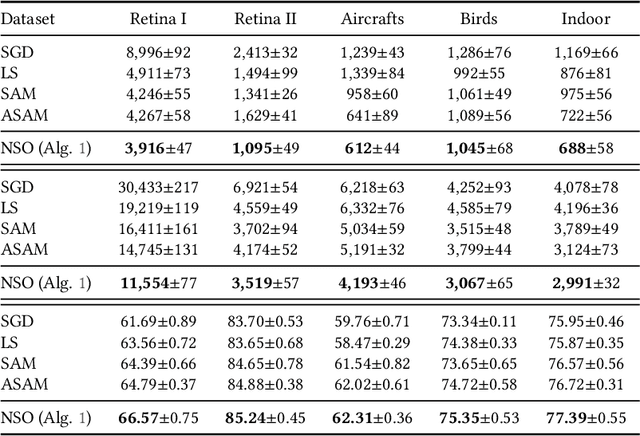
We consider finding flat, local minimizers by adding average weight perturbations. Given a nonconvex function $f: \mathbb{R}^d \rightarrow \mathbb{R}$ and a $d$-dimensional distribution $\mathcal{P}$ which is symmetric at zero, we perturb the weight of $f$ and define $F(W) = \mathbb{E}[f({W + U})]$, where $U$ is a random sample from $\mathcal{P}$. This injection induces regularization through the Hessian trace of $f$ for small, isotropic Gaussian perturbations. Thus, the weight-perturbed function biases to minimizers with low Hessian trace. Several prior works have studied settings related to this weight-perturbed function by designing algorithms to improve generalization. Still, convergence rates are not known for finding minima under the average perturbations of the function $F$. This paper considers an SGD-like algorithm that injects random noise before computing gradients while leveraging the symmetry of $\mathcal{P}$ to reduce variance. We then provide a rigorous analysis, showing matching upper and lower bounds of our algorithm for finding an approximate first-order stationary point of $F$ when the gradient of $f$ is Lipschitz-continuous. We empirically validate our algorithm for several image classification tasks with various architectures. Compared to sharpness-aware minimization, we note a 12.6% and 7.8% drop in the Hessian trace and top eigenvalue of the found minima, respectively, averaged over eight datasets. Ablation studies validate the benefit of the design of our algorithm.
UOD: Universal One-shot Detection of Anatomical Landmarks
Jun 14, 2023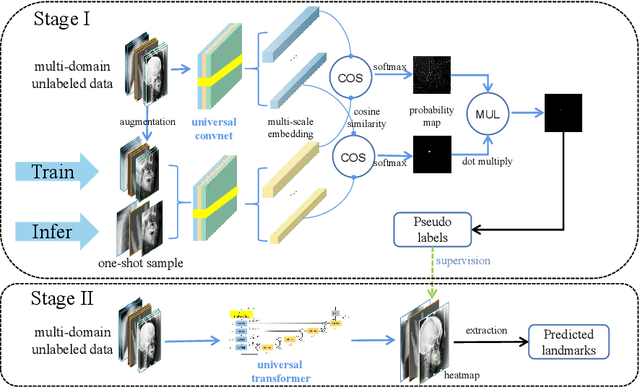
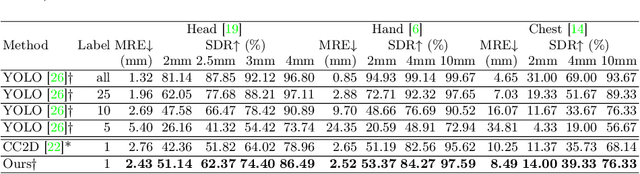
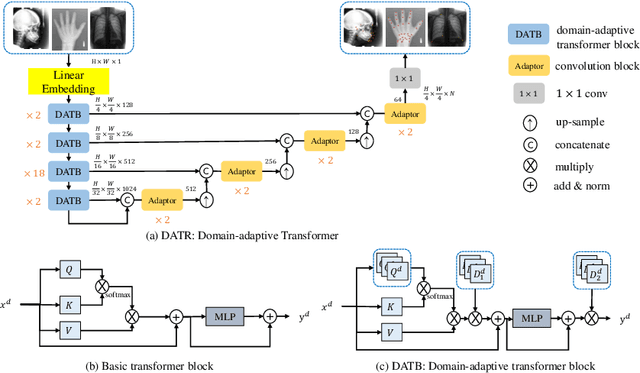
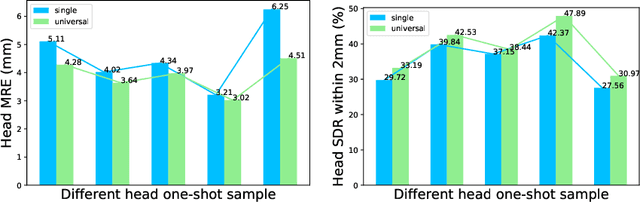
One-shot medical landmark detection gains much attention and achieves great success for its label-efficient training process. However, existing one-shot learning methods are highly specialized in a single domain and suffer domain preference heavily in the situation of multi-domain unlabeled data. Moreover, one-shot learning is not robust that it faces performance drop when annotating a sub-optimal image. To tackle these issues, we resort to developing a domain-adaptive one-shot landmark detection framework for handling multi-domain medical images, named Universal One-shot Detection (UOD). UOD consists of two stages and two corresponding universal models which are designed as combinations of domain-specific modules and domain-shared modules. In the first stage, a domain-adaptive convolution model is self-supervised learned to generate pseudo landmark labels. In the second stage, we design a domain-adaptive transformer to eliminate domain preference and build the global context for multi-domain data. Even though only one annotated sample from each domain is available for training, the domain-shared modules help UOD aggregate all one-shot samples to detect more robust and accurate landmarks. We investigated both qualitatively and quantitatively the proposed UOD on three widely-used public X-ray datasets in different anatomical domains (i.e., head, hand, chest) and obtained state-of-the-art performances in each domain.
Continual Learning for Abdominal Multi-Organ and Tumor Segmentation
Jun 01, 2023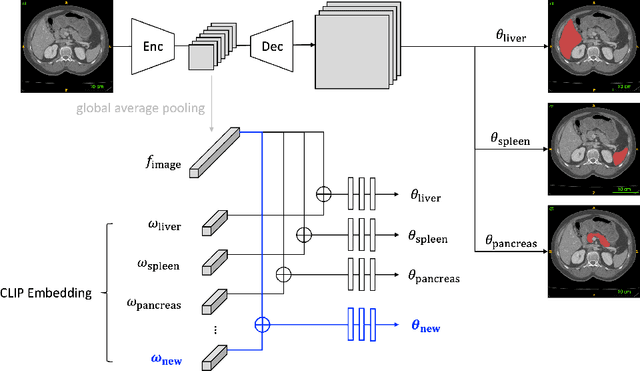
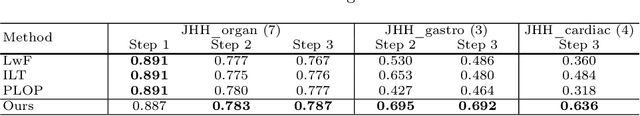
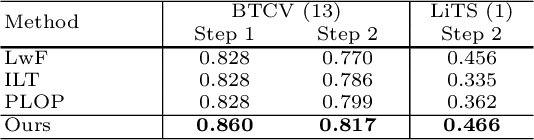
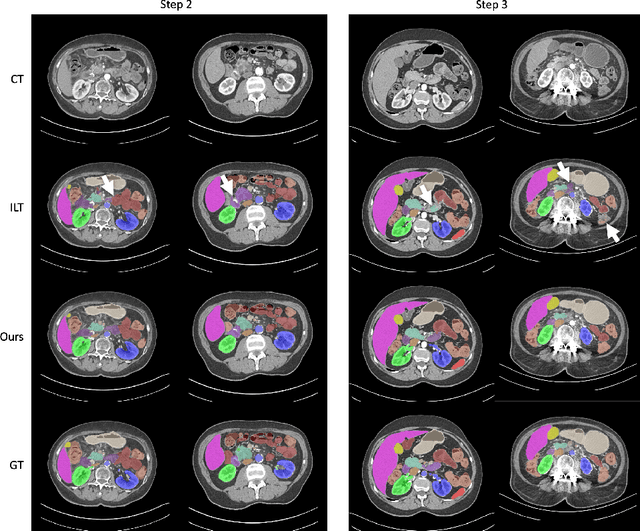
The ability to dynamically extend a model to new data and classes is critical for multiple organ and tumor segmentation. However, due to privacy regulations, accessing previous data and annotations can be problematic in the medical domain. This poses a significant barrier to preserving the high segmentation accuracy of the old classes when learning from new classes because of the catastrophic forgetting problem. In this paper, we first empirically demonstrate that simply using high-quality pseudo labels can fairly mitigate this problem in the setting of organ segmentation. Furthermore, we put forward an innovative architecture designed specifically for continuous organ and tumor segmentation, which incurs minimal computational overhead. Our proposed design involves replacing the conventional output layer with a suite of lightweight, class-specific heads, thereby offering the flexibility to accommodate newly emerging classes. These heads enable independent predictions for newly introduced and previously learned classes, effectively minimizing the impact of new classes on old ones during the course of continual learning. We further propose incorporating Contrastive Language-Image Pretraining (CLIP) embeddings into the organ-specific heads. These embeddings encapsulate the semantic information of each class, informed by extensive image-text co-training. The proposed method is evaluated on both in-house and public abdominal CT datasets under organ and tumor segmentation tasks. Empirical results suggest that the proposed design improves the segmentation performance of a baseline neural network on newly-introduced and previously-learned classes along the learning trajectory.
F?D: On understanding the role of deep feature spaces on face generation evaluation
Jun 01, 2023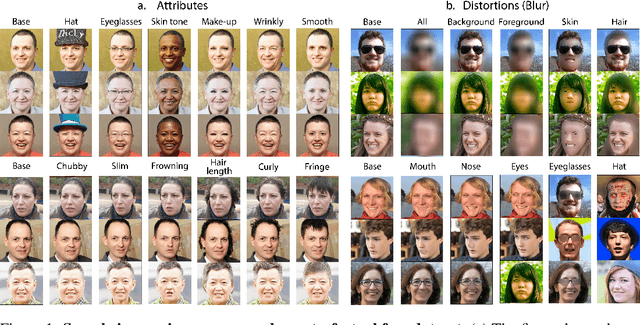
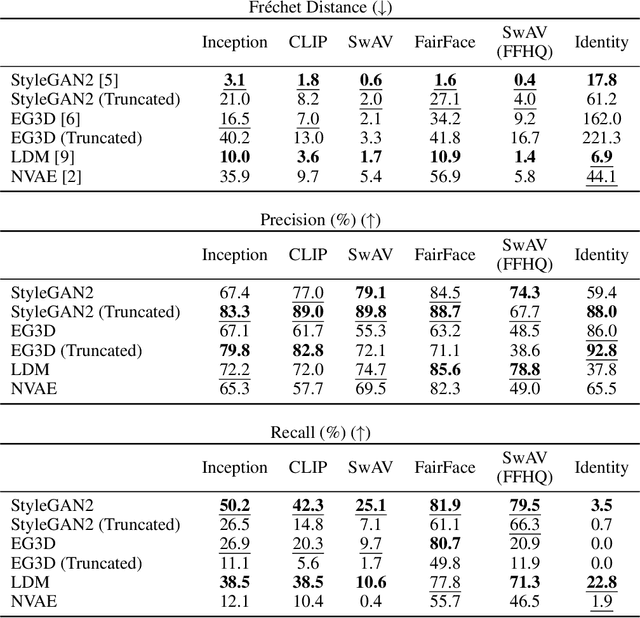
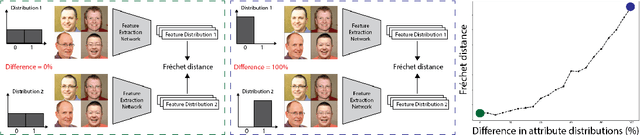
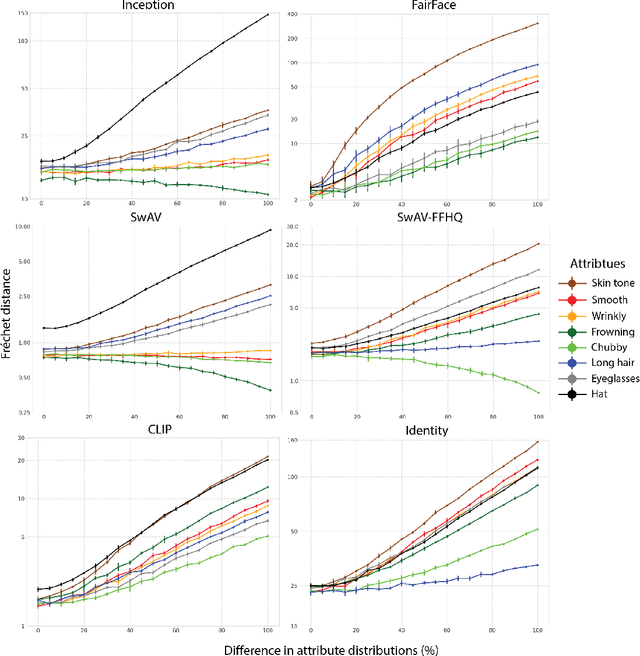
Perceptual metrics, like the Fr\'echet Inception Distance (FID), are widely used to assess the similarity between synthetically generated and ground truth (real) images. The key idea behind these metrics is to compute errors in a deep feature space that captures perceptually and semantically rich image features. Despite their popularity, the effect that different deep features and their design choices have on a perceptual metric has not been well studied. In this work, we perform a causal analysis linking differences in semantic attributes and distortions between face image distributions to Fr\'echet distances (FD) using several popular deep feature spaces. A key component of our analysis is the creation of synthetic counterfactual faces using deep face generators. Our experiments show that the FD is heavily influenced by its feature space's training dataset and objective function. For example, FD using features extracted from ImageNet-trained models heavily emphasize hats over regions like the eyes and mouth. Moreover, FD using features from a face gender classifier emphasize hair length more than distances in an identity (recognition) feature space. Finally, we evaluate several popular face generation models across feature spaces and find that StyleGAN2 consistently ranks higher than other face generators, except with respect to identity (recognition) features. This suggests the need for considering multiple feature spaces when evaluating generative models and using feature spaces that are tuned to nuances of the domain of interest.
LayoutDiffuse: Adapting Foundational Diffusion Models for Layout-to-Image Generation
Feb 16, 2023
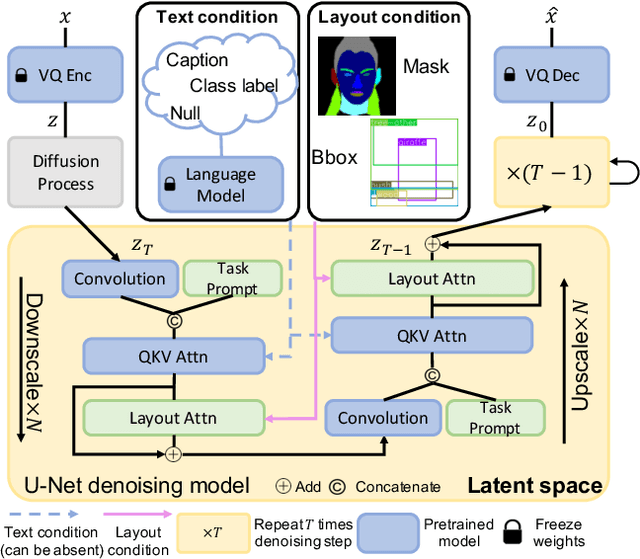
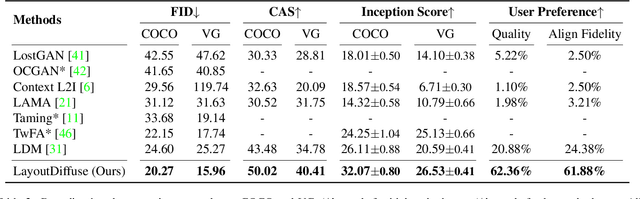
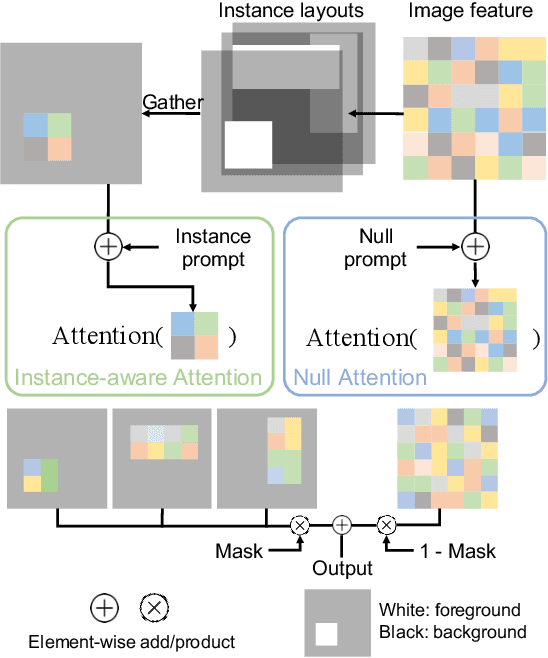
Layout-to-image generation refers to the task of synthesizing photo-realistic images based on semantic layouts. In this paper, we propose LayoutDiffuse that adapts a foundational diffusion model pretrained on large-scale image or text-image datasets for layout-to-image generation. By adopting a novel neural adaptor based on layout attention and task-aware prompts, our method trains efficiently, generates images with both high perceptual quality and layout alignment, and needs less data. Experiments on three datasets show that our method significantly outperforms other 10 generative models based on GANs, VQ-VAE, and diffusion models.
RealFusion: 360° Reconstruction of Any Object from a Single Image
Feb 21, 2023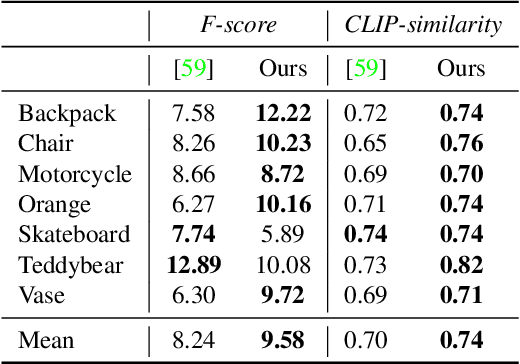
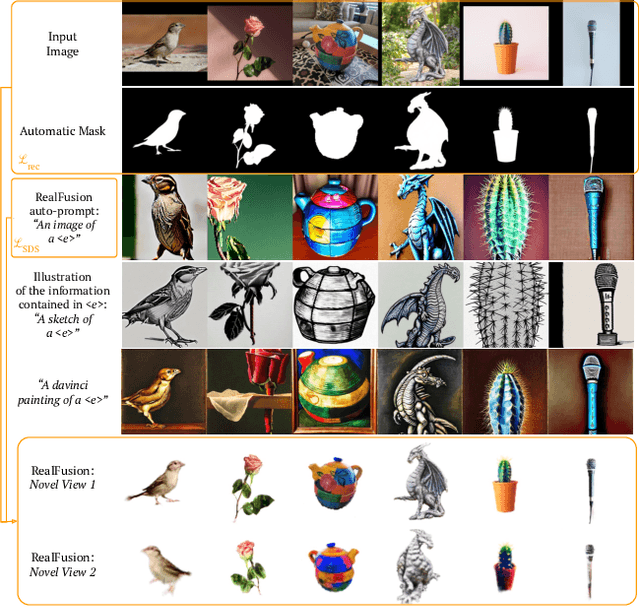
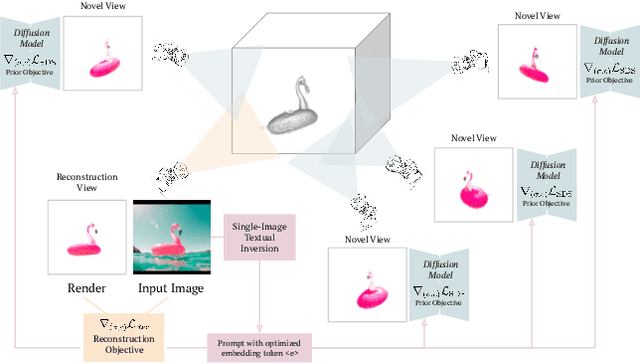


We consider the problem of reconstructing a full 360{\deg} photographic model of an object from a single image of it. We do so by fitting a neural radiance field to the image, but find this problem to be severely ill-posed. We thus take an off-the-self conditional image generator based on diffusion and engineer a prompt that encourages it to ``dream up'' novel views of the object. Using an approach inspired by DreamFields and DreamFusion, we fuse the given input view, the conditional prior, and other regularizers in a final, consistent reconstruction. We demonstrate state-of-the-art reconstruction results on benchmark images when compared to prior methods for monocular 3D reconstruction of objects. Qualitatively, our reconstructions provide a faithful match of the input view and a plausible extrapolation of its appearance and 3D shape, including to the side of the object not visible in the image.
Artificial Intelligence can facilitate selfish decisions by altering the appearance of interaction partners
Jun 07, 2023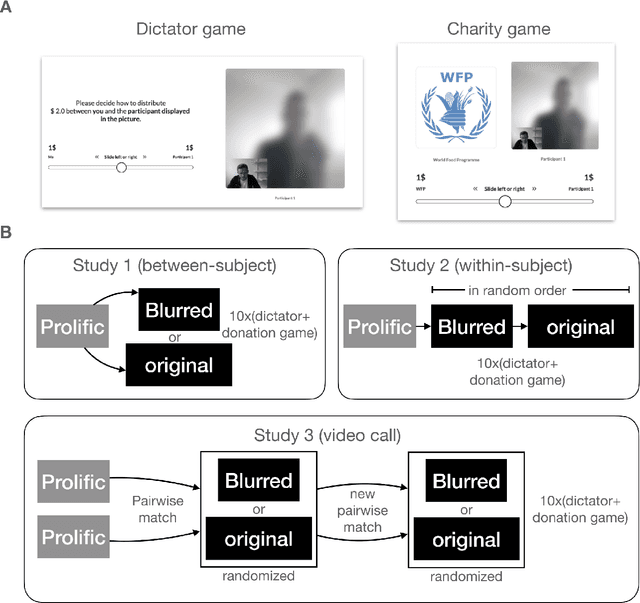
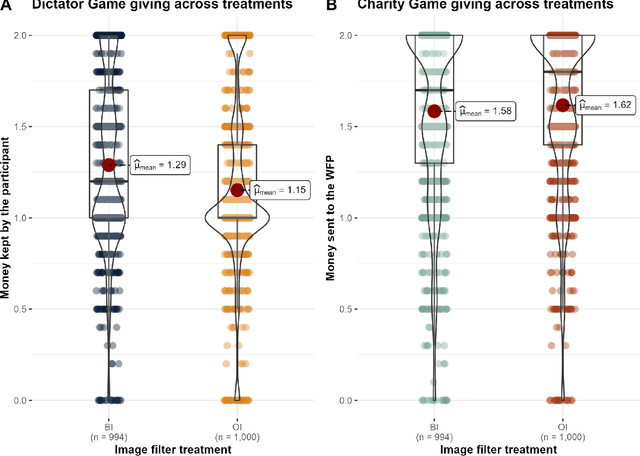
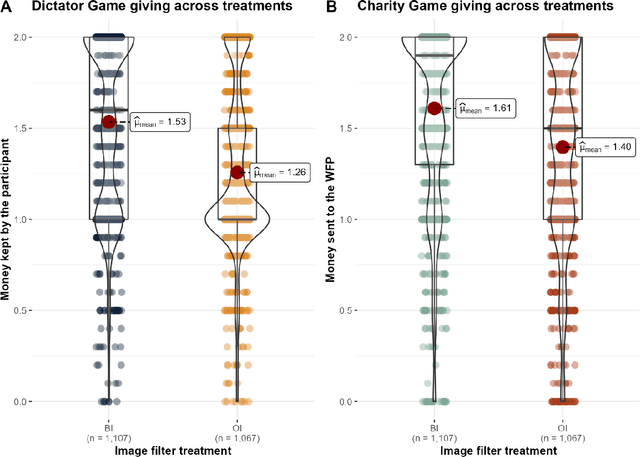
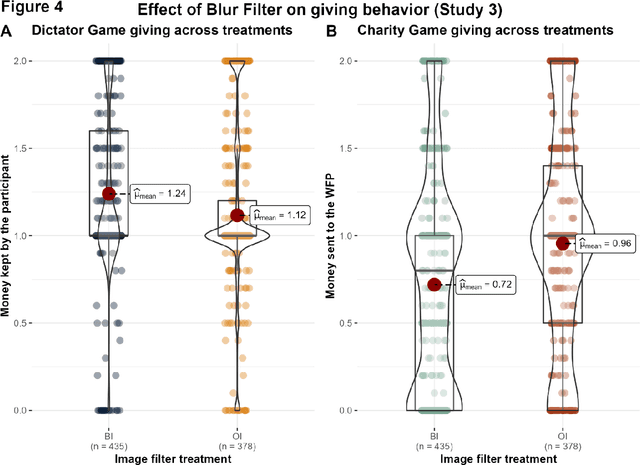
The increasing prevalence of image-altering filters on social media and video conferencing technologies has raised concerns about the ethical and psychological implications of using Artificial Intelligence (AI) to manipulate our perception of others. In this study, we specifically investigate the potential impact of blur filters, a type of appearance-altering technology, on individuals' behavior towards others. Our findings consistently demonstrate a significant increase in selfish behavior directed towards individuals whose appearance is blurred, suggesting that blur filters can facilitate moral disengagement through depersonalization. These results emphasize the need for broader ethical discussions surrounding AI technologies that modify our perception of others, including issues of transparency, consent, and the awareness of being subject to appearance manipulation by others. We also emphasize the importance of anticipatory experiments in informing the development of responsible guidelines and policies prior to the widespread adoption of such technologies.
Echocardiography Segmentation Using Neural ODE-based Diffeomorphic Registration Field
Jun 16, 2023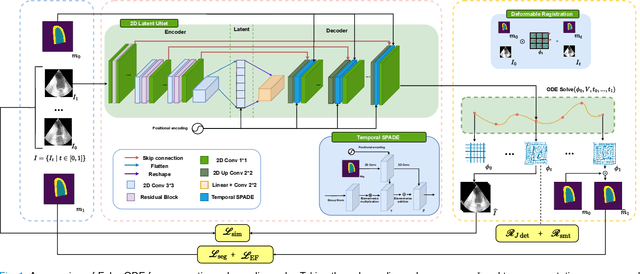
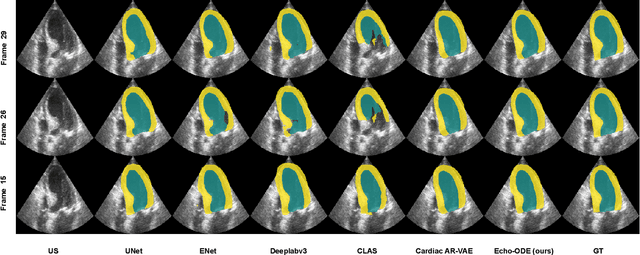
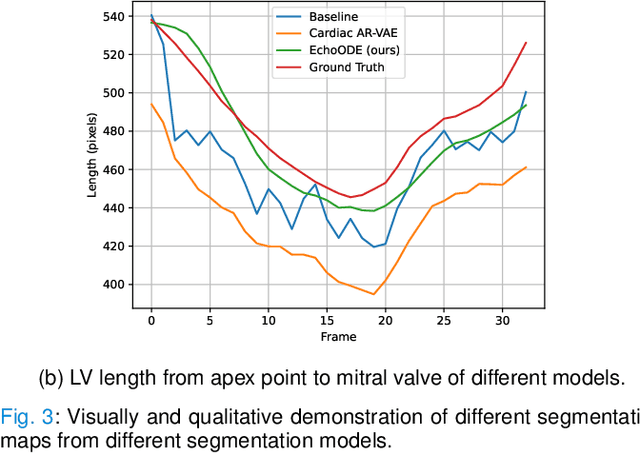
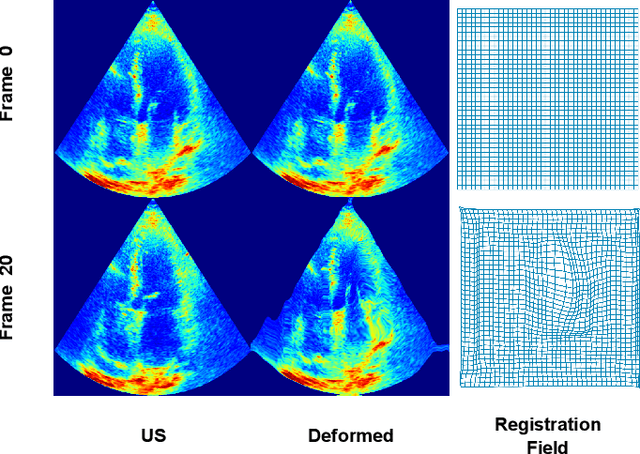
Convolutional neural networks (CNNs) have recently proven their excellent ability to segment 2D cardiac ultrasound images. However, the majority of attempts to perform full-sequence segmentation of cardiac ultrasound videos either rely on models trained only on keyframe images or fail to maintain the topology over time. To address these issues, in this work, we consider segmentation of ultrasound video as a registration estimation problem and present a novel method for diffeomorphic image registration using neural ordinary differential equations (Neural ODE). In particular, we consider the registration field vector field between frames as a continuous trajectory ODE. The estimated registration field is then applied to the segmentation mask of the first frame to obtain a segment for the whole cardiac cycle. The proposed method, Echo-ODE, introduces several key improvements compared to the previous state-of-the-art. Firstly, by solving a continuous ODE, the proposed method achieves smoother segmentation, preserving the topology of segmentation maps over the whole sequence (Hausdorff distance: 3.7-4.4). Secondly, it maintains temporal consistency between frames without explicitly optimizing for temporal consistency attributes, achieving temporal consistency in 91% of the videos in the dataset. Lastly, the proposed method is able to maintain the clinical accuracy of the segmentation maps (MAE of the LVEF: 2.7-3.1). The results show that our method surpasses the previous state-of-the-art in multiple aspects, demonstrating the importance of spatial-temporal data processing for the implementation of Neural ODEs in medical imaging applications. These findings open up new research directions for solving echocardiography segmentation tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge