"Image": models, code, and papers
Improving visual image reconstruction from human brain activity using latent diffusion models via multiple decoded inputs
Jun 20, 2023
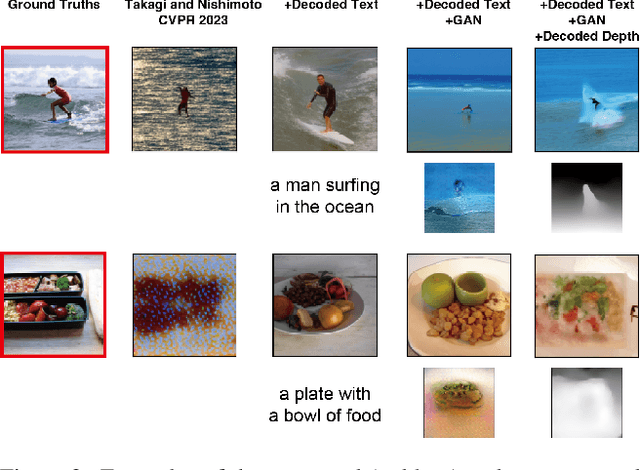
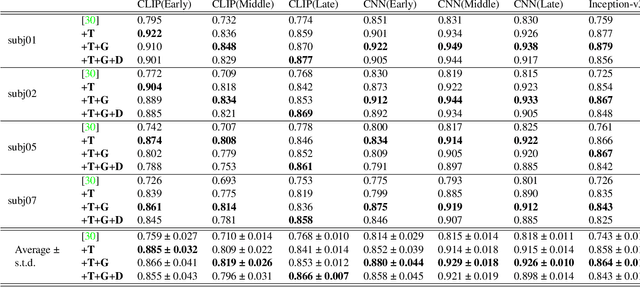

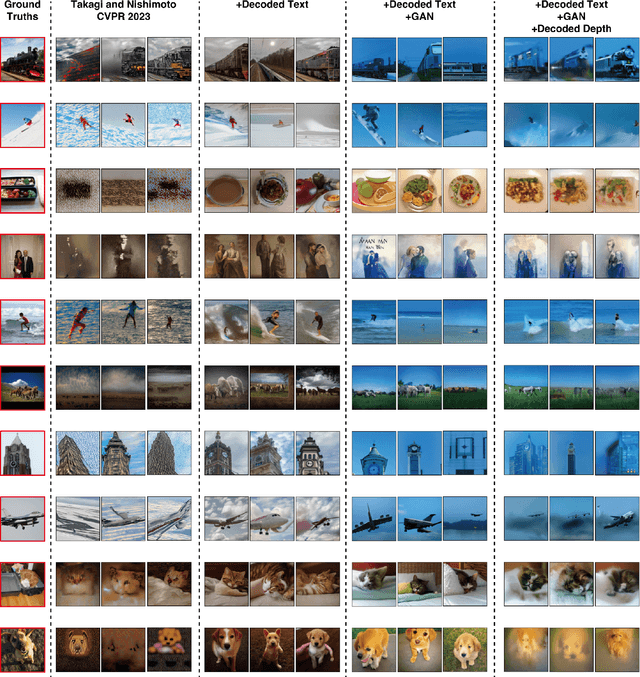
The integration of deep learning and neuroscience has been advancing rapidly, which has led to improvements in the analysis of brain activity and the understanding of deep learning models from a neuroscientific perspective. The reconstruction of visual experience from human brain activity is an area that has particularly benefited: the use of deep learning models trained on large amounts of natural images has greatly improved its quality, and approaches that combine the diverse information contained in visual experiences have proliferated rapidly in recent years. In this technical paper, by taking advantage of the simple and generic framework that we proposed (Takagi and Nishimoto, CVPR 2023), we examine the extent to which various additional decoding techniques affect the performance of visual experience reconstruction. Specifically, we combined our earlier work with the following three techniques: using decoded text from brain activity, nonlinear optimization for structural image reconstruction, and using decoded depth information from brain activity. We confirmed that these techniques contributed to improving accuracy over the baseline. We also discuss what researchers should consider when performing visual reconstruction using deep generative models trained on large datasets. Please check our webpage at https://sites.google.com/view/stablediffusion-with-brain/. Code is also available at https://github.com/yu-takagi/StableDiffusionReconstruction.
MOCA: Self-supervised Representation Learning by Predicting Masked Online Codebook Assignments
Jul 18, 2023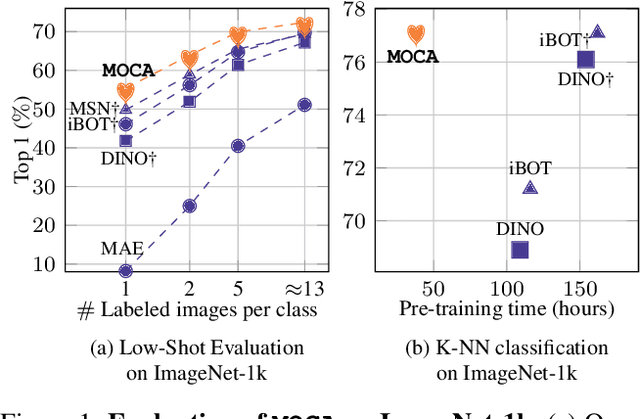

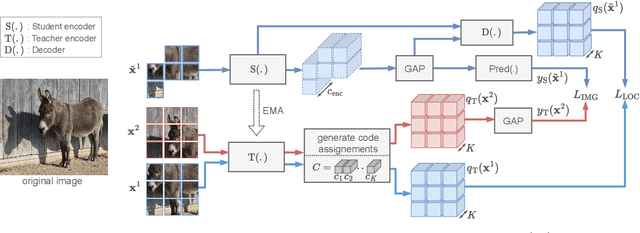
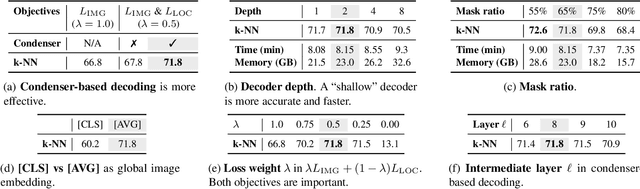
Self-supervised learning can be used for mitigating the greedy needs of Vision Transformer networks for very large fully-annotated datasets. Different classes of self-supervised learning offer representations with either good contextual reasoning properties, e.g., using masked image modeling strategies, or invariance to image perturbations, e.g., with contrastive methods. In this work, we propose a single-stage and standalone method, MOCA, which unifies both desired properties using novel mask-and-predict objectives defined with high-level features (instead of pixel-level details). Moreover, we show how to effectively employ both learning paradigms in a synergistic and computation-efficient way. Doing so, we achieve new state-of-the-art results on low-shot settings and strong experimental results in various evaluation protocols with a training that is at least 3 times faster than prior methods.
Negative-prompt Inversion: Fast Image Inversion for Editing with Text-guided Diffusion Models
May 26, 2023
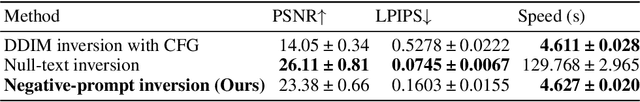
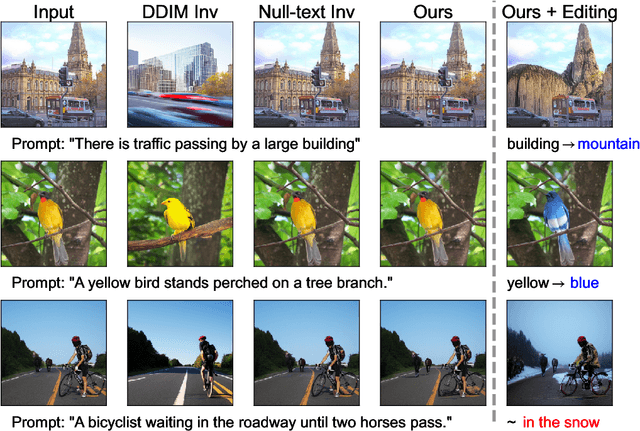

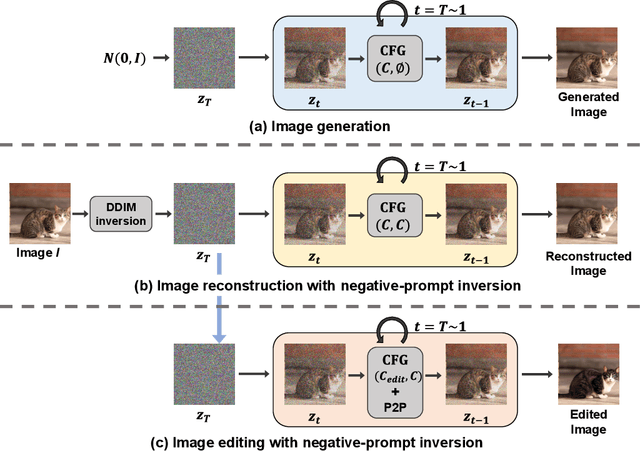

In image editing employing diffusion models, it is crucial to preserve the reconstruction quality of the original image while changing its style. Although existing methods ensure reconstruction quality through optimization, a drawback of these is the significant amount of time required for optimization. In this paper, we propose negative-prompt inversion, a method capable of achieving equivalent reconstruction solely through forward propagation without optimization, thereby enabling much faster editing processes. We experimentally demonstrate that the reconstruction quality of our method is comparable to that of existing methods, allowing for inversion at a resolution of 512 pixels and with 50 sampling steps within approximately 5 seconds, which is more than 30 times faster than null-text inversion. Reduction of the computation time by the proposed method further allows us to use a larger number of sampling steps in diffusion models to improve the reconstruction quality with a moderate increase in computation time.
Learning Full-Head 3D GANs from a Single-View Portrait Dataset
Jul 27, 2023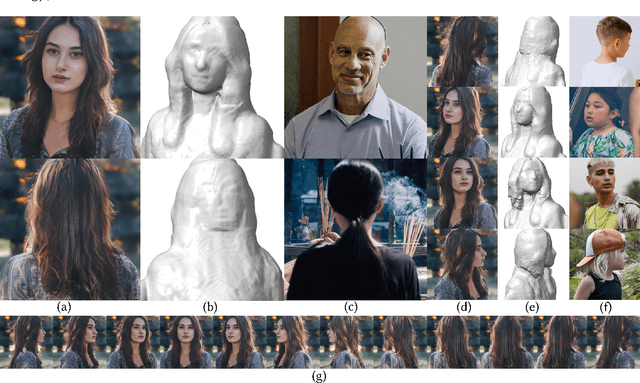
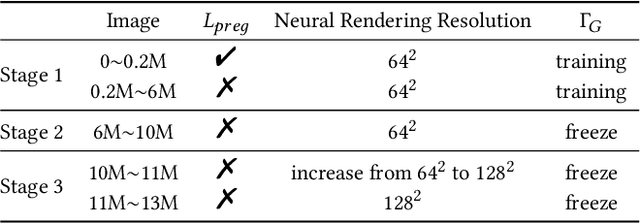


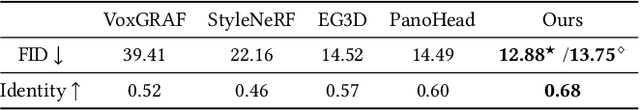
33D-aware face generators are commonly trained on 2D real-life face image datasets. Nevertheless, existing facial recognition methods often struggle to extract face data captured from various camera angles. Furthermore, in-the-wild images with diverse body poses introduce a high-dimensional challenge for 3D-aware generators, making it difficult to utilize data that contains complete neck and shoulder regions. Consequently, these face image datasets often contain only near-frontal face data, which poses challenges for 3D-aware face generators to construct \textit{full-head} 3D portraits. To this end, we first create the dataset {$\it{360}^{\circ}$}-\textit{Portrait}-\textit{HQ} (\textit{$\it{360}^{\circ}$PHQ}), which consists of high-quality single-view real portraits annotated with a variety of camera parameters {(the yaw angles span the entire $360^{\circ}$ range)} and body poses. We then propose \textit{3DPortraitGAN}, the first 3D-aware full-head portrait generator that learns a canonical 3D avatar distribution from the body-pose-various \textit{$\it{360}^{\circ}$PHQ} dataset with body pose self-learning. Our model can generate view-consistent portrait images from all camera angles (${360}^{\circ}$) with a full-head 3D representation. We incorporate a mesh-guided deformation field into volumetric rendering to produce deformed results to generate portrait images that conform to the body pose distribution of the dataset using our canonical generator. We integrate two pose predictors into our framework to predict more accurate body poses to address the issue of inaccurately estimated body poses in our dataset. Our experiments show that the proposed framework can generate view-consistent, realistic portrait images with complete geometry from all camera angles and accurately predict portrait body pose.
Multimodal Pathology Image Search Between H&E Slides and Multiplexed Immunofluorescent Images
Jun 11, 2023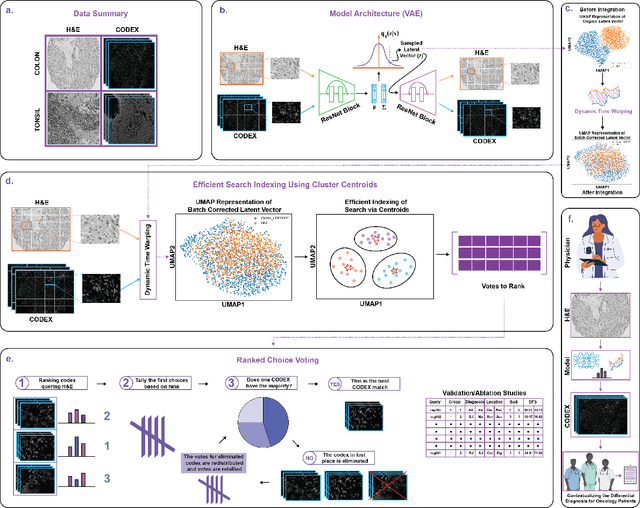
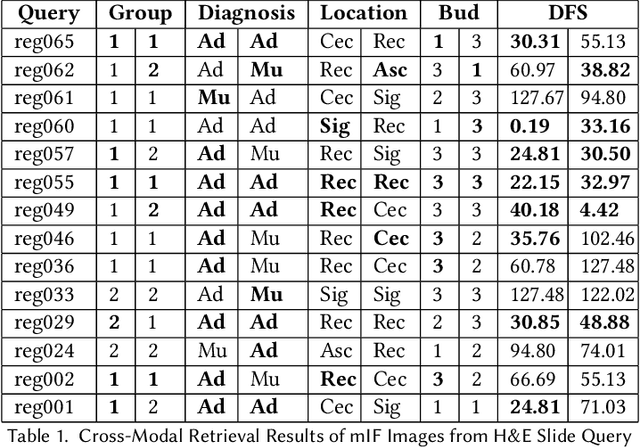
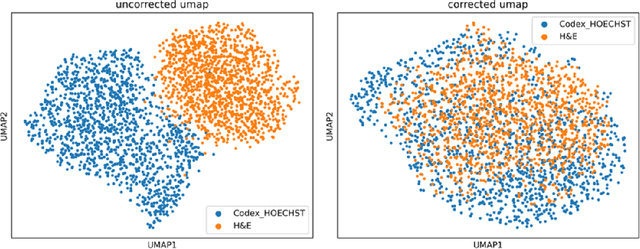
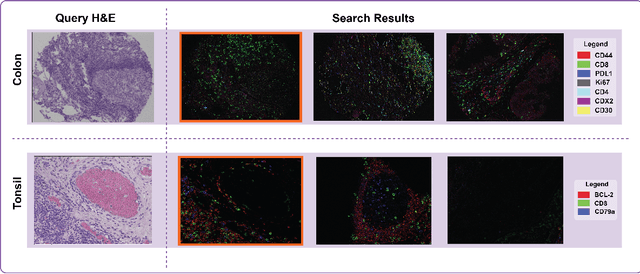
We present an approach for multimodal pathology image search, using dynamic time warping (DTW) on Variational Autoencoder (VAE) latent space that is fed into a ranked choice voting scheme to retrieve multiplexed immunofluorescent imaging (mIF) that is most similar to a query H&E slide. Through training the VAE and applying DTW, we align and compare mIF and H&E slides. Our method improves differential diagnosis and therapeutic decisions by integrating morphological H&E data with immunophenotyping from mIF, providing clinicians a rich perspective of disease states. This facilitates an understanding of the spatial relationships in tissue samples and could revolutionize the diagnostic process, enhancing precision and enabling personalized therapy selection. Our technique demonstrates feasibility using colorectal cancer and healthy tonsil samples. An exhaustive ablation study was conducted on a search engine designed to explore the correlation between multiplexed Immunofluorescence (mIF) and Hematoxylin and Eosin (H&E) staining, in order to validate its ability to map these distinct modalities into a unified vector space. Despite extreme class imbalance, the system demonstrated robustness and utility by returning similar results across various data features, which suggests potential for future use in multimodal histopathology data analysis.
Real-time Strawberry Detection Based on Improved YOLOv5s Architecture for Robotic Harvesting in open-field environment
Aug 08, 2023
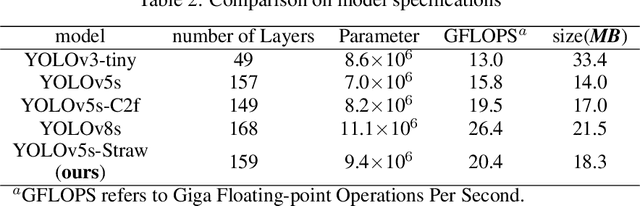
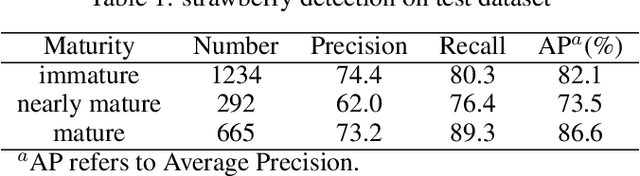


This study proposed a YOLOv5-based custom object detection model to detect strawberries in an outdoor environment. The original architecture of the YOLOv5s was modified by replacing the C3 module with the C2f module in the backbone network, which provided a better feature gradient flow. Secondly, the Spatial Pyramid Pooling Fast in the final layer of the backbone network of YOLOv5s was combined with Cross Stage Partial Net to improve the generalization ability over the strawberry dataset in this study. The proposed architecture was named YOLOv5s-Straw. The RGB images dataset of the strawberry canopy with three maturity classes (immature, nearly mature, and mature) was collected in open-field environment and augmented through a series of operations including brightness reduction, brightness increase, and noise adding. To verify the superiority of the proposed method for strawberry detection in open-field environment, four competitive detection models (YOLOv3-tiny, YOLOv5s, YOLOv5s-C2f, and YOLOv8s) were trained, and tested under the same computational environment and compared with YOLOv5s-Straw. The results showed that the highest mean average precision of 80.3% was achieved using the proposed architecture whereas the same was achieved with YOLOv3-tiny, YOLOv5s, YOLOv5s-C2f, and YOLOv8s were 73.4%, 77.8%, 79.8%, 79.3%, respectively. Specifically, the average precision of YOLOv5s-Straw was 82.1% in the immature class, 73.5% in the nearly mature class, and 86.6% in the mature class, which were 2.3% and 3.7%, respectively, higher than that of the latest YOLOv8s. The model included 8.6*10^6 network parameters with an inference speed of 18ms per image while the inference speed of YOLOv8s had a slower inference speed of 21.0ms and heavy parameters of 11.1*10^6, which indicates that the proposed model is fast enough for real time strawberry detection and localization for the robotic picking.
SDC-UDA: Volumetric Unsupervised Domain Adaptation Framework for Slice-Direction Continuous Cross-Modality Medical Image Segmentation
May 18, 2023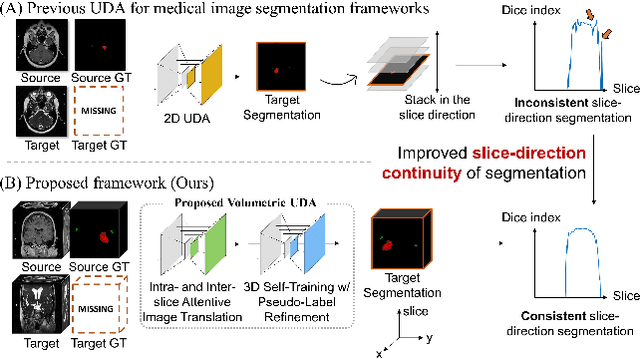
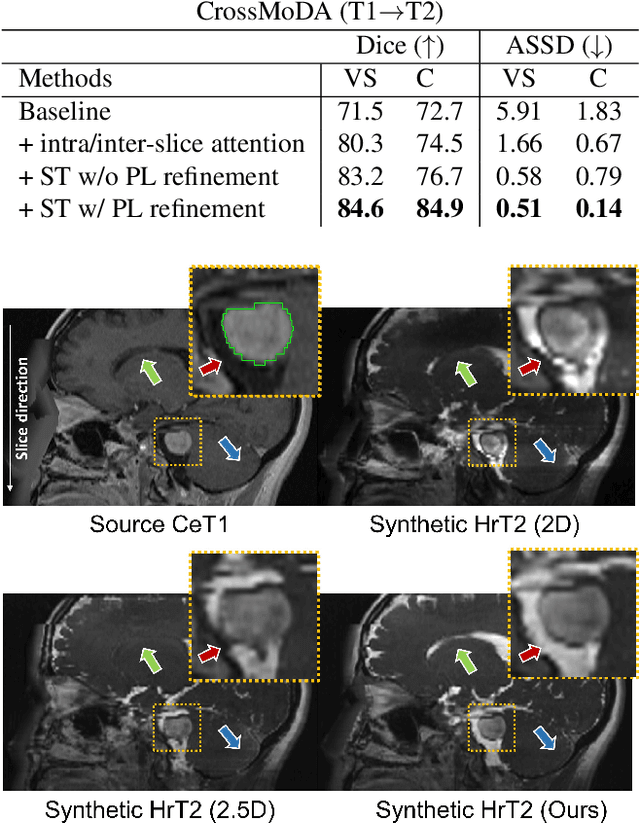
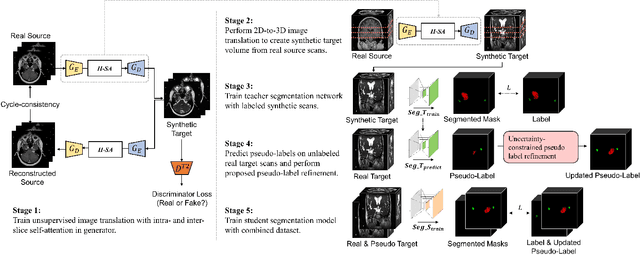
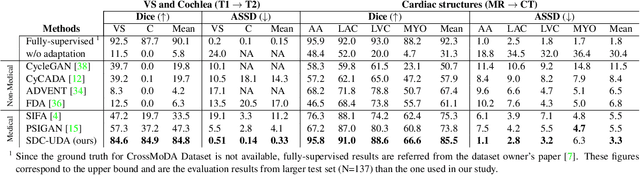
Recent advances in deep learning-based medical image segmentation studies achieve nearly human-level performance in fully supervised manner. However, acquiring pixel-level expert annotations is extremely expensive and laborious in medical imaging fields. Unsupervised domain adaptation (UDA) can alleviate this problem, which makes it possible to use annotated data in one imaging modality to train a network that can successfully perform segmentation on target imaging modality with no labels. In this work, we propose SDC-UDA, a simple yet effective volumetric UDA framework for slice-direction continuous cross-modality medical image segmentation which combines intra- and inter-slice self-attentive image translation, uncertainty-constrained pseudo-label refinement, and volumetric self-training. Our method is distinguished from previous methods on UDA for medical image segmentation in that it can obtain continuous segmentation in the slice direction, thereby ensuring higher accuracy and potential in clinical practice. We validate SDC-UDA with multiple publicly available cross-modality medical image segmentation datasets and achieve state-of-the-art segmentation performance, not to mention the superior slice-direction continuity of prediction compared to previous studies.
Bidirectional Temporal Diffusion Model for Temporally Consistent Human Animation
Jul 19, 2023
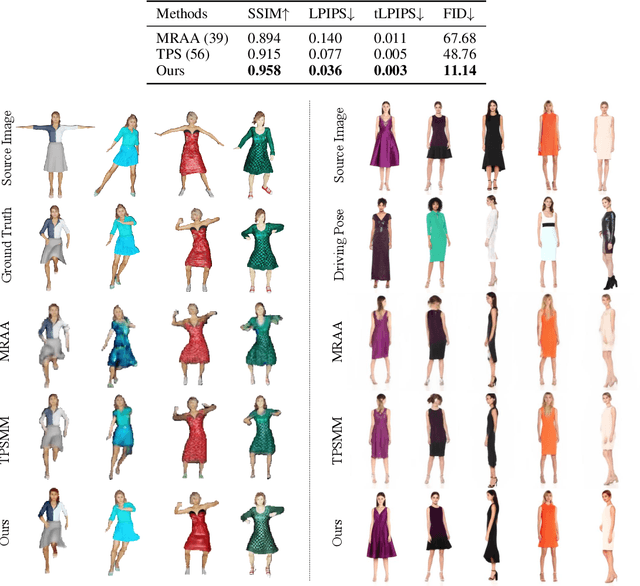
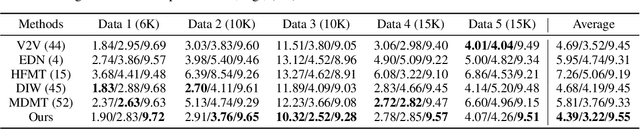
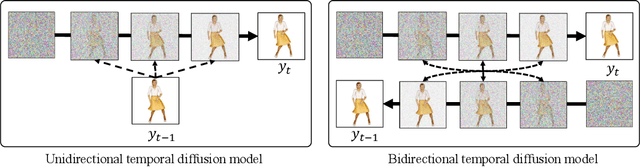
We introduce a method to generate temporally coherent human animation from a single image, a video, or a random noise. This problem has been formulated as modeling of an auto-regressive generation, i.e., to regress past frames to decode future frames. However, such unidirectional generation is highly prone to motion drifting over time, generating unrealistic human animation with significant artifacts such as appearance distortion. We claim that bidirectional temporal modeling enforces temporal coherence on a generative network by largely suppressing the motion ambiguity of human appearance. To prove our claim, we design a novel human animation framework using a denoising diffusion model: a neural network learns to generate the image of a person by denoising temporal Gaussian noises whose intermediate results are cross-conditioned bidirectionally between consecutive frames. In the experiments, our method demonstrates strong performance compared to existing unidirectional approaches with realistic temporal coherence
EchoGLAD: Hierarchical Graph Neural Networks for Left Ventricle Landmark Detection on Echocardiograms
Jul 23, 2023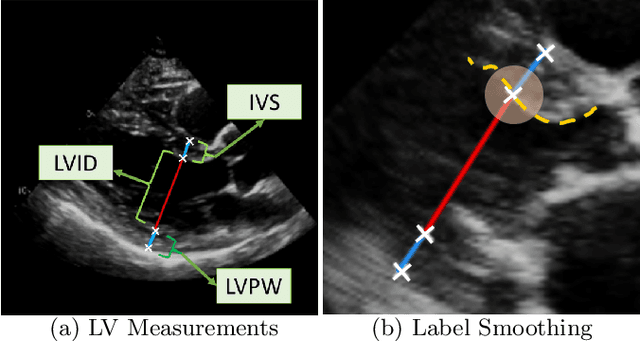
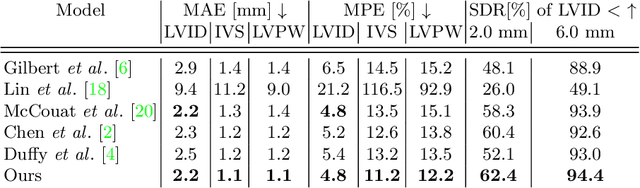
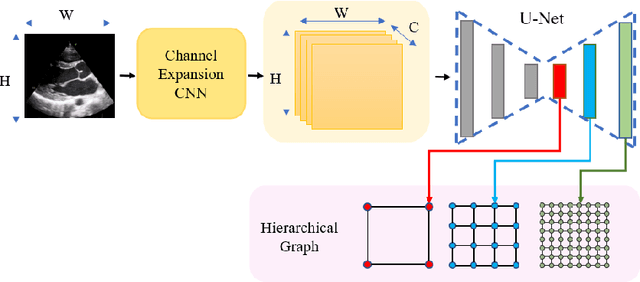
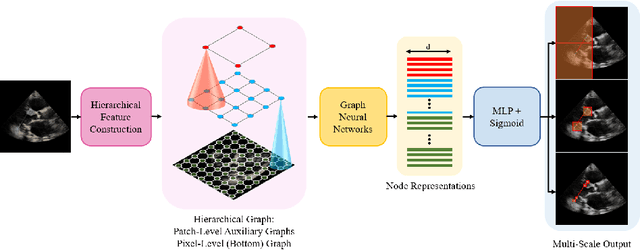
The functional assessment of the left ventricle chamber of the heart requires detecting four landmark locations and measuring the internal dimension of the left ventricle and the approximate mass of the surrounding muscle. The key challenge of automating this task with machine learning is the sparsity of clinical labels, i.e., only a few landmark pixels in a high-dimensional image are annotated, leading many prior works to heavily rely on isotropic label smoothing. However, such a label smoothing strategy ignores the anatomical information of the image and induces some bias. To address this challenge, we introduce an echocardiogram-based, hierarchical graph neural network (GNN) for left ventricle landmark detection (EchoGLAD). Our main contributions are: 1) a hierarchical graph representation learning framework for multi-resolution landmark detection via GNNs; 2) induced hierarchical supervision at different levels of granularity using a multi-level loss. We evaluate our model on a public and a private dataset under the in-distribution (ID) and out-of-distribution (OOD) settings. For the ID setting, we achieve the state-of-the-art mean absolute errors (MAEs) of 1.46 mm and 1.86 mm on the two datasets. Our model also shows better OOD generalization than prior works with a testing MAE of 4.3 mm.
Assessing Intra-class Diversity and Quality of Synthetically Generated Images in a Biomedical and Non-biomedical Setting
Jul 23, 2023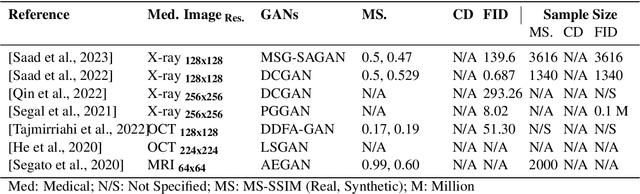
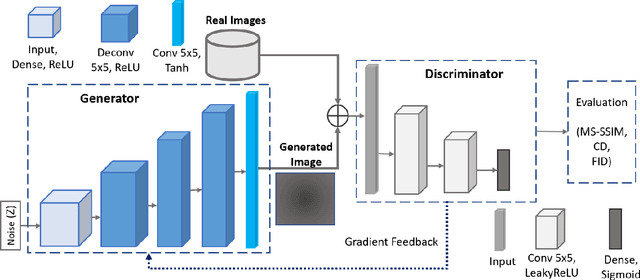
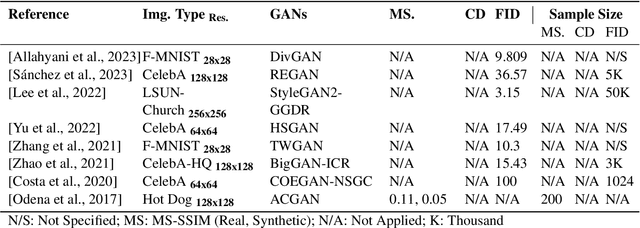
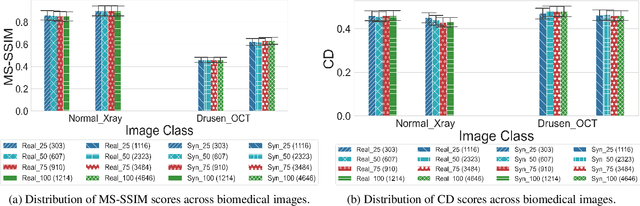
In biomedical image analysis, data imbalance is common across several imaging modalities. Data augmentation is one of the key solutions in addressing this limitation. Generative Adversarial Networks (GANs) are increasingly being relied upon for data augmentation tasks. Biomedical image features are sensitive to evaluating the efficacy of synthetic images. These features can have a significant impact on metric scores when evaluating synthetic images across different biomedical imaging modalities. Synthetically generated images can be evaluated by comparing the diversity and quality of real images. Multi-scale Structural Similarity Index Measure and Cosine Distance are used to evaluate intra-class diversity, while Frechet Inception Distance is used to evaluate the quality of synthetic images. Assessing these metrics for biomedical and non-biomedical imaging is important to investigate an informed strategy in evaluating the diversity and quality of synthetic images. In this work, an empirical assessment of these metrics is conducted for the Deep Convolutional GAN in a biomedical and non-biomedical setting. The diversity and quality of synthetic images are evaluated using different sample sizes. This research intends to investigate the variance in diversity and quality across biomedical and non-biomedical imaging modalities. Results demonstrate that the metrics scores for diversity and quality vary significantly across biomedical-to-biomedical and biomedical-to-non-biomedical imaging modalities.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge