"Image": models, code, and papers
TD-GEM: Text-Driven Garment Editing Mapper
May 29, 2023
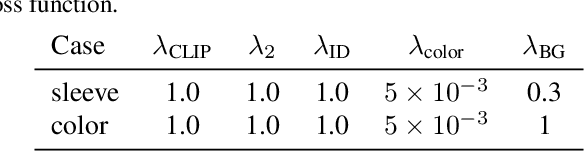

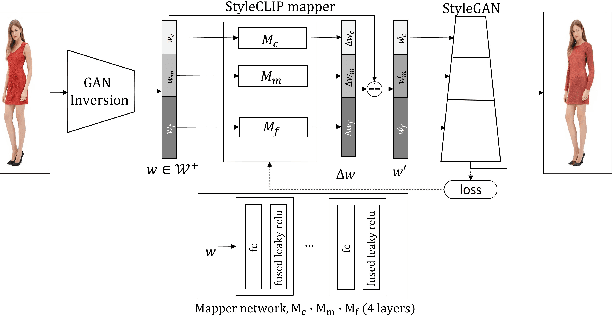
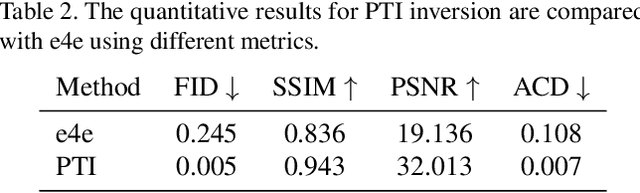
Language-based fashion image editing allows users to try out variations of desired garments through provided text prompts. Inspired by research on manipulating latent representations in StyleCLIP and HairCLIP, we focus on these latent spaces for editing fashion items of full-body human datasets. Currently, there is a gap in handling fashion image editing due to the complexity of garment shapes and textures and the diversity of human poses. In this paper, we propose an editing optimizer scheme method called Text-Driven Garment Editing Mapper (TD-GEM), aiming to edit fashion items in a disentangled way. To this end, we initially obtain a latent representation of an image through generative adversarial network inversions such as Encoder for Editing (e4e) or Pivotal Tuning Inversion (PTI) for more accurate results. An optimization-based Contrasive Language-Image Pre-training (CLIP) is then utilized to guide the latent representation of a fashion image in the direction of a target attribute expressed in terms of a text prompt. Our TD-GEM manipulates the image accurately according to the target attribute, while other parts of the image are kept untouched. In the experiments, we evaluate TD-GEM on two different attributes (i.e., "color" and "sleeve length"), which effectively generates realistic images compared to the recent manipulation schemes.
Adversarial Attack and Defense for Medical Image Analysis: Methods and Applications
Mar 24, 2023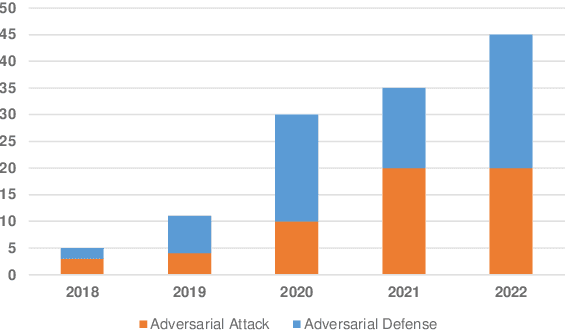
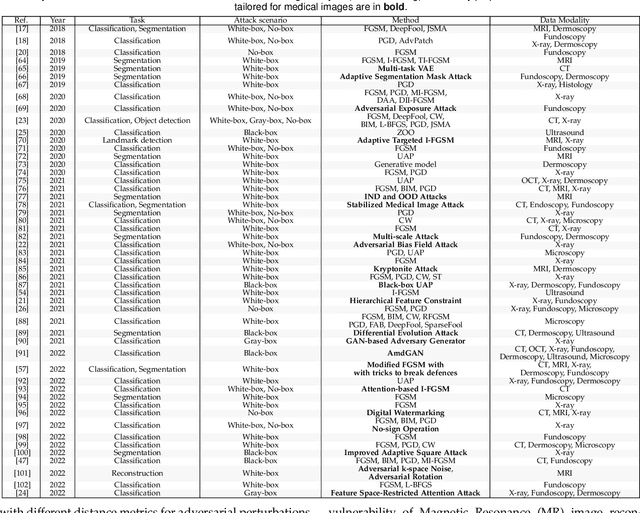
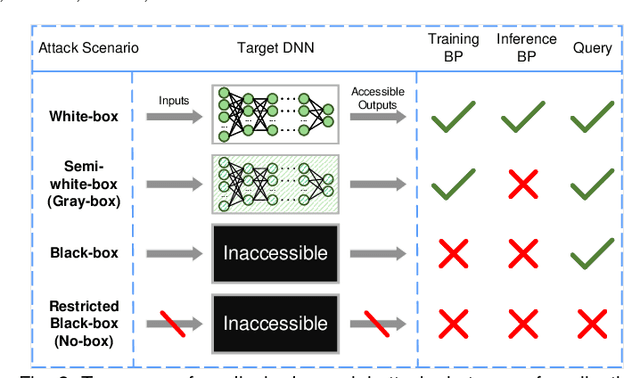
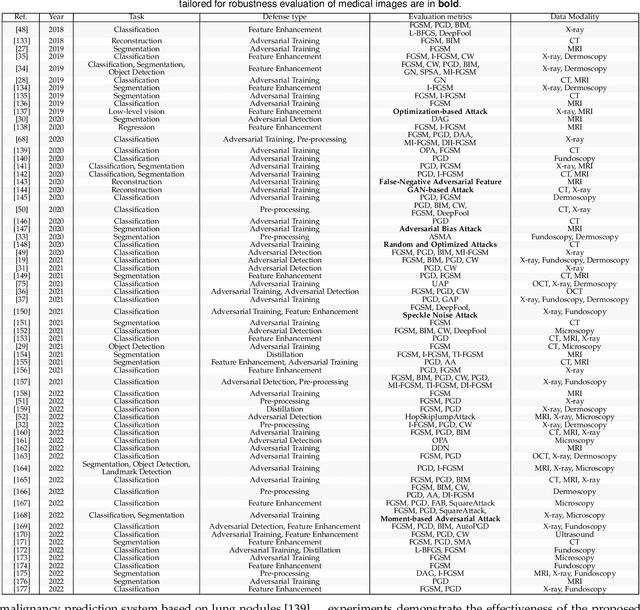
Deep learning techniques have achieved superior performance in computer-aided medical image analysis, yet they are still vulnerable to imperceptible adversarial attacks, resulting in potential misdiagnosis in clinical practice. Oppositely, recent years have also witnessed remarkable progress in defense against these tailored adversarial examples in deep medical diagnosis systems. In this exposition, we present a comprehensive survey on recent advances in adversarial attack and defense for medical image analysis with a novel taxonomy in terms of the application scenario. We also provide a unified theoretical framework for different types of adversarial attack and defense methods for medical image analysis. For a fair comparison, we establish a new benchmark for adversarially robust medical diagnosis models obtained by adversarial training under various scenarios. To the best of our knowledge, this is the first survey paper that provides a thorough evaluation of adversarially robust medical diagnosis models. By analyzing qualitative and quantitative results, we conclude this survey with a detailed discussion of current challenges for adversarial attack and defense in medical image analysis systems to shed light on future research directions.
Probabilistic Risk Assessment of an Obstacle Detection System for GoA 4 Freight Trains
Jun 26, 2023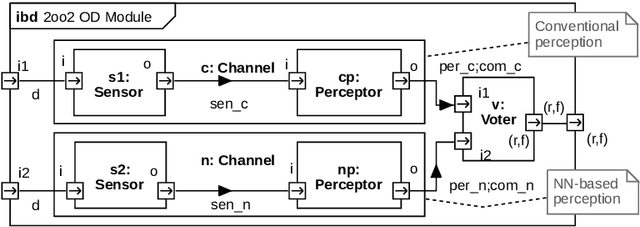

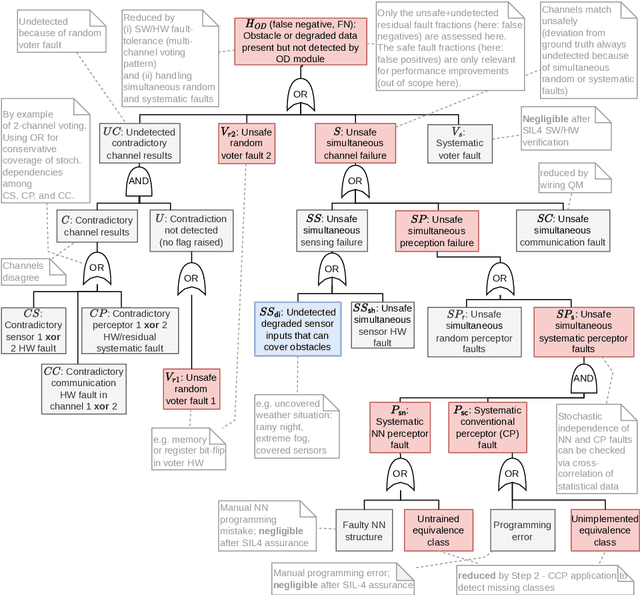
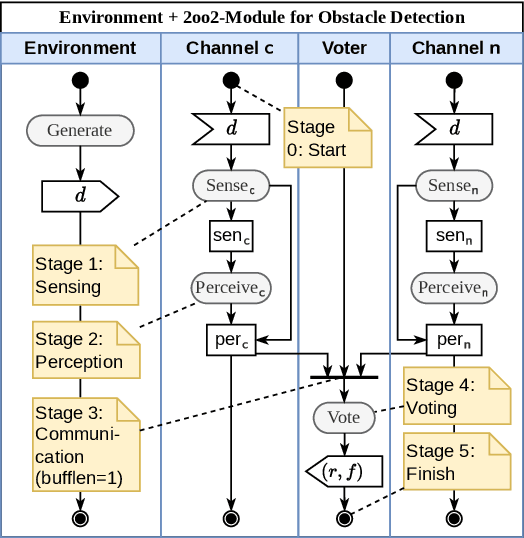
In this paper, a quantitative risk assessment approach is discussed for the design of an obstacle detection function for low-speed freight trains with grade of automation (GoA)~4. In this 5-step approach, starting with single detection channels and ending with a three-out-of-three (3oo3) model constructed of three independent dual-channel modules and a voter, a probabilistic assessment is exemplified, using a combination of statistical methods and parametric stochastic model checking. It is illustrated that, under certain not unreasonable assumptions, the resulting hazard rate becomes acceptable for specific application settings. The statistical approach for assessing the residual risk of misclassifications in convolutional neural networks and conventional image processing software suggests that high confidence can be placed into the safety-critical obstacle detection function, even though its implementation involves realistic machine learning uncertainties.
Toward Fairness Through Fair Multi-Exit Framework for Dermatological Disease Diagnosis
Jun 26, 2023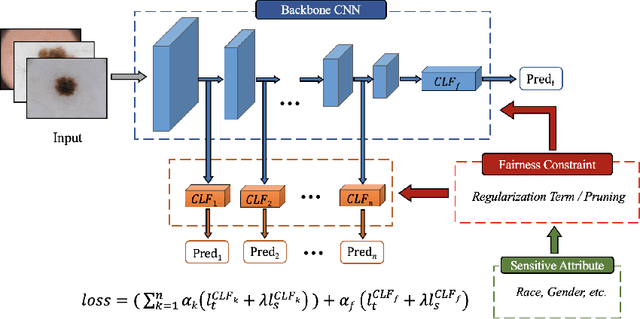
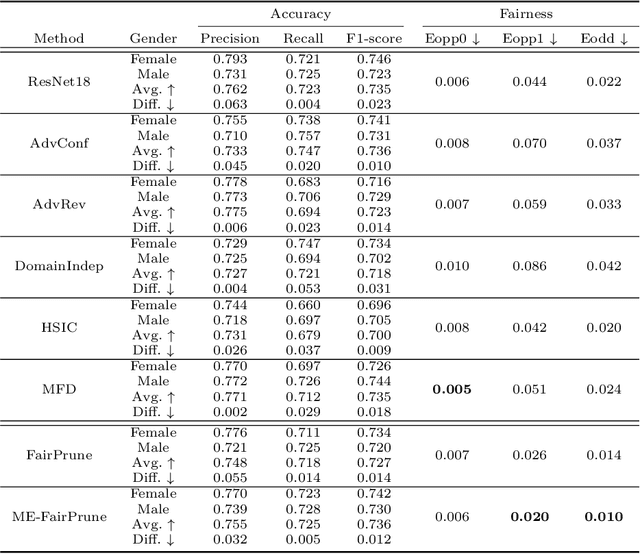
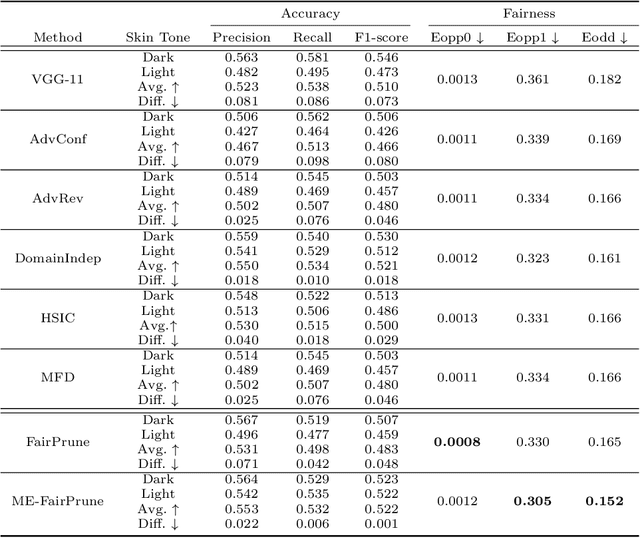
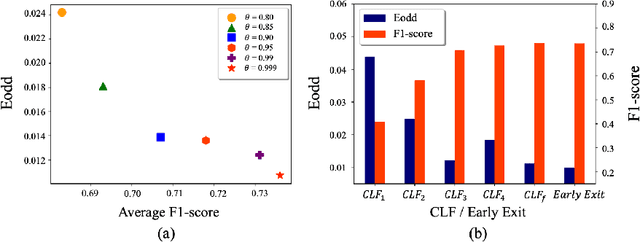
Fairness has become increasingly pivotal in medical image recognition. However, without mitigating bias, deploying unfair medical AI systems could harm the interests of underprivileged populations. In this paper, we observe that while features extracted from the deeper layers of neural networks generally offer higher accuracy, fairness conditions deteriorate as we extract features from deeper layers. This phenomenon motivates us to extend the concept of multi-exit frameworks. Unlike existing works mainly focusing on accuracy, our multi-exit framework is fairness-oriented; the internal classifiers are trained to be more accurate and fairer, with high extensibility to apply to most existing fairness-aware frameworks. During inference, any instance with high confidence from an internal classifier is allowed to exit early. Experimental results show that the proposed framework can improve the fairness condition over the state-of-the-art in two dermatological disease datasets.
Visually grounded few-shot word acquisition with fewer shots
May 25, 2023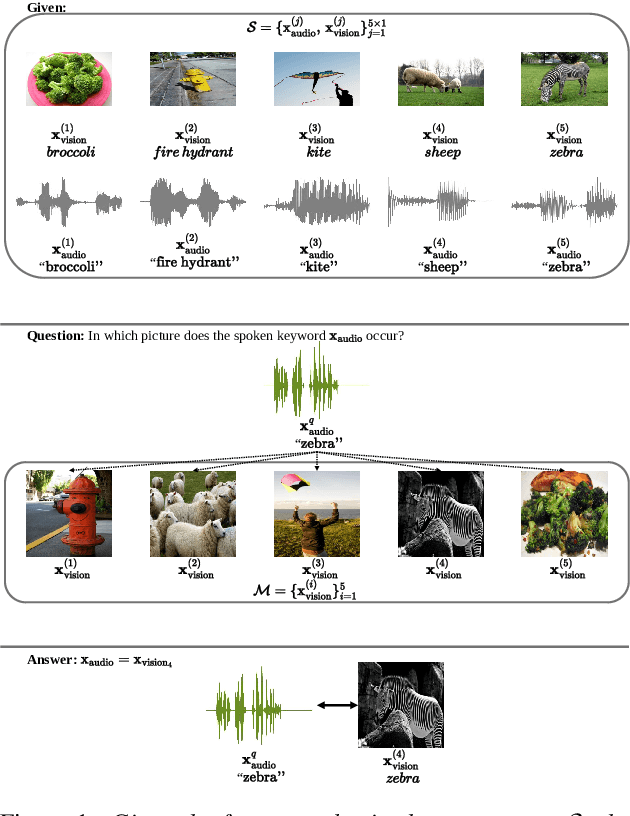

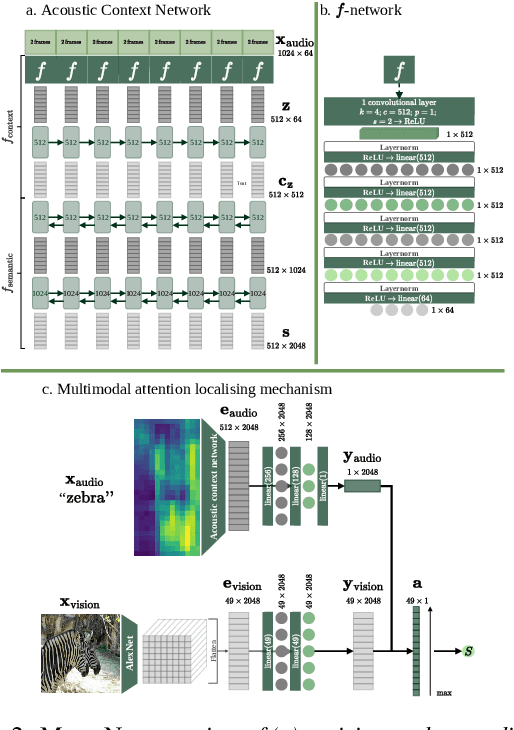
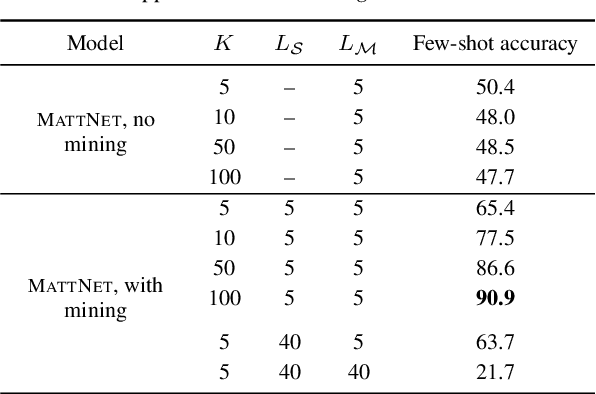
We propose a visually grounded speech model that acquires new words and their visual depictions from just a few word-image example pairs. Given a set of test images and a spoken query, we ask the model which image depicts the query word. Previous work has simplified this problem by either using an artificial setting with digit word-image pairs or by using a large number of examples per class. We propose an approach that can work on natural word-image pairs but with less examples, i.e. fewer shots. Our approach involves using the given word-image example pairs to mine new unsupervised word-image training pairs from large collections of unlabelled speech and images. Additionally, we use a word-to-image attention mechanism to determine word-image similarity. With this new model, we achieve better performance with fewer shots than any existing approach.
Factor Decomposed Generative Adversarial Networks for Text-to-Image Synthesis
Mar 24, 2023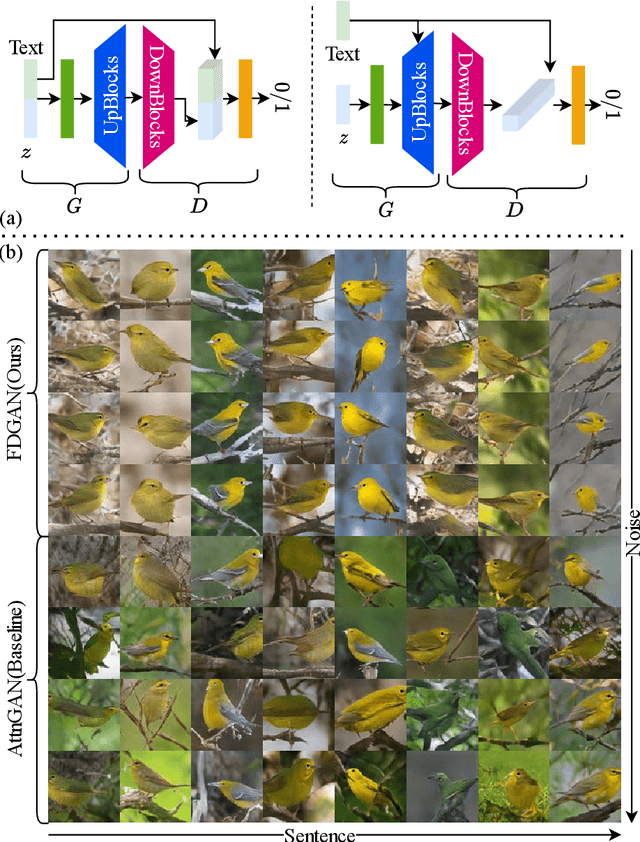
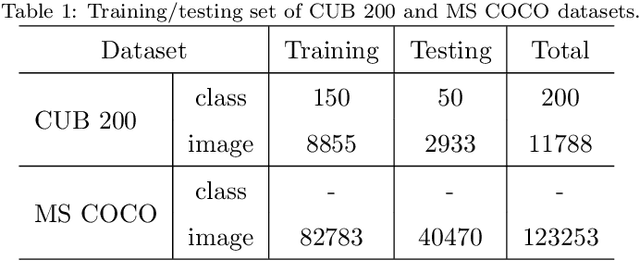
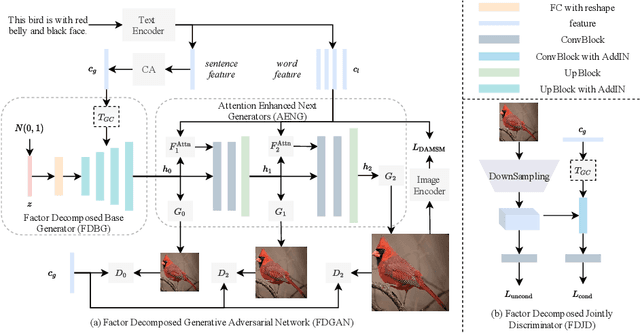
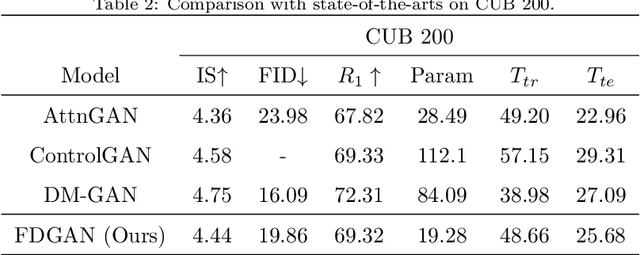
Prior works about text-to-image synthesis typically concatenated the sentence embedding with the noise vector, while the sentence embedding and the noise vector are two different factors, which control the different aspects of the generation. Simply concatenating them will entangle the latent factors and encumber the generative model. In this paper, we attempt to decompose these two factors and propose Factor Decomposed Generative Adversarial Networks~(FDGAN). To achieve this, we firstly generate images from the noise vector and then apply the sentence embedding in the normalization layer for both generator and discriminators. We also design an additive norm layer to align and fuse the text-image features. The experimental results show that decomposing the noise and the sentence embedding can disentangle latent factors in text-to-image synthesis, and make the generative model more efficient. Compared with the baseline, FDGAN can achieve better performance, while fewer parameters are used.
Tame a Wild Camera: In-the-Wild Monocular Camera Calibration
Jun 19, 2023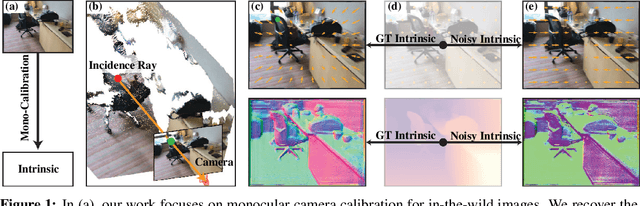

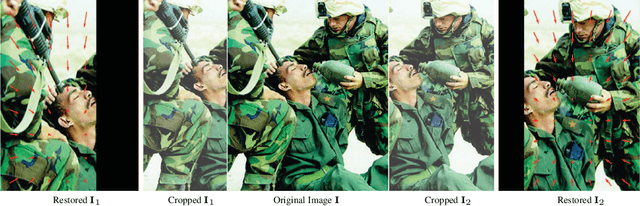
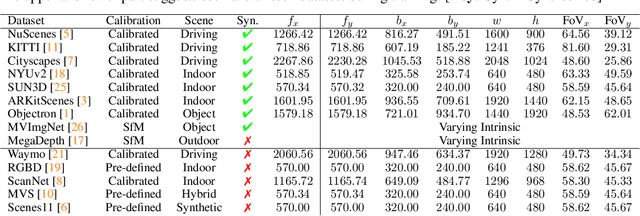
3D sensing for monocular in-the-wild images, e.g., depth estimation and 3D object detection, has become increasingly important. However, the unknown intrinsic parameter hinders their development and deployment. Previous methods for the monocular camera calibration rely on specific 3D objects or strong geometry prior, such as using a checkerboard or imposing a Manhattan World assumption. This work solves the problem from the other perspective by exploiting the monocular 3D prior. Our method is assumption-free and calibrates the complete $4$ Degree-of-Freedom (DoF) intrinsic parameters. First, we demonstrate intrinsic is solved from two well-studied monocular priors, i.e., monocular depthmap, and surface normal map. However, this solution imposes a low-bias and low-variance requirement for depth estimation. Alternatively, we introduce a novel monocular 3D prior, the incidence field, defined as the incidence rays between points in 3D space and pixels in the 2D imaging plane. The incidence field is a pixel-wise parametrization of the intrinsic invariant to image cropping and resizing. With the estimated incidence field, a robust RANSAC algorithm recovers intrinsic. We demonstrate the effectiveness of our method by showing superior performance on synthetic and zero-shot testing datasets. Beyond calibration, we demonstrate downstream applications in image manipulation detection & restoration, uncalibrated two-view pose estimation, and 3D sensing. Codes, models, and data will be held in https://github.com/ShngJZ/WildCamera.
An Accelerated Pipeline for Multi-label Renal Pathology Image Segmentation at the Whole Slide Image Level
May 23, 2023Deep-learning techniques have been used widely to alleviate the labour-intensive and time-consuming manual annotation required for pixel-level tissue characterization. Our previous study introduced an efficient single dynamic network - Omni-Seg - that achieved multi-class multi-scale pathological segmentation with less computational complexity. However, the patch-wise segmentation paradigm still applies to Omni-Seg, and the pipeline is time-consuming when providing segmentation for Whole Slide Images (WSIs). In this paper, we propose an enhanced version of the Omni-Seg pipeline in order to reduce the repetitive computing processes and utilize a GPU to accelerate the model's prediction for both better model performance and faster speed. Our proposed method's innovative contribution is two-fold: (1) a Docker is released for an end-to-end slide-wise multi-tissue segmentation for WSIs; and (2) the pipeline is deployed on a GPU to accelerate the prediction, achieving better segmentation quality in less time. The proposed accelerated implementation reduced the average processing time (at the testing stage) on a standard needle biopsy WSI from 2.3 hours to 22 minutes, using 35 WSIs from the Kidney Tissue Atlas (KPMP) Datasets. The source code and the Docker have been made publicly available at https://github.com/ddrrnn123/Omni-Seg.
Autoencoders for Real-Time SUEP Detection
Jun 23, 2023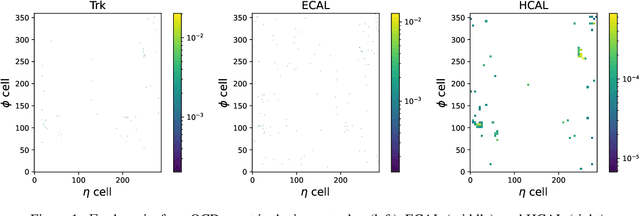
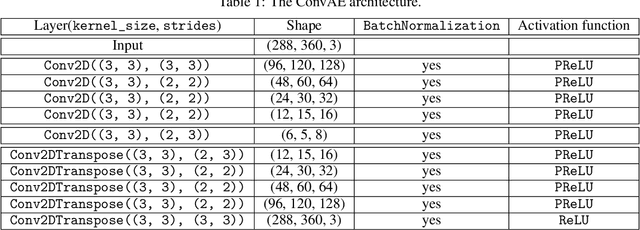
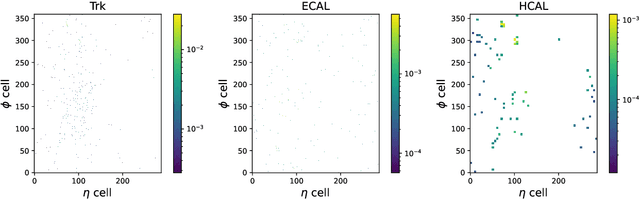
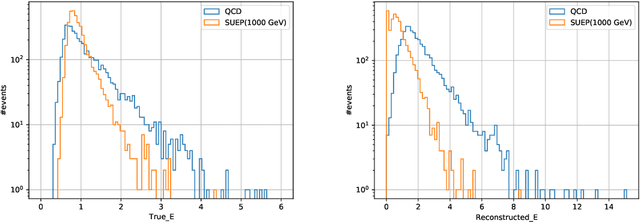
Confining dark sectors with pseudo-conformal dynamics can produce Soft Unclustered Energy Patterns, or SUEPs, at the Large Hadron Collider: the production of dark quarks in proton-proton collisions leading to a dark shower and the high-multiplicity production of dark hadrons. The final experimental signature is spherically-symmetric energy deposits by an anomalously large number of soft Standard Model particles with a transverse energy of a few hundred MeV. The dominant background for the SUEP search, if it gets produced via gluon-gluon fusion, is multi-jet QCD events. We have developed a deep learning-based Anomaly Detection technique to reject QCD jets and identify any anomalous signature, including SUEP, in real-time in the High-Level Trigger system of the Compact Muon Solenoid experiment at the Large Hadron Collider. A deep convolutional neural autoencoder network has been trained using QCD events by taking transverse energy deposits in the inner tracker, electromagnetic calorimeter, and hadron calorimeter sub-detectors as 3-channel image data. To tackle the biggest challenge of the task, due to the sparse nature of the data: only ~0.5% of the total ~300 k image pixels have non-zero values, a non-standard loss function, the inverse of the so-called Dice Loss, has been exploited. The trained autoencoder with learned spatial features of QCD jets can detect 40% of the SUEP events, with a QCD event mistagging rate as low as 2%. The model inference time has been measured using the Intel CoreTM i5-9600KF processor and found to be ~20 ms, which perfectly satisfies the High-Level Trigger system's latency of O(100) ms. Given the virtue of the unsupervised learning of the autoencoders, the trained model can be applied to any new physics model that predicts an experimental signature anomalous to QCD jets.
Self-Supervised Pre-Training for Deep Image Prior-Based Robust PET Image Denoising
Feb 27, 2023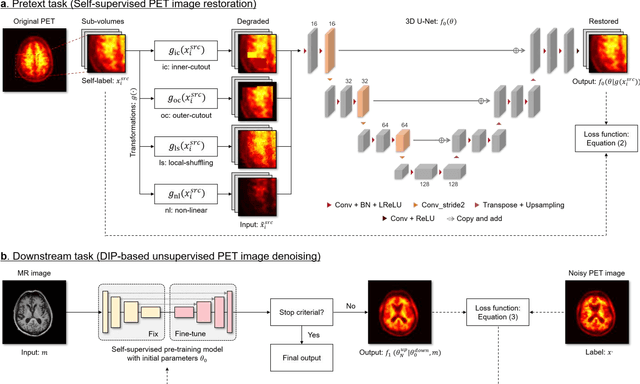
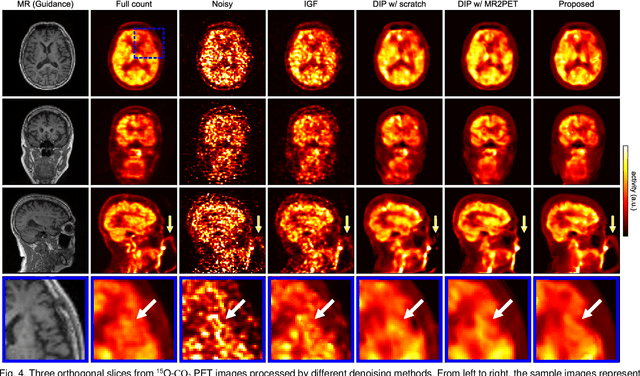
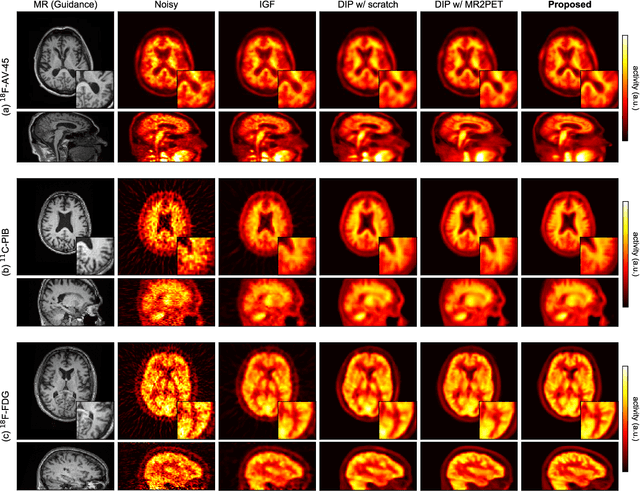
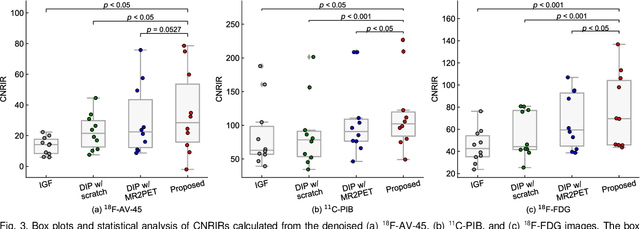

Deep image prior (DIP) has been successfully applied to positron emission tomography (PET) image restoration, enabling represent implicit prior using only convolutional neural network architecture without training dataset, whereas the general supervised approach requires massive low- and high-quality PET image pairs. To answer the increased need for PET imaging with DIP, it is indispensable to improve the performance of the underlying DIP itself. Here, we propose a self-supervised pre-training model to improve the DIP-based PET image denoising performance. Our proposed pre-training model acquires transferable and generalizable visual representations from only unlabeled PET images by restoring various degraded PET images in a self-supervised approach. We evaluated the proposed method using clinical brain PET data with various radioactive tracers ($^{18}$F-florbetapir, $^{11}$C-Pittsburgh compound-B, $^{18}$F-fluoro-2-deoxy-D-glucose, and $^{15}$O-CO$_{2}$) acquired from different PET scanners. The proposed method using the self-supervised pre-training model achieved robust and state-of-the-art denoising performance while retaining spatial details and quantification accuracy compared to other unsupervised methods and pre-training model. These results highlight the potential that the proposed method is particularly effective against rare diseases and probes and helps reduce the scan time or the radiotracer dose without affecting the patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge