"Image": models, code, and papers
Elongated Physiological Structure Segmentation via Spatial and Scale Uncertainty-aware Network
May 30, 2023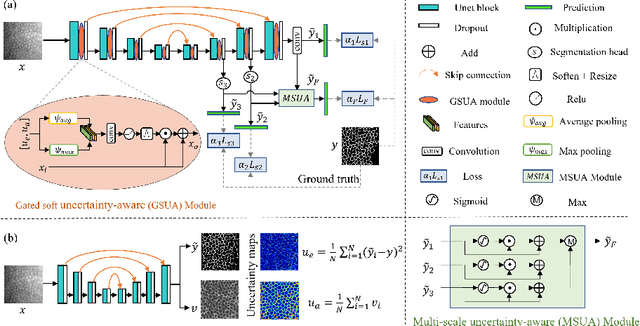
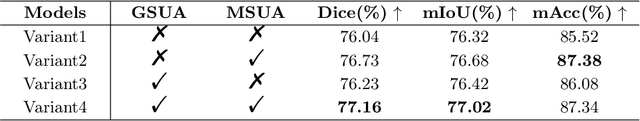
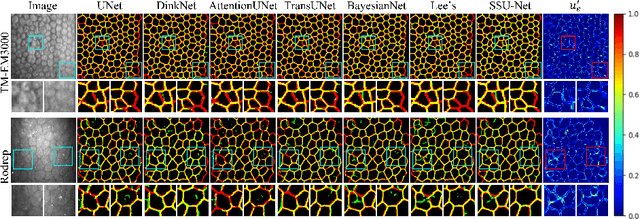
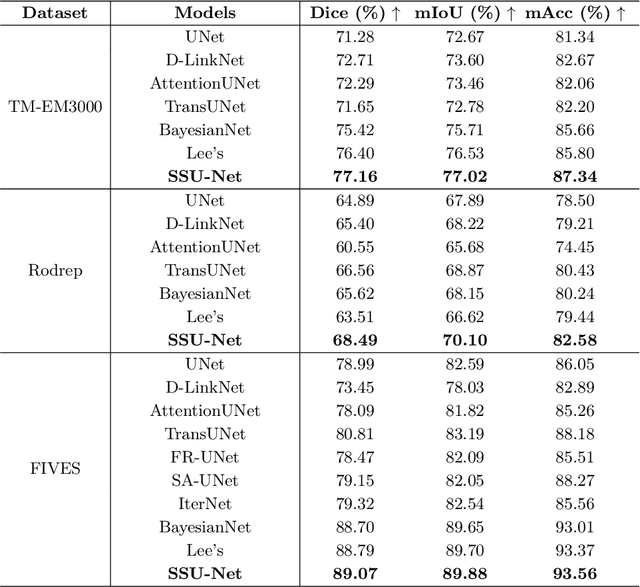
Robust and accurate segmentation for elongated physiological structures is challenging, especially in the ambiguous region, such as the corneal endothelium microscope image with uneven illumination or the fundus image with disease interference. In this paper, we present a spatial and scale uncertainty-aware network (SSU-Net) that fully uses both spatial and scale uncertainty to highlight ambiguous regions and integrate hierarchical structure contexts. First, we estimate epistemic and aleatoric spatial uncertainty maps using Monte Carlo dropout to approximate Bayesian networks. Based on these spatial uncertainty maps, we propose the gated soft uncertainty-aware (GSUA) module to guide the model to focus on ambiguous regions. Second, we extract the uncertainty under different scales and propose the multi-scale uncertainty-aware (MSUA) fusion module to integrate structure contexts from hierarchical predictions, strengthening the final prediction. Finally, we visualize the uncertainty map of final prediction, providing interpretability for segmentation results. Experiment results show that the SSU-Net performs best on cornea endothelial cell and retinal vessel segmentation tasks. Moreover, compared with counterpart uncertainty-based methods, SSU-Net is more accurate and robust.
Joint Optimization of Class-Specific Training- and Test-Time Data Augmentation in Segmentation
May 30, 2023
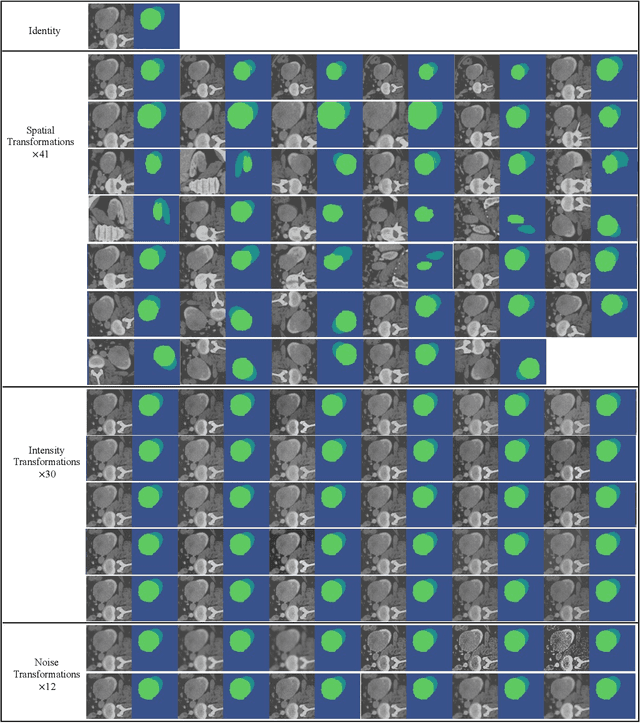
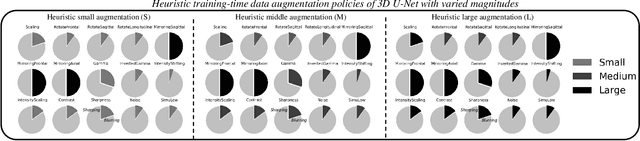
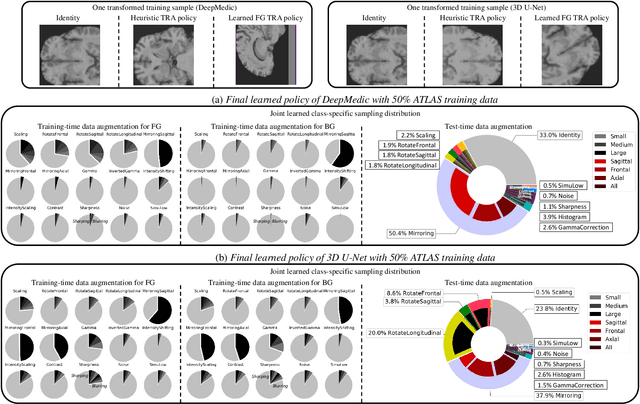
This paper presents an effective and general data augmentation framework for medical image segmentation. We adopt a computationally efficient and data-efficient gradient-based meta-learning scheme to explicitly align the distribution of training and validation data which is used as a proxy for unseen test data. We improve the current data augmentation strategies with two core designs. First, we learn class-specific training-time data augmentation (TRA) effectively increasing the heterogeneity within the training subsets and tackling the class imbalance common in segmentation. Second, we jointly optimize TRA and test-time data augmentation (TEA), which are closely connected as both aim to align the training and test data distribution but were so far considered separately in previous works. We demonstrate the effectiveness of our method on four medical image segmentation tasks across different scenarios with two state-of-the-art segmentation models, DeepMedic and nnU-Net. Extensive experimentation shows that the proposed data augmentation framework can significantly and consistently improve the segmentation performance when compared to existing solutions. Code is publicly available.
Compression with Bayesian Implicit Neural Representations
May 30, 2023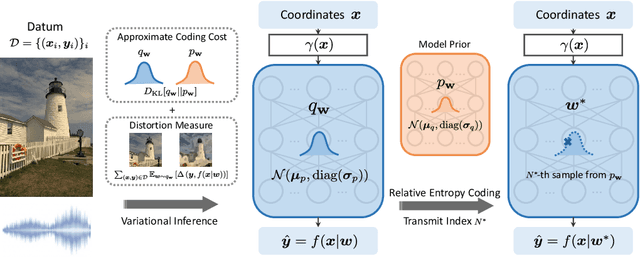
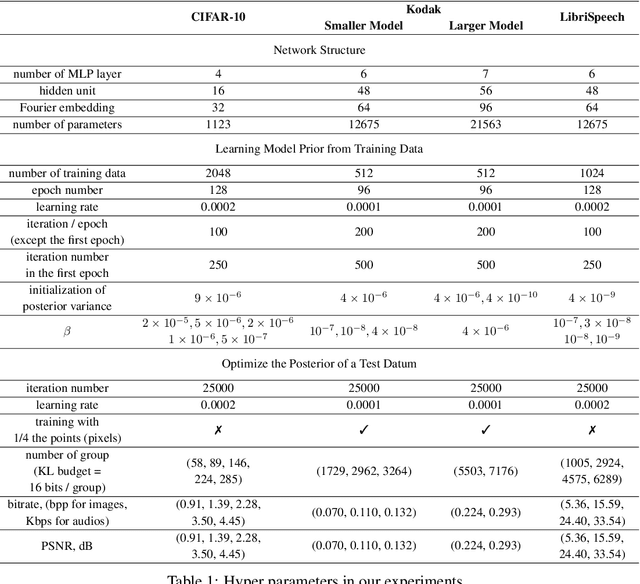
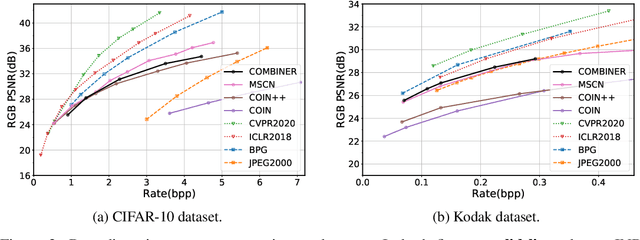

Many common types of data can be represented as functions that map coordinates to signal values, such as pixel locations to RGB values in the case of an image. Based on this view, data can be compressed by overfitting a compact neural network to its functional representation and then encoding the network weights. However, most current solutions for this are inefficient, as quantization to low-bit precision substantially degrades the reconstruction quality. To address this issue, we propose overfitting variational Bayesian neural networks to the data and compressing an approximate posterior weight sample using relative entropy coding instead of quantizing and entropy coding it. This strategy enables direct optimization of the rate-distortion performance by minimizing the $\beta$-ELBO, and target different rate-distortion trade-offs for a given network architecture by adjusting $\beta$. Moreover, we introduce an iterative algorithm for learning prior weight distributions and employ a progressive refinement process for the variational posterior that significantly enhances performance. Experiments show that our method achieves strong performance on image and audio compression while retaining simplicity.
User-defined Event Sampling and Uncertainty Quantification in Diffusion Models for Physical Dynamical Systems
Jun 13, 2023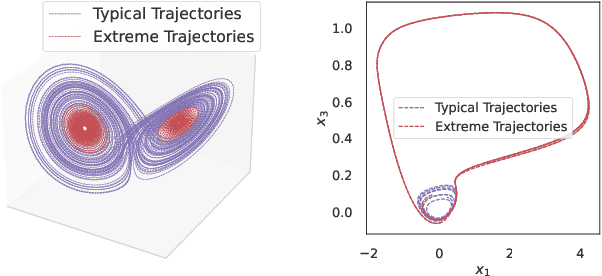
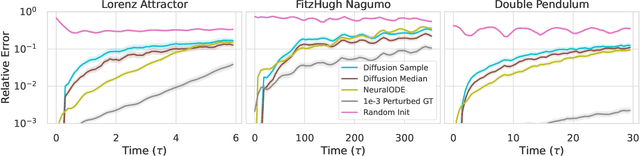
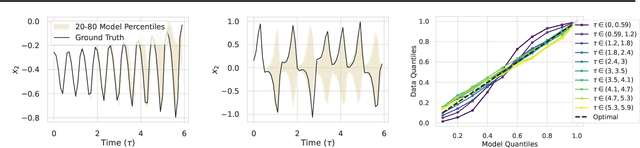
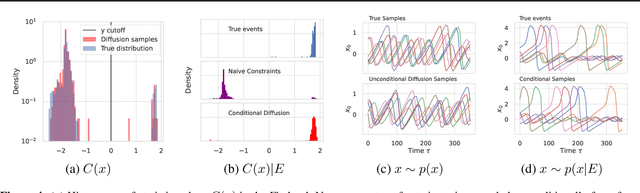
Diffusion models are a class of probabilistic generative models that have been widely used as a prior for image processing tasks like text conditional generation and inpainting. We demonstrate that these models can be adapted to make predictions and provide uncertainty quantification for chaotic dynamical systems. In these applications, diffusion models can implicitly represent knowledge about outliers and extreme events; however, querying that knowledge through conditional sampling or measuring probabilities is surprisingly difficult. Existing methods for conditional sampling at inference time seek mainly to enforce the constraints, which is insufficient to match the statistics of the distribution or compute the probability of the chosen events. To achieve these ends, optimally one would use the conditional score function, but its computation is typically intractable. In this work, we develop a probabilistic approximation scheme for the conditional score function which provably converges to the true distribution as the noise level decreases. With this scheme we are able to sample conditionally on nonlinear userdefined events at inference time, and matches data statistics even when sampling from the tails of the distribution.
MetaMorph: Learning Metamorphic Image Transformation With Appearance Changes
Mar 08, 2023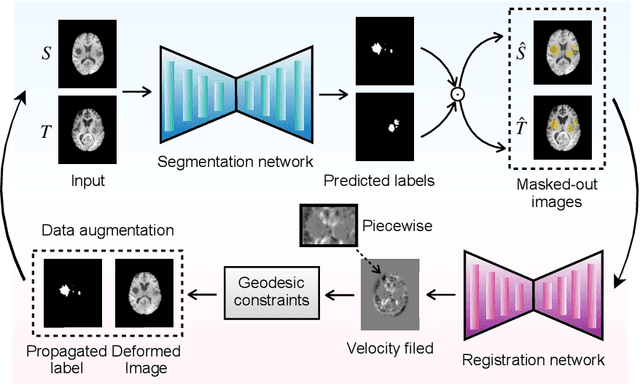
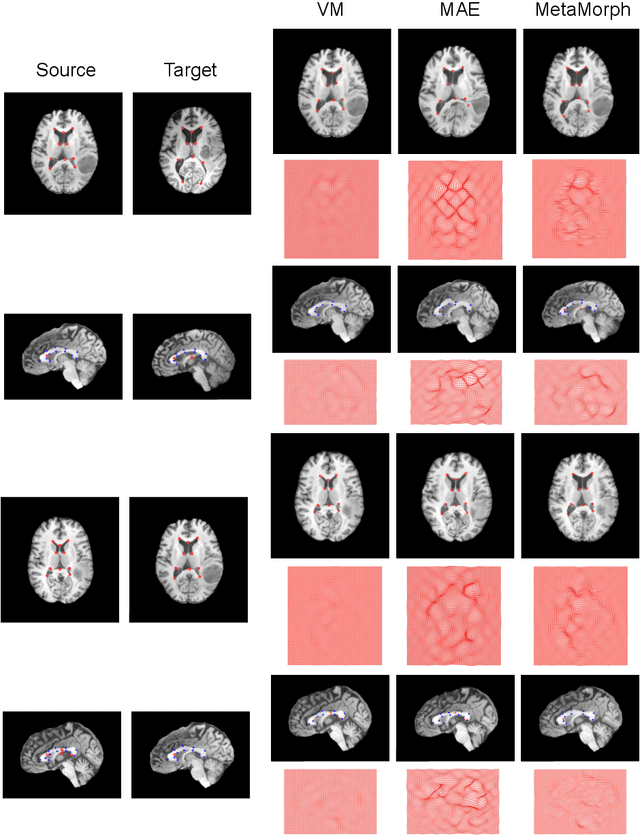
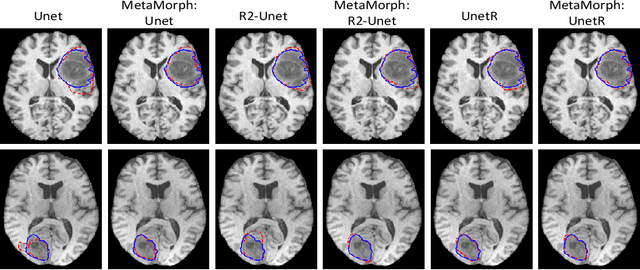
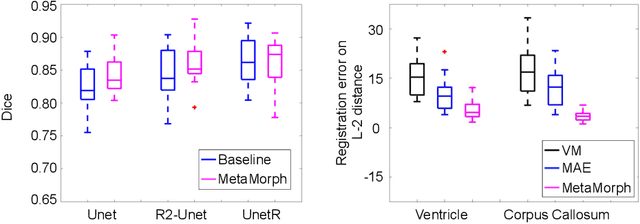
This paper presents a novel predictive model, MetaMorph, for metamorphic registration of images with appearance changes (i.e., caused by brain tumors). In contrast to previous learning-based registration methods that have little or no control over appearance-changes, our model introduces a new regularization that can effectively suppress the negative effects of appearance changing areas. In particular, we develop a piecewise regularization on the tangent space of diffeomorphic transformations (also known as initial velocity fields) via learned segmentation maps of abnormal regions. The geometric transformation and appearance changes are treated as joint tasks that are mutually beneficial. Our model MetaMorph is more robust and accurate when searching for an optimal registration solution under the guidance of segmentation, which in turn improves the segmentation performance by providing appropriately augmented training labels. We validate MetaMorph on real 3D human brain tumor magnetic resonance imaging (MRI) scans. Experimental results show that our model outperforms the state-of-the-art learning-based registration models. The proposed MetaMorph has great potential in various image-guided clinical interventions, e.g., real-time image-guided navigation systems for tumor removal surgery.
Forget-Me-Not: Learning to Forget in Text-to-Image Diffusion Models
Mar 30, 2023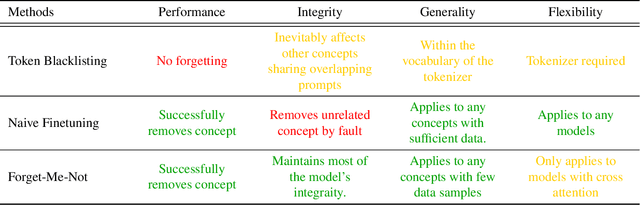
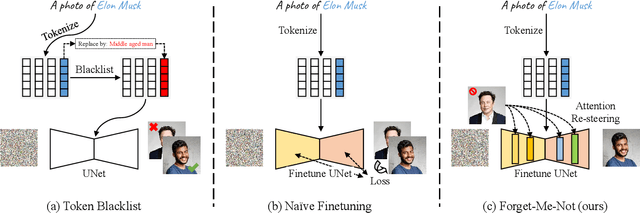
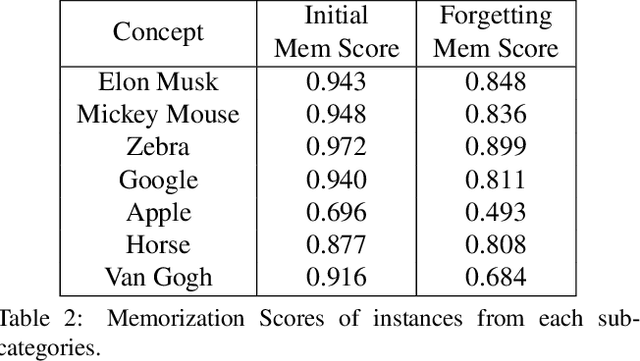
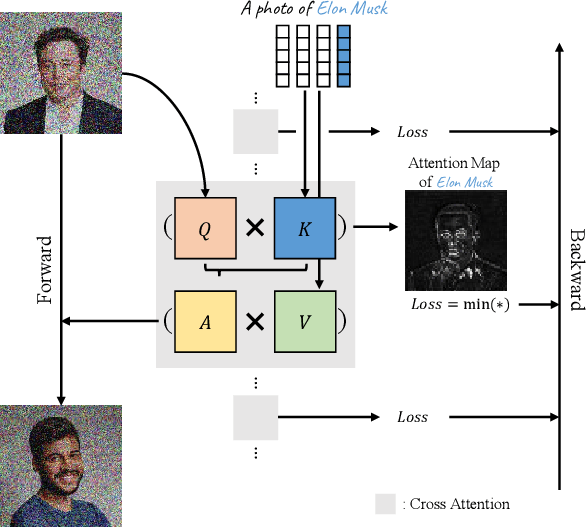
The unlearning problem of deep learning models, once primarily an academic concern, has become a prevalent issue in the industry. The significant advances in text-to-image generation techniques have prompted global discussions on privacy, copyright, and safety, as numerous unauthorized personal IDs, content, artistic creations, and potentially harmful materials have been learned by these models and later utilized to generate and distribute uncontrolled content. To address this challenge, we propose \textbf{Forget-Me-Not}, an efficient and low-cost solution designed to safely remove specified IDs, objects, or styles from a well-configured text-to-image model in as little as 30 seconds, without impairing its ability to generate other content. Alongside our method, we introduce the \textbf{Memorization Score (M-Score)} and \textbf{ConceptBench} to measure the models' capacity to generate general concepts, grouped into three primary categories: ID, object, and style. Using M-Score and ConceptBench, we demonstrate that Forget-Me-Not can effectively eliminate targeted concepts while maintaining the model's performance on other concepts. Furthermore, Forget-Me-Not offers two practical extensions: a) removal of potentially harmful or NSFW content, and b) enhancement of model accuracy, inclusion and diversity through \textbf{concept correction and disentanglement}. It can also be adapted as a lightweight model patch for Stable Diffusion, allowing for concept manipulation and convenient distribution. To encourage future research in this critical area and promote the development of safe and inclusive generative models, we will open-source our code and ConceptBench at \href{https://github.com/SHI-Labs/Forget-Me-Not}{https://github.com/SHI-Labs/Forget-Me-Not}.
Beyond Surface Statistics: Scene Representations in a Latent Diffusion Model
Jun 09, 2023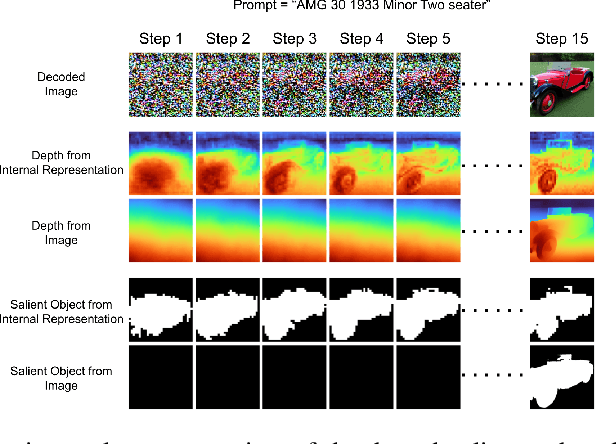
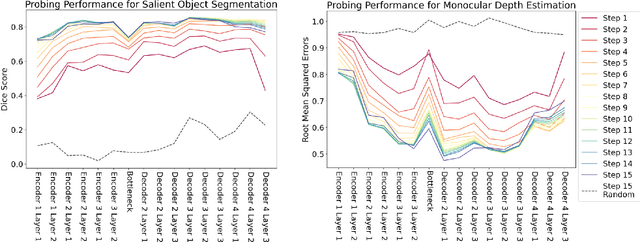
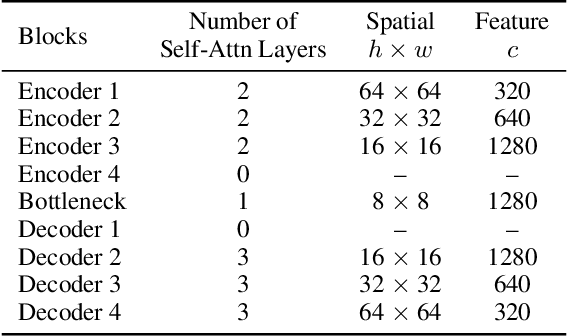
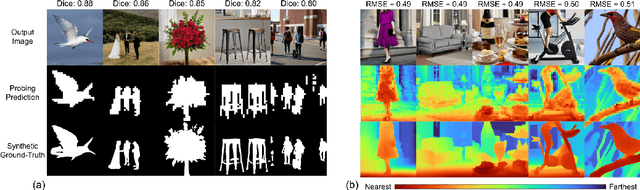
Latent diffusion models (LDMs) exhibit an impressive ability to produce realistic images, yet the inner workings of these models remain mysterious. Even when trained purely on images without explicit depth information, they typically output coherent pictures of 3D scenes. In this work, we investigate a basic interpretability question: does an LDM create and use an internal representation of simple scene geometry? Using linear probes, we find evidence that the internal activations of the LDM encode linear representations of both 3D depth data and a salient-object / background distinction. These representations appear surprisingly early in the denoising process$-$well before a human can easily make sense of the noisy images. Intervention experiments further indicate these representations play a causal role in image synthesis, and may be used for simple high-level editing of an LDM's output.
Boosting GUI Prototyping with Diffusion Models
Jun 09, 2023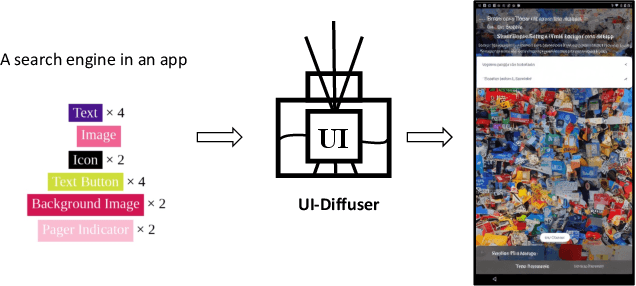
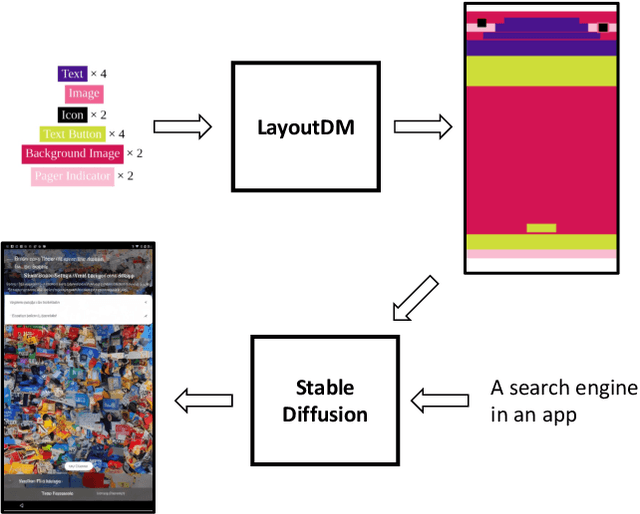
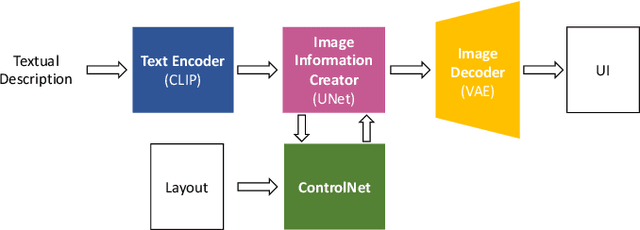
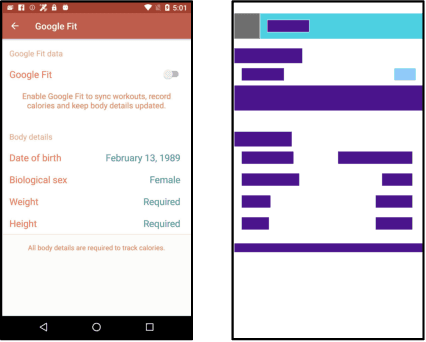
GUI (graphical user interface) prototyping is a widely-used technique in requirements engineering for gathering and refining requirements, reducing development risks and increasing stakeholder engagement. However, GUI prototyping can be a time-consuming and costly process. In recent years, deep learning models such as Stable Diffusion have emerged as a powerful text-to-image tool capable of generating detailed images based on text prompts. In this paper, we propose UI-Diffuser, an approach that leverages Stable Diffusion to generate mobile UIs through simple textual descriptions and UI components. Preliminary results show that UI-Diffuser provides an efficient and cost-effective way to generate mobile GUI designs while reducing the need for extensive prototyping efforts. This approach has the potential to significantly improve the speed and efficiency of GUI prototyping in requirements engineering.
DAGrid: Directed Accumulator Grid
Jun 05, 2023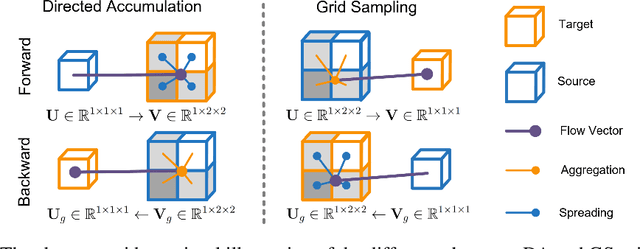
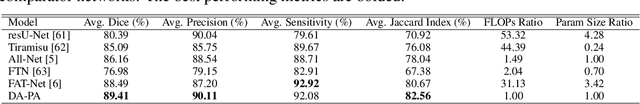
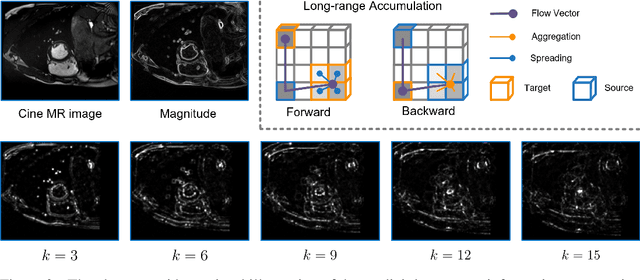
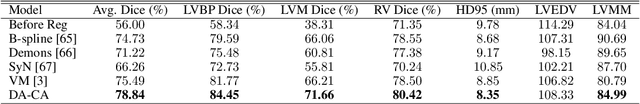
Recent research highlights that the Directed Accumulator (DA), through its parametrization of geometric priors into neural networks, has notably improved the performance of medical image recognition, particularly with small and imbalanced datasets. However, DA's potential in pixel-wise dense predictions is unexplored. To bridge this gap, we present the Directed Accumulator Grid (DAGrid), which allows geometric-preserving filtering in neural networks, thus broadening the scope of DA's applications to include pixel-level dense prediction tasks. DAGrid utilizes homogeneous data types in conjunction with designed sampling grids to construct geometrically transformed representations, retaining intricate geometric information and promoting long-range information propagation within the neural networks. Contrary to its symmetric counterpart, grid sampling, which might lose information in the sampling process, DAGrid aggregates all pixels, ensuring a comprehensive representation in the transformed space. The parallelization of DAGrid on modern GPUs is facilitated using CUDA programming, and also back propagation is enabled for deep neural network training. Empirical results show DAGrid-enhanced neural networks excel in supervised skin lesion segmentation and unsupervised cardiac image registration. Specifically, the network incorporating DAGrid has realized a 70.8% reduction in network parameter size and a 96.8% decrease in FLOPs, while concurrently improving the Dice score for skin lesion segmentation by 1.0% compared to state-of-the-art transformers. Furthermore, it has achieved improvements of 4.4% and 8.2% in the average Dice score and Dice score of the left ventricular mass, respectively, indicating an increase in registration accuracy for cardiac images. The source code is available at https://github.com/tinymilky/DeDA.
Masked Collaborative Contrast for Weakly Supervised Semantic Segmentation
May 19, 2023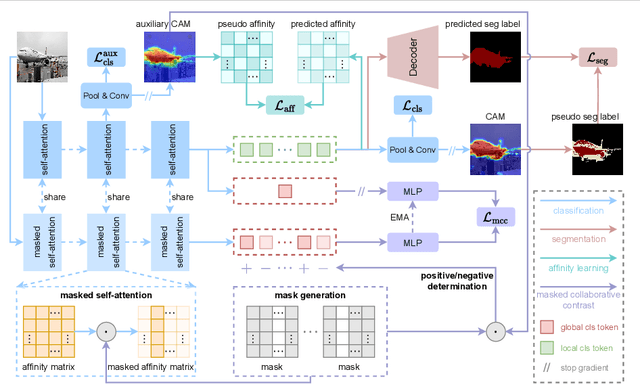
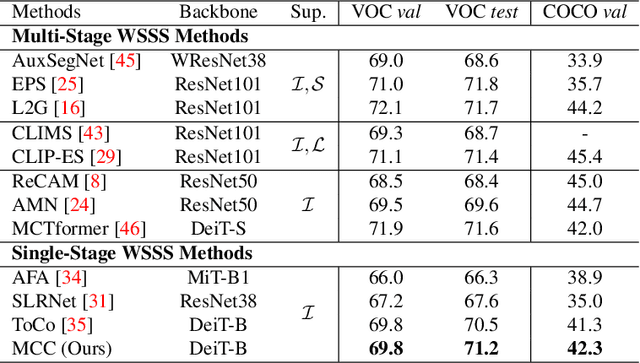
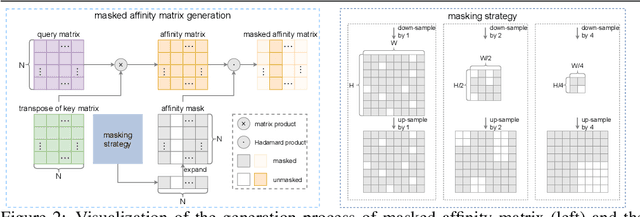
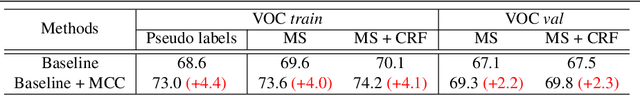
This study introduces an efficacious approach, Masked Collaborative Contrast (MCC), to emphasize semantic regions in weakly supervised semantic segmentation. MCC adroitly incorporates concepts from masked image modeling and contrastive learning to devise Transformer blocks that induce keys to contract towards semantically pertinent regions. Unlike prevalent techniques that directly eradicate patch regions in the input image when generating masks, we scrutinize the neighborhood relations of patch tokens by exploring masks considering keys on the affinity matrix. Moreover, we generate positive and negative samples in contrastive learning by utilizing the masked local output and contrasting it with the global output. Elaborate experiments on commonly employed datasets evidences that the proposed MCC mechanism effectively aligns global and local perspectives within the image, attaining impressive performance. The source code is available at \url{https://github.com/fwu11/MCC}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge