"Image": models, code, and papers
Locally Differentially Private Distributed Online Learning with Guaranteed Optimality
Jun 25, 2023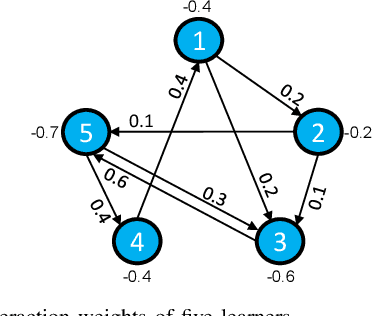
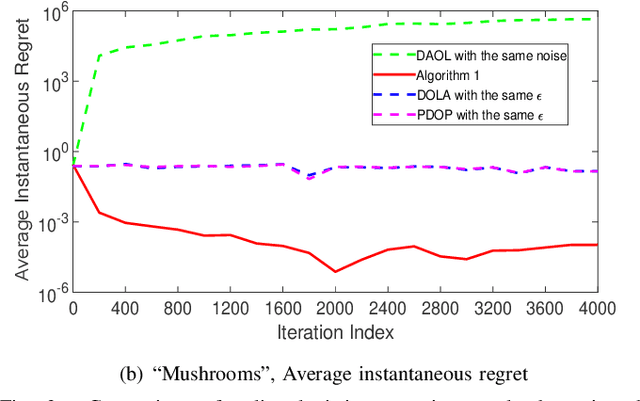
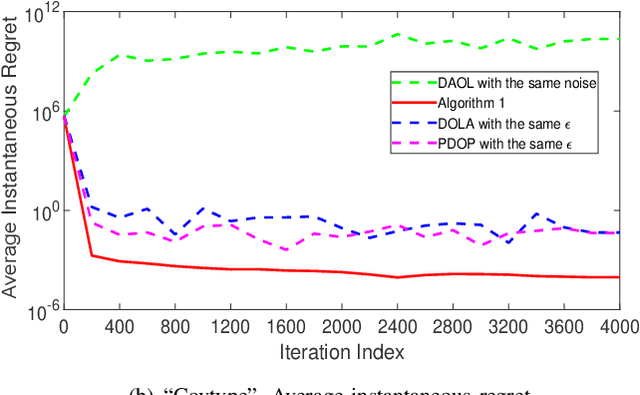
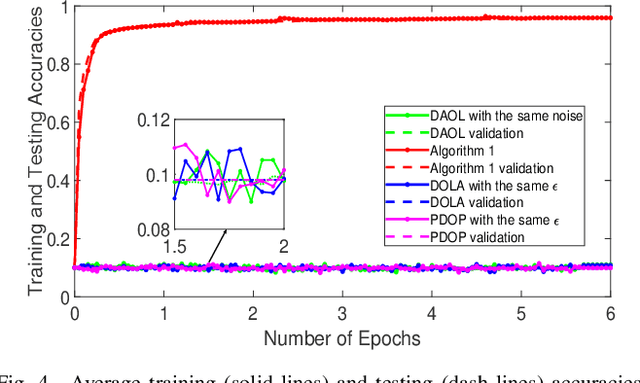
Distributed online learning is gaining increased traction due to its unique ability to process large-scale datasets and streaming data. To address the growing public awareness and concern on privacy protection, plenty of private distributed online learning algorithms have been proposed, mostly based on differential privacy which has emerged as the ``gold standard" for privacy protection. However, these algorithms often face the dilemma of trading learning accuracy for privacy. By exploiting the unique characteristics of online learning, this paper proposes an approach that tackles the dilemma and ensures both differential privacy and learning accuracy in distributed online learning. More specifically, while ensuring a diminishing expected instantaneous regret, the approach can simultaneously ensure a finite cumulative privacy budget, even on the infinite time horizon. To cater for the fully distributed setting, we adopt the local differential-privacy framework which avoids the reliance on a trusted data curator, and hence, provides stronger protection than the classic ``centralized" (global) differential privacy. To the best of our knowledge, this is the first algorithm that successfully ensures both rigorous local differential privacy and learning accuracy. The effectiveness of the proposed algorithm is evaluated using machine learning tasks, including logistic regression on the ``Mushrooms" and ``Covtype" datasets and CNN based image classification on the ``MNIST" and ``CIFAR-10" datasets.
Medical supervised masked autoencoders: Crafting a better masking strategy and efficient fine-tuning schedule for medical image classification
May 10, 2023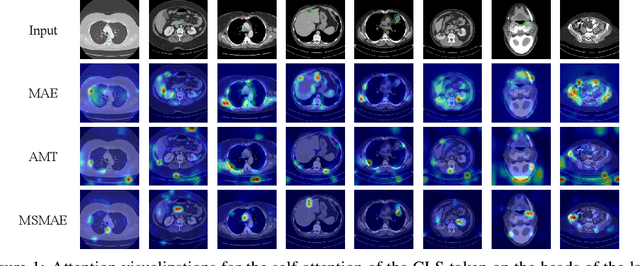
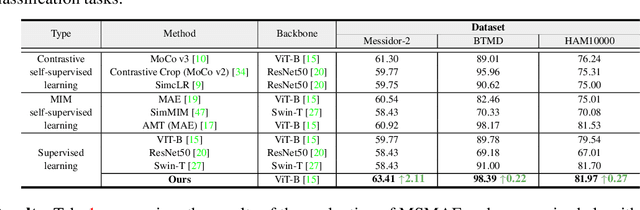
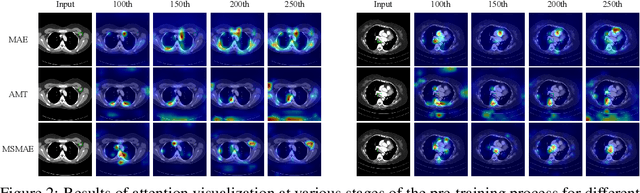
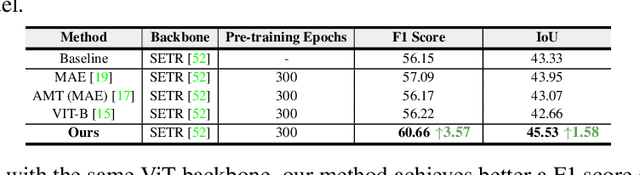
Masked autoencoders (MAEs) have displayed significant potential in the classification and semantic segmentation of medical images in the last year. Due to the high similarity of human tissues, even slight changes in medical images may represent diseased tissues, necessitating fine-grained inspection to pinpoint diseased tissues. The random masking strategy of MAEs is likely to result in areas of lesions being overlooked by the model. At the same time, inconsistencies between the pre-training and fine-tuning phases impede the performance and efficiency of MAE in medical image classification. To address these issues, we propose a medical supervised masked autoencoder (MSMAE) in this paper. In the pre-training phase, MSMAE precisely masks medical images via the attention maps obtained from supervised training, contributing to the representation learning of human tissue in the lesion area. During the fine-tuning phase, MSMAE is also driven by attention to the accurate masking of medical images. This improves the computational efficiency of the MSMAE while increasing the difficulty of fine-tuning, which indirectly improves the quality of MSMAE medical diagnosis. Extensive experiments demonstrate that MSMAE achieves state-of-the-art performance in case with three official medical datasets for various diseases. Meanwhile, transfer learning for MSMAE also demonstrates the great potential of our approach for medical semantic segmentation tasks. Moreover, the MSMAE accelerates the inference time in the fine-tuning phase by 11.2% and reduces the number of floating-point operations (FLOPs) by 74.08% compared to a traditional MAE.
MindDiffuser: Controlled Image Reconstruction from Human Brain Activity with Semantic and Structural Diffusion
Mar 24, 2023
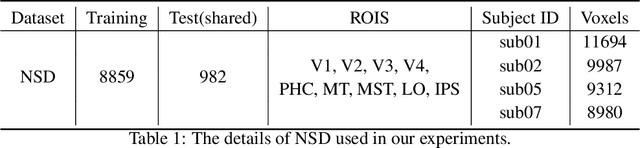

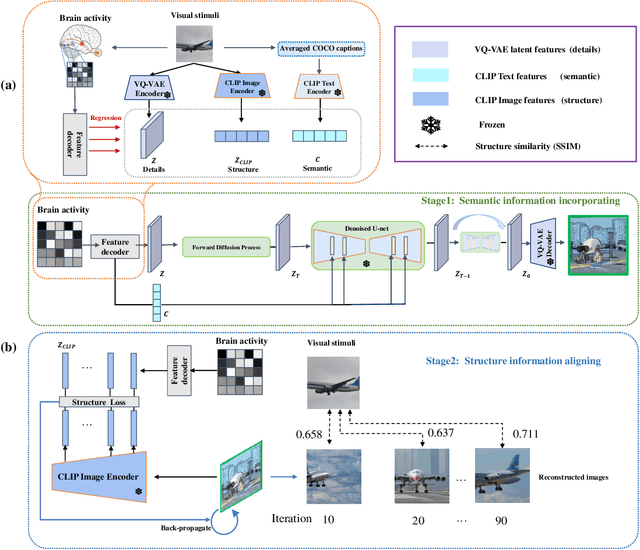
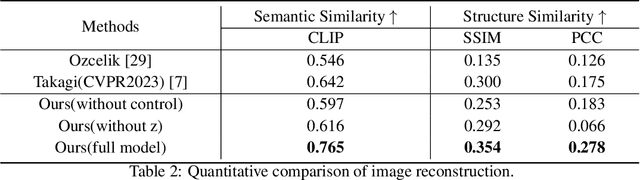
Reconstructing visual stimuli from measured functional magnetic resonance imaging (fMRI) has been a meaningful and challenging task. Previous studies have successfully achieved reconstructions with structures similar to the original images, such as the outlines and size of some natural images. However, these reconstructions lack explicit semantic information and are difficult to discern. In recent years, many studies have utilized multi-modal pre-trained models with stronger generative capabilities to reconstruct images that are semantically similar to the original ones. However, these images have uncontrollable structural information such as position and orientation. To address both of the aforementioned issues simultaneously, we propose a two-stage image reconstruction model called MindDiffuser, utilizing Stable Diffusion. In Stage 1, the VQ-VAE latent representations and the CLIP text embeddings decoded from fMRI are put into the image-to-image process of Stable Diffusion, which yields a preliminary image that contains semantic and structural information. In Stage 2, we utilize the low-level CLIP visual features decoded from fMRI as supervisory information, and continually adjust the two features in Stage 1 through backpropagation to align the structural information. The results of both qualitative and quantitative analyses demonstrate that our proposed model has surpassed the current state-of-the-art models in terms of reconstruction results on Natural Scenes Dataset (NSD). Furthermore, the results of ablation experiments indicate that each component of our model is effective for image reconstruction.
Democratizing Pathological Image Segmentation with Lay Annotators via Molecular-empowered Learning
May 31, 2023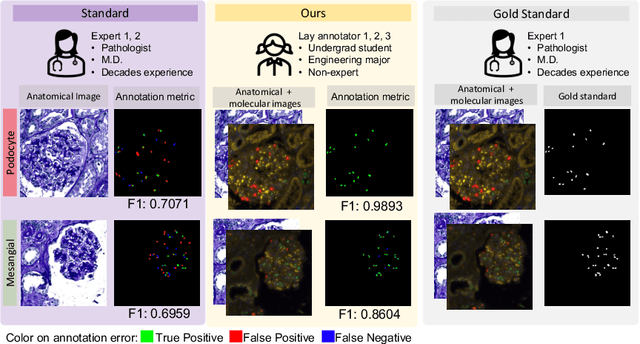
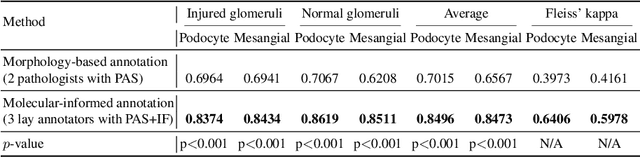
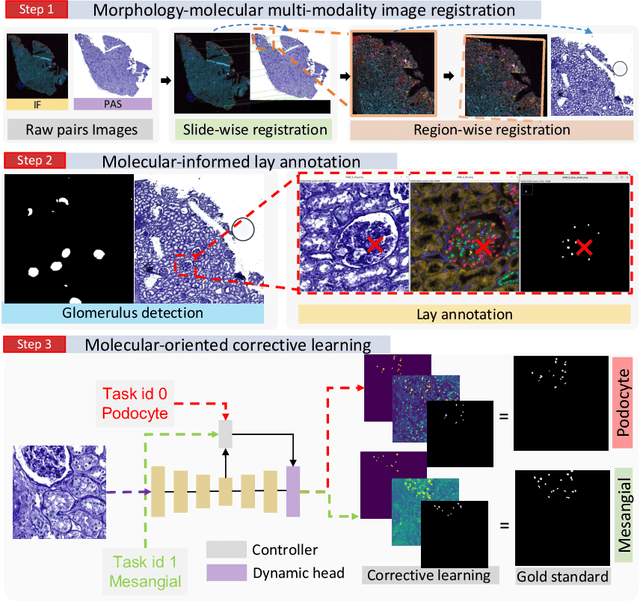
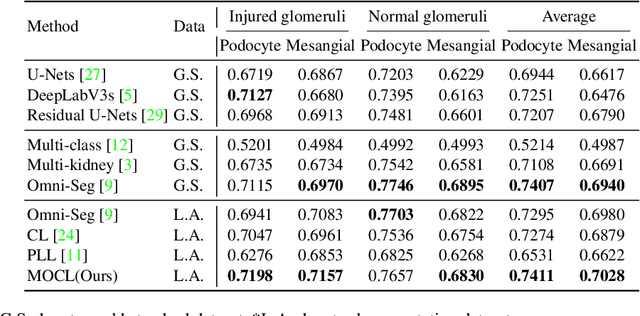
Multi-class cell segmentation in high-resolution Giga-pixel whole slide images (WSI) is critical for various clinical applications. Training such an AI model typically requires labor-intensive pixel-wise manual annotation from experienced domain experts (e.g., pathologists). Moreover, such annotation is error-prone when differentiating fine-grained cell types (e.g., podocyte and mesangial cells) via the naked human eye. In this study, we assess the feasibility of democratizing pathological AI deployment by only using lay annotators (annotators without medical domain knowledge). The contribution of this paper is threefold: (1) We proposed a molecular-empowered learning scheme for multi-class cell segmentation using partial labels from lay annotators; (2) The proposed method integrated Giga-pixel level molecular-morphology cross-modality registration, molecular-informed annotation, and molecular-oriented segmentation model, so as to achieve significantly superior performance via 3 lay annotators as compared with 2 experienced pathologists; (3) A deep corrective learning (learning with imperfect label) method is proposed to further improve the segmentation performance using partially annotated noisy data. From the experimental results, our learning method achieved F1 = 0.8496 using molecular-informed annotations from lay annotators, which is better than conventional morphology-based annotations (F1 = 0.7051) from experienced pathologists. Our method democratizes the development of a pathological segmentation deep model to the lay annotator level, which consequently scales up the learning process similar to a non-medical computer vision task. The official implementation and cell annotations are publicly available at https://github.com/hrlblab/MolecularEL.
MedGen3D: A Deep Generative Framework for Paired 3D Image and Mask Generation
Apr 08, 2023
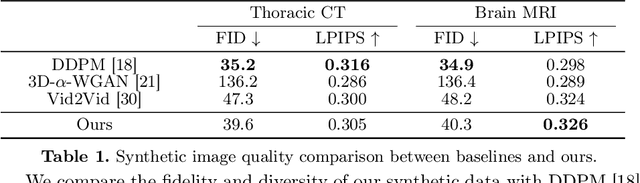
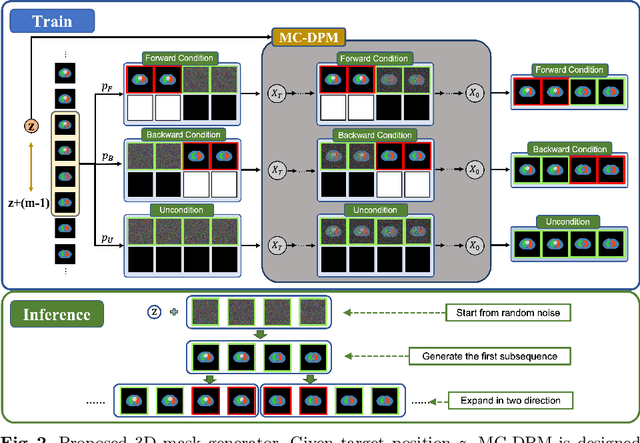
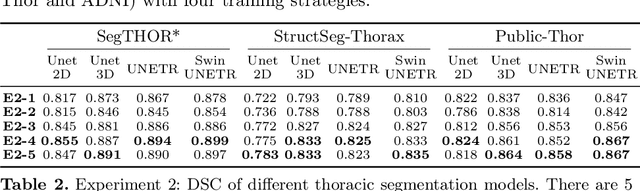
Acquiring and annotating sufficient labeled data is crucial in developing accurate and robust learning-based models, but obtaining such data can be challenging in many medical image segmentation tasks. One promising solution is to synthesize realistic data with ground-truth mask annotations. However, no prior studies have explored generating complete 3D volumetric images with masks. In this paper, we present MedGen3D, a deep generative framework that can generate paired 3D medical images and masks. First, we represent the 3D medical data as 2D sequences and propose the Multi-Condition Diffusion Probabilistic Model (MC-DPM) to generate multi-label mask sequences adhering to anatomical geometry. Then, we use an image sequence generator and semantic diffusion refiner conditioned on the generated mask sequences to produce realistic 3D medical images that align with the generated masks. Our proposed framework guarantees accurate alignment between synthetic images and segmentation maps. Experiments on 3D thoracic CT and brain MRI datasets show that our synthetic data is both diverse and faithful to the original data, and demonstrate the benefits for downstream segmentation tasks. We anticipate that MedGen3D's ability to synthesize paired 3D medical images and masks will prove valuable in training deep learning models for medical imaging tasks.
Object-Centric Learning for Real-World Videos by Predicting Temporal Feature Similarities
Jun 07, 2023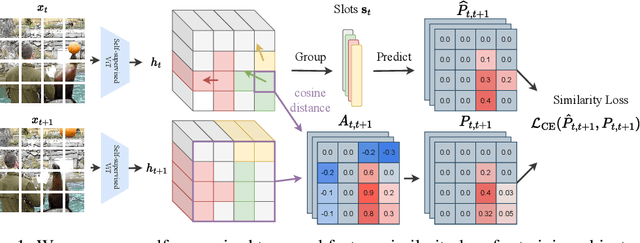
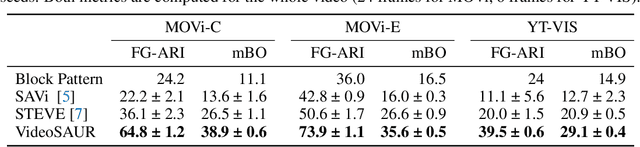
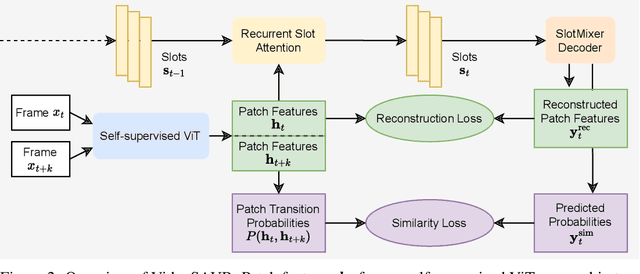

Unsupervised video-based object-centric learning is a promising avenue to learn structured representations from large, unlabeled video collections, but previous approaches have only managed to scale to real-world datasets in restricted domains. Recently, it was shown that the reconstruction of pre-trained self-supervised features leads to object-centric representations on unconstrained real-world image datasets. Building on this approach, we propose a novel way to use such pre-trained features in the form of a temporal feature similarity loss. This loss encodes temporal correlations between image patches and is a natural way to introduce a motion bias for object discovery. We demonstrate that this loss leads to state-of-the-art performance on the challenging synthetic MOVi datasets. When used in combination with the feature reconstruction loss, our model is the first object-centric video model that scales to unconstrained video datasets such as YouTube-VIS.
Medical Federated Model with Mixture of Personalized and Sharing Components
Jun 26, 2023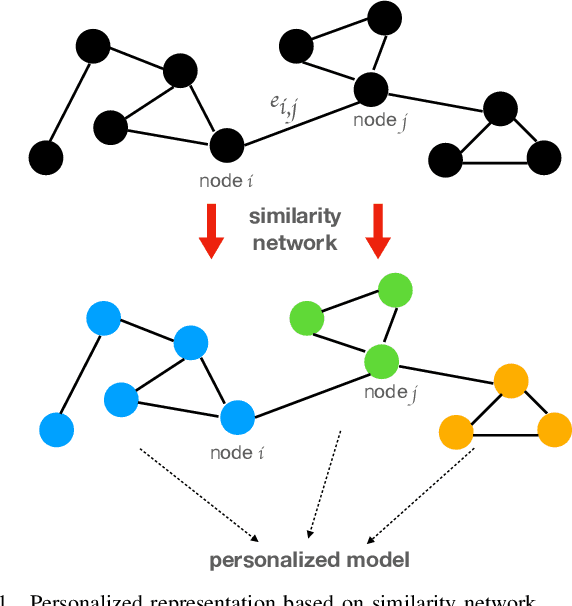
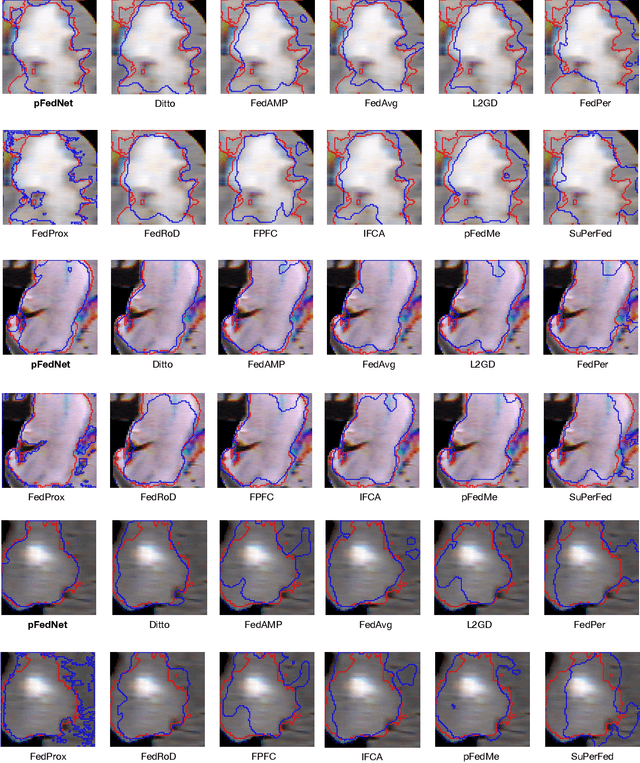
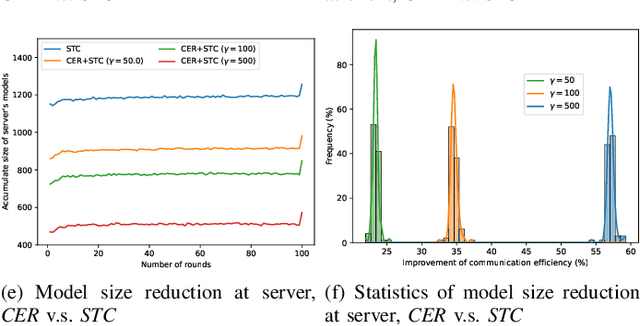

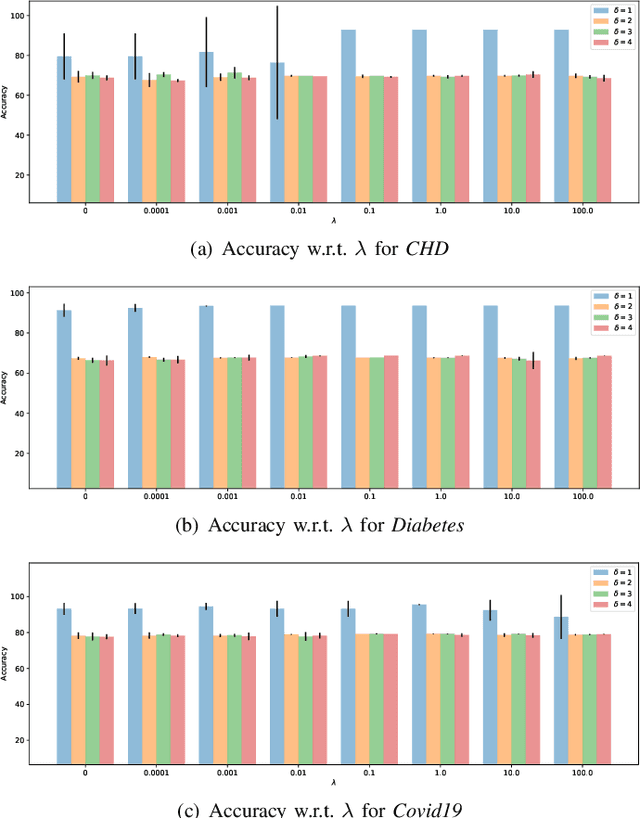
Although data-driven methods usually have noticeable performance on disease diagnosis and treatment, they are suspected of leakage of privacy due to collecting data for model training. Recently, federated learning provides a secure and trustable alternative to collaboratively train model without any exchange of medical data among multiple institutes. Therefore, it has draw much attention due to its natural merit on privacy protection. However, when heterogenous medical data exists between different hospitals, federated learning usually has to face with degradation of performance. In the paper, we propose a new personalized framework of federated learning to handle the problem. It successfully yields personalized models based on awareness of similarity between local data, and achieves better tradeoff between generalization and personalization than existing methods. After that, we further design a differentially sparse regularizer to improve communication efficiency during procedure of model training. Additionally, we propose an effective method to reduce the computational cost, which improves computation efficiency significantly. Furthermore, we collect 5 real medical datasets, including 2 public medical image datasets and 3 private multi-center clinical diagnosis datasets, and evaluate its performance by conducting nodule classification, tumor segmentation, and clinical risk prediction tasks. Comparing with 13 existing related methods, the proposed method successfully achieves the best model performance, and meanwhile up to 60% improvement of communication efficiency. Source code is public, and can be accessed at: https://github.com/ApplicationTechnologyOfMedicalBigData/pFedNet-code.
STU-Net: Scalable and Transferable Medical Image Segmentation Models Empowered by Large-Scale Supervised Pre-training
Apr 13, 2023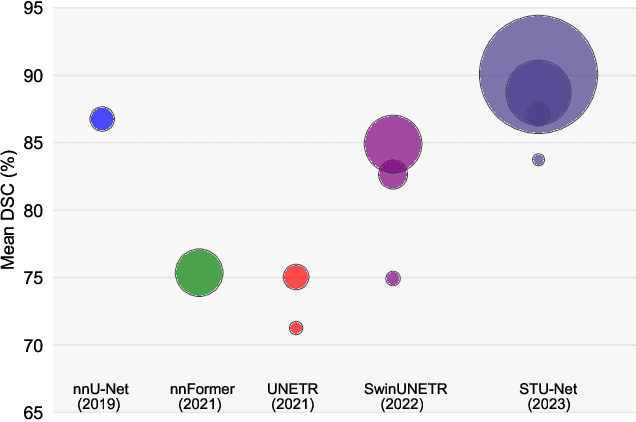
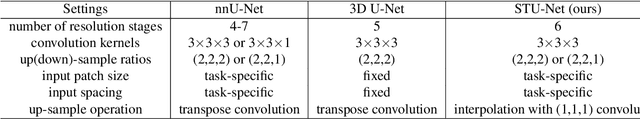

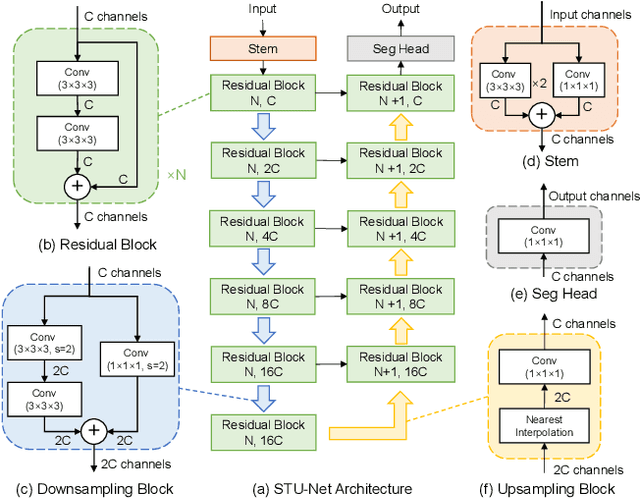

Large-scale models pre-trained on large-scale datasets have profoundly advanced the development of deep learning. However, the state-of-the-art models for medical image segmentation are still small-scale, with their parameters only in the tens of millions. Further scaling them up to higher orders of magnitude is rarely explored. An overarching goal of exploring large-scale models is to train them on large-scale medical segmentation datasets for better transfer capacities. In this work, we design a series of Scalable and Transferable U-Net (STU-Net) models, with parameter sizes ranging from 14 million to 1.4 billion. Notably, the 1.4B STU-Net is the largest medical image segmentation model to date. Our STU-Net is based on nnU-Net framework due to its popularity and impressive performance. We first refine the default convolutional blocks in nnU-Net to make them scalable. Then, we empirically evaluate different scaling combinations of network depth and width, discovering that it is optimal to scale model depth and width together. We train our scalable STU-Net models on a large-scale TotalSegmentator dataset and find that increasing model size brings a stronger performance gain. This observation reveals that a large model is promising in medical image segmentation. Furthermore, we evaluate the transferability of our model on 14 downstream datasets for direct inference and 3 datasets for further fine-tuning, covering various modalities and segmentation targets. We observe good performance of our pre-trained model in both direct inference and fine-tuning. The code and pre-trained models are available at https://github.com/Ziyan-Huang/STU-Net.
S-TLLR: STDP-inspired Temporal Local Learning Rule for Spiking Neural Networks
Jun 27, 2023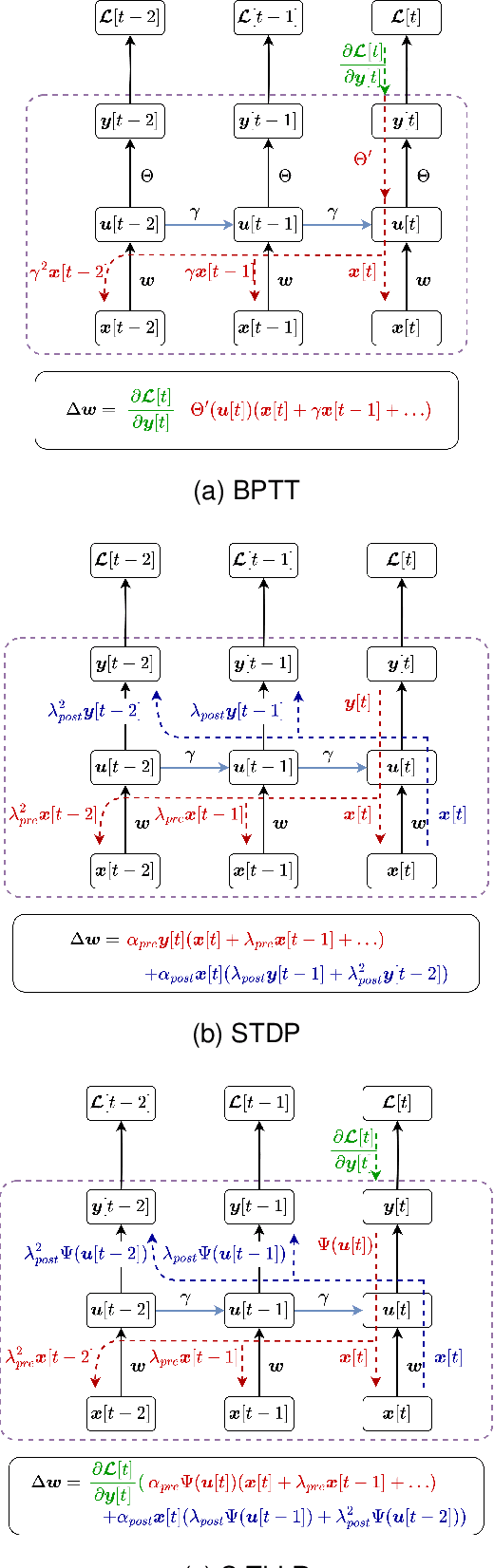
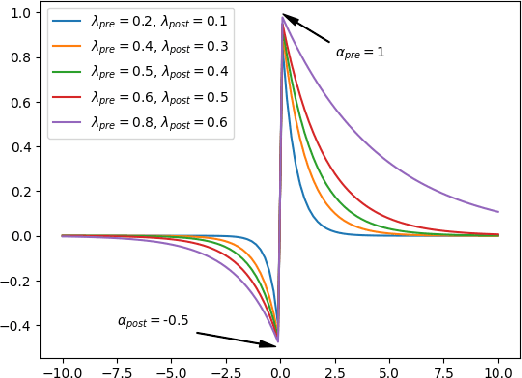
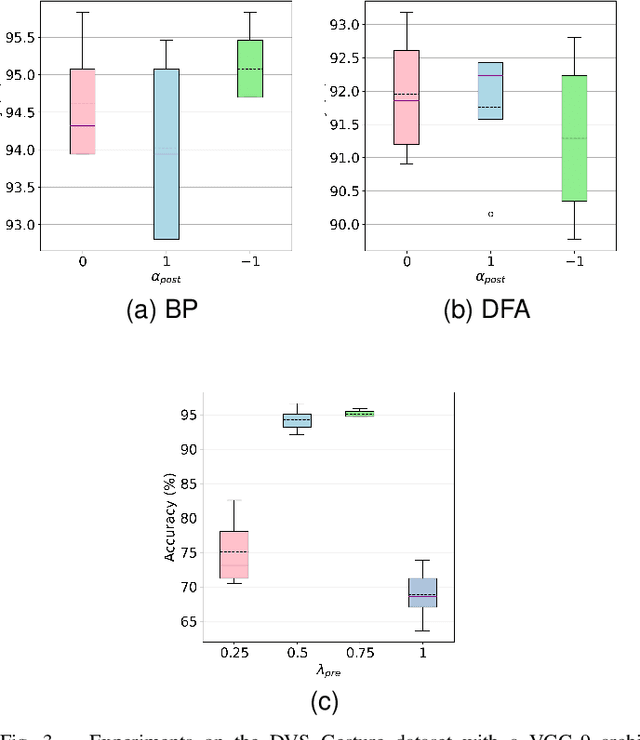
Spiking Neural Networks (SNNs) are biologically plausible models that have been identified as potentially apt for the deployment for energy-efficient intelligence at the edge, particularly for sequential learning tasks. However, training of SNNs poses a significant challenge due to the necessity for precise temporal and spatial credit assignment. Back-propagation through time (BPTT) algorithm, whilst being the most widely used method for addressing these issues, incurs a high computational cost due to its temporal dependency. Moreover, BPTT and its approximations solely utilize causal information derived from the spiking activity to compute the synaptic updates, thus neglecting non-causal relationships. In this work, we propose S-TLLR, a novel three-factor temporal local learning rule inspired by the Spike-Timing Dependent Plasticity (STDP) mechanism, aimed at training SNNs on event-based learning tasks. S-TLLR considers both causal and non-causal relationships between pre and post-synaptic activities, achieving performance comparable to BPTT and enhancing performance relative to methods using only causal information. Furthermore, S-TLLR has low memory and time complexity, which is independent of the number of time steps, rendering it suitable for online learning on low-power devices. To demonstrate the scalability of our proposed method, we have conducted extensive evaluations on event-based datasets spanning a wide range of applications, such as image and gesture recognition, audio classification, and optical flow estimation. In all the experiments, S-TLLR achieved high accuracy with a reduction in the number of computations between $1.1-10\times$.
Novel Hybrid-Learning Algorithms for Improved Millimeter-Wave Imaging Systems
Jun 27, 2023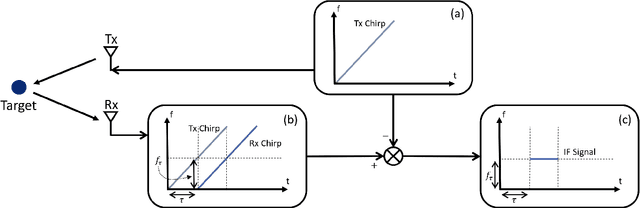
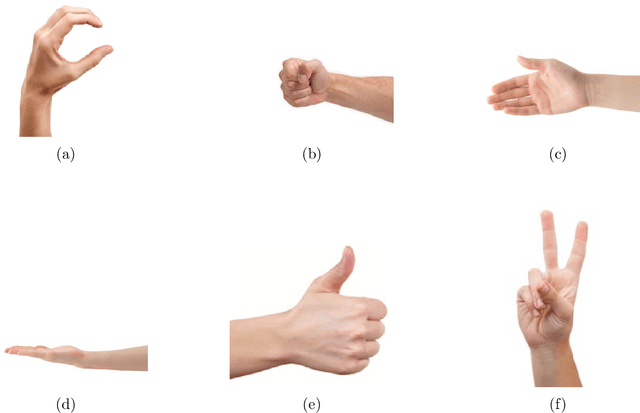
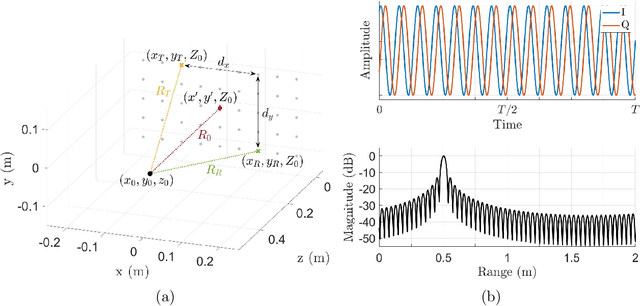

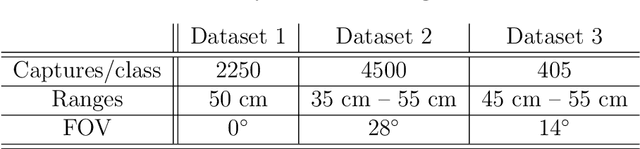
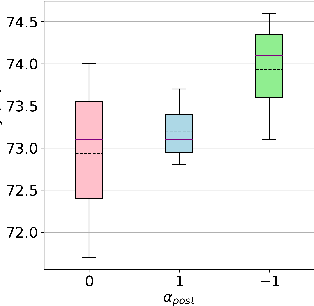
Increasing attention is being paid to millimeter-wave (mmWave), 30 GHz to 300 GHz, and terahertz (THz), 300 GHz to 10 THz, sensing applications including security sensing, industrial packaging, medical imaging, and non-destructive testing. Traditional methods for perception and imaging are challenged by novel data-driven algorithms that offer improved resolution, localization, and detection rates. Over the past decade, deep learning technology has garnered substantial popularity, particularly in perception and computer vision applications. Whereas conventional signal processing techniques are more easily generalized to various applications, hybrid approaches where signal processing and learning-based algorithms are interleaved pose a promising compromise between performance and generalizability. Furthermore, such hybrid algorithms improve model training by leveraging the known characteristics of radio frequency (RF) waveforms, thus yielding more efficiently trained deep learning algorithms and offering higher performance than conventional methods. This dissertation introduces novel hybrid-learning algorithms for improved mmWave imaging systems applicable to a host of problems in perception and sensing. Various problem spaces are explored, including static and dynamic gesture classification; precise hand localization for human computer interaction; high-resolution near-field mmWave imaging using forward synthetic aperture radar (SAR); SAR under irregular scanning geometries; mmWave image super-resolution using deep neural network (DNN) and Vision Transformer (ViT) architectures; and data-level multiband radar fusion using a novel hybrid-learning architecture. Furthermore, we introduce several novel approaches for deep learning model training and dataset synthesis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge