"Image": models, code, and papers
GaussianImage: 1000 FPS Image Representation and Compression by 2D Gaussian Splatting
Mar 14, 2024
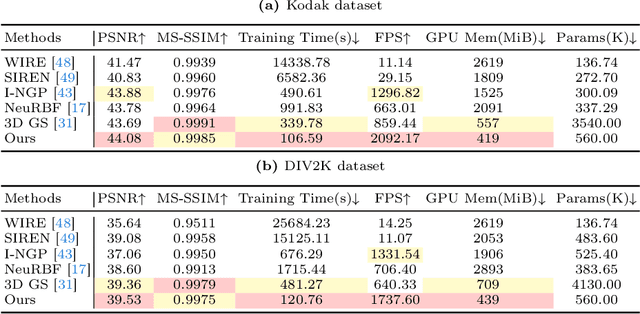

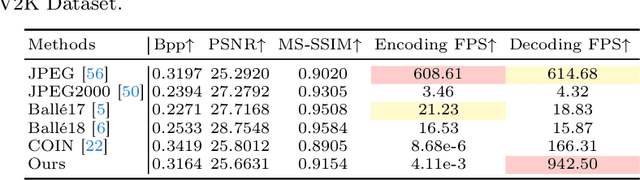
Implicit neural representations (INRs) recently achieved great success in image representation and compression, offering high visual quality and fast rendering speeds with 10-1000 FPS, assuming sufficient GPU resources are available. However, this requirement often hinders their use on low-end devices with limited memory. In response, we propose a groundbreaking paradigm of image representation and compression by 2D Gaussian Splatting, named GaussianImage. We first introduce 2D Gaussian to represent the image, where each Gaussian has 8 parameters including position, covariance and color. Subsequently, we unveil a novel rendering algorithm based on accumulated summation. Remarkably, our method with a minimum of 3$\times$ lower GPU memory usage and 5$\times$ faster fitting time not only rivals INRs (e.g., WIRE, I-NGP) in representation performance, but also delivers a faster rendering speed of 1500-2000 FPS regardless of parameter size. Furthermore, we integrate existing vector quantization technique to build an image codec. Experimental results demonstrate that our codec attains rate-distortion performance comparable to compression-based INRs such as COIN and COIN++, while facilitating decoding speeds of approximately 1000 FPS. Additionally, preliminary proof of concept shows that our codec surpasses COIN and COIN++ in performance when using partial bits-back coding.
An Optimization Framework to Enforce Multi-View Consistency for Texturing 3D Meshes Using Pre-Trained Text-to-Image Models
Mar 22, 2024A fundamental problem in the texturing of 3D meshes using pre-trained text-to-image models is to ensure multi-view consistency. State-of-the-art approaches typically use diffusion models to aggregate multi-view inputs, where common issues are the blurriness caused by the averaging operation in the aggregation step or inconsistencies in local features. This paper introduces an optimization framework that proceeds in four stages to achieve multi-view consistency. Specifically, the first stage generates an over-complete set of 2D textures from a predefined set of viewpoints using an MV-consistent diffusion process. The second stage selects a subset of views that are mutually consistent while covering the underlying 3D model. We show how to achieve this goal by solving semi-definite programs. The third stage performs non-rigid alignment to align the selected views across overlapping regions. The fourth stage solves an MRF problem to associate each mesh face with a selected view. In particular, the third and fourth stages are iterated, with the cuts obtained in the fourth stage encouraging non-rigid alignment in the third stage to focus on regions close to the cuts. Experimental results show that our approach significantly outperforms baseline approaches both qualitatively and quantitatively.
Automated Report Generation for Lung Cytological Images Using a CNN Vision Classifier and Multiple-Transformer Text Decoders: Preliminary Study
Mar 26, 2024Cytology plays a crucial role in lung cancer diagnosis. Pulmonary cytology involves cell morphological characterization in the specimen and reporting the corresponding findings, which are extremely burdensome tasks. In this study, we propose a report-generation technique for lung cytology images. In total, 71 benign and 135 malignant pulmonary cytology specimens were collected. Patch images were extracted from the captured specimen images, and the findings were assigned to each image as a dataset for report generation. The proposed method consists of a vision model and a text decoder. In the former, a convolutional neural network (CNN) is used to classify a given image as benign or malignant, and the features related to the image are extracted from the intermediate layer. Independent text decoders for benign and malignant cells are prepared for text generation, and the text decoder switches according to the CNN classification results. The text decoder is configured using a Transformer that uses the features obtained from the CNN for report generation. Based on the evaluation results, the sensitivity and specificity were 100% and 96.4%, respectively, for automated benign and malignant case classification, and the saliency map indicated characteristic benign and malignant areas. The grammar and style of the generated texts were confirmed as correct and in better agreement with gold standard compared to existing LLM-based image-captioning methods and single-text-decoder ablation model. These results indicate that the proposed method is useful for pulmonary cytology classification and reporting.
Eta Inversion: Designing an Optimal Eta Function for Diffusion-based Real Image Editing
Mar 14, 2024
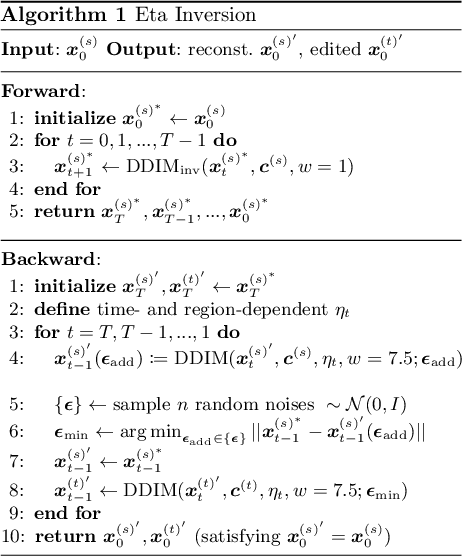
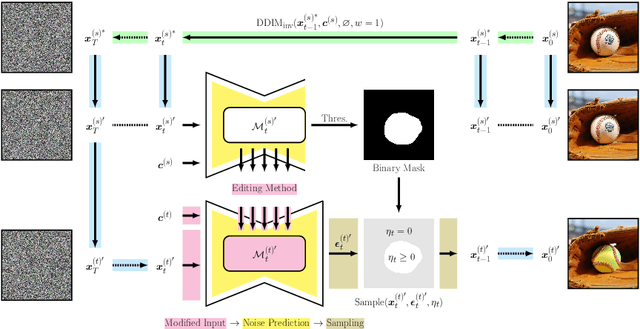
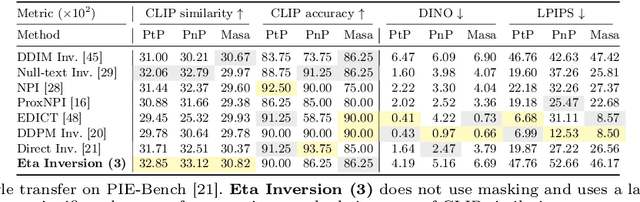

Diffusion models have achieved remarkable success in the domain of text-guided image generation and, more recently, in text-guided image editing. A commonly adopted strategy for editing real images involves inverting the diffusion process to obtain a noisy representation of the original image, which is then denoised to achieve the desired edits. However, current methods for diffusion inversion often struggle to produce edits that are both faithful to the specified text prompt and closely resemble the source image. To overcome these limitations, we introduce a novel and adaptable diffusion inversion technique for real image editing, which is grounded in a theoretical analysis of the role of $\eta$ in the DDIM sampling equation for enhanced editability. By designing a universal diffusion inversion method with a time- and region-dependent $\eta$ function, we enable flexible control over the editing extent. Through a comprehensive series of quantitative and qualitative assessments, involving a comparison with a broad array of recent methods, we demonstrate the superiority of our approach. Our method not only sets a new benchmark in the field but also significantly outperforms existing strategies. Our code is available at https://github.com/furiosa-ai/eta-inversion
Unleashing the Potential of SAM for Medical Adaptation via Hierarchical Decoding
Mar 27, 2024The Segment Anything Model (SAM) has garnered significant attention for its versatile segmentation abilities and intuitive prompt-based interface. However, its application in medical imaging presents challenges, requiring either substantial training costs and extensive medical datasets for full model fine-tuning or high-quality prompts for optimal performance. This paper introduces H-SAM: a prompt-free adaptation of SAM tailored for efficient fine-tuning of medical images via a two-stage hierarchical decoding procedure. In the initial stage, H-SAM employs SAM's original decoder to generate a prior probabilistic mask, guiding a more intricate decoding process in the second stage. Specifically, we propose two key designs: 1) A class-balanced, mask-guided self-attention mechanism addressing the unbalanced label distribution, enhancing image embedding; 2) A learnable mask cross-attention mechanism spatially modulating the interplay among different image regions based on the prior mask. Moreover, the inclusion of a hierarchical pixel decoder in H-SAM enhances its proficiency in capturing fine-grained and localized details. This approach enables SAM to effectively integrate learned medical priors, facilitating enhanced adaptation for medical image segmentation with limited samples. Our H-SAM demonstrates a 4.78% improvement in average Dice compared to existing prompt-free SAM variants for multi-organ segmentation using only 10% of 2D slices. Notably, without using any unlabeled data, H-SAM even outperforms state-of-the-art semi-supervised models relying on extensive unlabeled training data across various medical datasets. Our code is available at https://github.com/Cccccczh404/H-SAM.
LightIt: Illumination Modeling and Control for Diffusion Models
Mar 25, 2024
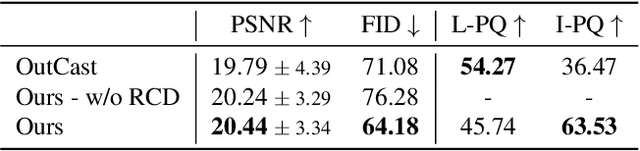
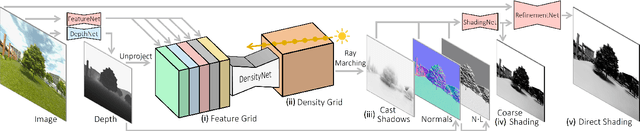

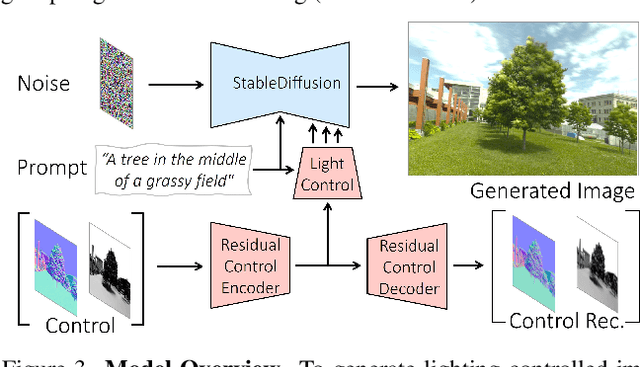
We introduce LightIt, a method for explicit illumination control for image generation. Recent generative methods lack lighting control, which is crucial to numerous artistic aspects of image generation such as setting the overall mood or cinematic appearance. To overcome these limitations, we propose to condition the generation on shading and normal maps. We model the lighting with single bounce shading, which includes cast shadows. We first train a shading estimation module to generate a dataset of real-world images and shading pairs. Then, we train a control network using the estimated shading and normals as input. Our method demonstrates high-quality image generation and lighting control in numerous scenes. Additionally, we use our generated dataset to train an identity-preserving relighting model, conditioned on an image and a target shading. Our method is the first that enables the generation of images with controllable, consistent lighting and performs on par with specialized relighting state-of-the-art methods.
The Solution for the ICCV 2023 1st Scientific Figure Captioning Challenge
Mar 26, 2024In this paper, we propose a solution for improving the quality of captions generated for figures in papers. We adopt the approach of summarizing the textual content in the paper to generate image captions. Throughout our study, we encounter discrepancies in the OCR information provided in the official dataset. To rectify this, we employ the PaddleOCR toolkit to extract OCR information from all images. Moreover, we observe that certain textual content in the official paper pertains to images that are not relevant for captioning, thereby introducing noise during caption generation. To mitigate this issue, we leverage LLaMA to extract image-specific information by querying the textual content based on image mentions, effectively filtering out extraneous information. Additionally, we recognize a discrepancy between the primary use of maximum likelihood estimation during text generation and the evaluation metrics such as ROUGE employed to assess the quality of generated captions. To bridge this gap, we integrate the BRIO model framework, enabling a more coherent alignment between the generation and evaluation processes. Our approach ranked first in the final test with a score of 4.49.
Segment Any Medical Model Extended
Mar 26, 2024The Segment Anything Model (SAM) has drawn significant attention from researchers who work on medical image segmentation because of its generalizability. However, researchers have found that SAM may have limited performance on medical images compared to state-of-the-art non-foundation models. Regardless, the community sees potential in extending, fine-tuning, modifying, and evaluating SAM for analysis of medical imaging. An increasing number of works have been published focusing on the mentioned four directions, where variants of SAM are proposed. To this end, a unified platform helps push the boundary of the foundation model for medical images, facilitating the use, modification, and validation of SAM and its variants in medical image segmentation. In this work, we introduce SAMM Extended (SAMME), a platform that integrates new SAM variant models, adopts faster communication protocols, accommodates new interactive modes, and allows for fine-tuning of subcomponents of the models. These features can expand the potential of foundation models like SAM, and the results can be translated to applications such as image-guided therapy, mixed reality interaction, robotic navigation, and data augmentation.
MeDSLIP: Medical Dual-Stream Language-Image Pre-training for Fine-grained Alignment
Mar 15, 2024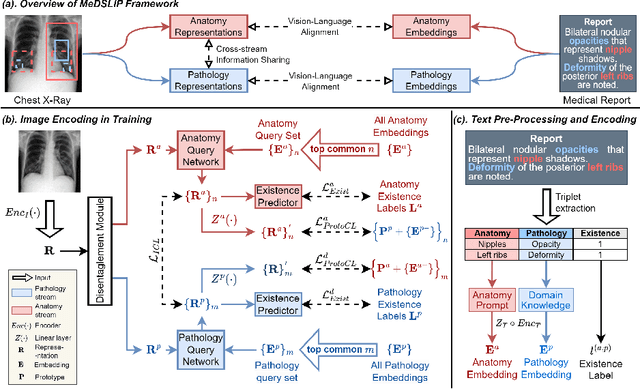
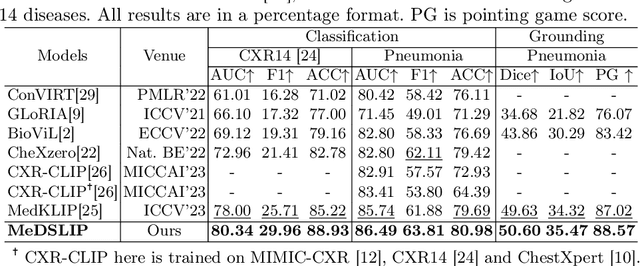
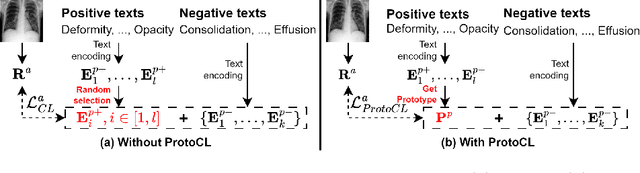
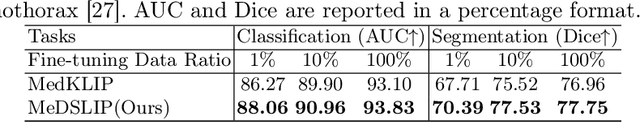
Vision-language pre-training (VLP) models have shown significant advancements in the medical domain. Yet, most VLP models align raw reports to images at a very coarse level, without modeling fine-grained relationships between anatomical and pathological concepts outlined in reports and the corresponding semantic counterparts in images. To address this problem, we propose a Medical Dual-Stream Language-Image Pre-training (MeDSLIP) framework. Specifically, MeDSLIP establishes vision-language fine-grained alignments via disentangling visual and textual representations into anatomy-relevant and pathology-relevant streams. Moreover, a novel vision-language Prototypical Contr-astive Learning (ProtoCL) method is adopted in MeDSLIP to enhance the alignment within the anatomical and pathological streams. MeDSLIP further employs cross-stream Intra-image Contrastive Learning (ICL) to ensure the consistent coexistence of paired anatomical and pathological concepts within the same image. Such a cross-stream regularization encourages the model to exploit the synchrony between two streams for a more comprehensive representation learning. MeDSLIP is evaluated under zero-shot and supervised fine-tuning settings on three public datasets: NIH CXR14, RSNA Pneumonia, and SIIM-ACR Pneumothorax. Under these settings, MeDSLIP outperforms six leading CNN-based models on classification, grounding, and segmentation tasks.
OCR is All you need: Importing Multi-Modality into Image-based Defect Detection System
Mar 18, 2024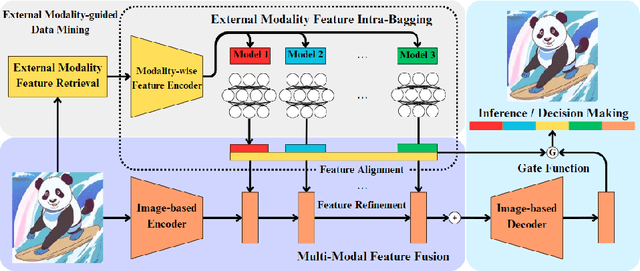
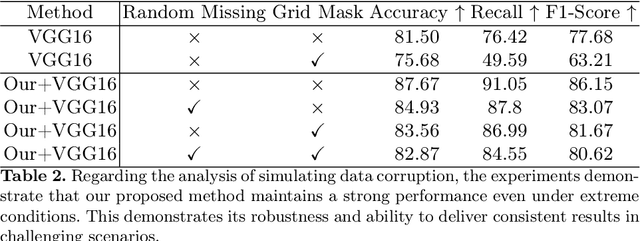
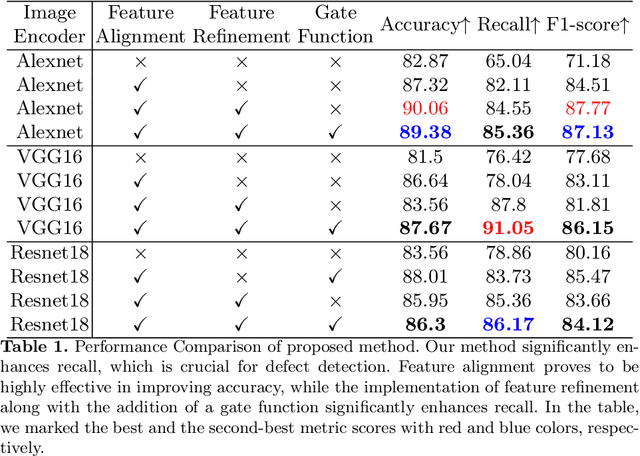
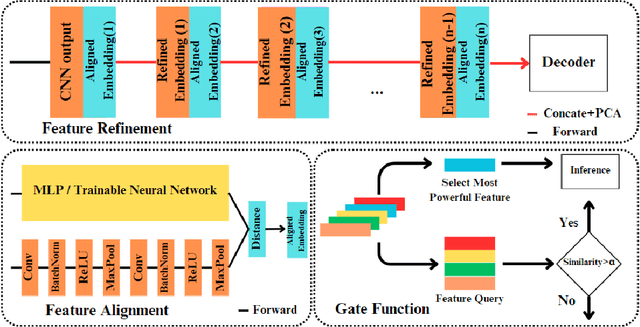

Automatic optical inspection (AOI) plays a pivotal role in the manufacturing process, predominantly leveraging high-resolution imaging instruments for scanning purposes. It detects anomalies by analyzing image textures or patterns, making it an essential tool in industrial manufacturing and quality control. Despite its importance, the deployment of models for AOI often faces challenges. These include limited sample sizes, which hinder effective feature learning, variations among source domains, and sensitivities to changes in lighting and camera positions during imaging. These factors collectively compromise the accuracy of model predictions. Traditional AOI often fails to capitalize on the rich mechanism-parameter information from machines or inside images, including statistical parameters, which typically benefit AOI classification. To address this, we introduce an external modality-guided data mining framework, primarily rooted in optical character recognition (OCR), to extract statistical features from images as a second modality to enhance performance, termed OANet (Ocr-Aoi-Net). A key aspect of our approach is the alignment of external modality features, extracted using a single modality-aware model, with image features encoded by a convolutional neural network. This synergy enables a more refined fusion of semantic representations from different modalities. We further introduce feature refinement and a gating function in our OANet to optimize the combination of these features, enhancing inference and decision-making capabilities. Experimental outcomes show that our methodology considerably boosts the recall rate of the defect detection model and maintains high robustness even in challenging scenarios.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge