"Image": models, code, and papers
ModeT: Learning Deformable Image Registration via Motion Decomposition Transformer
Jun 09, 2023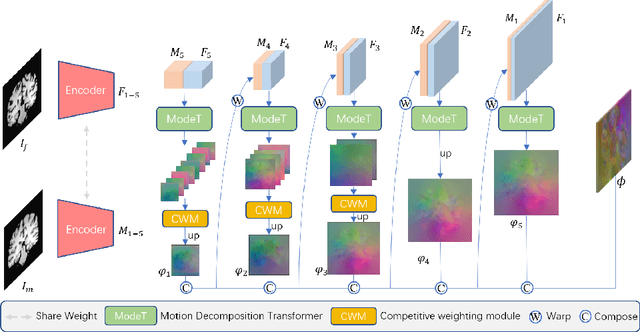
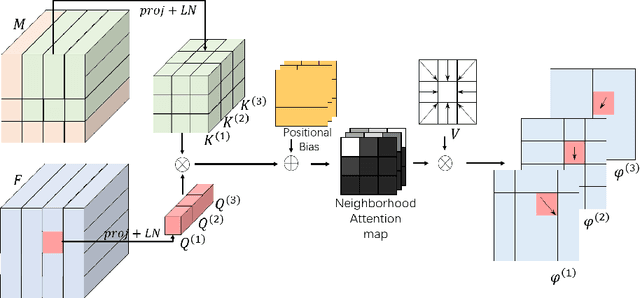
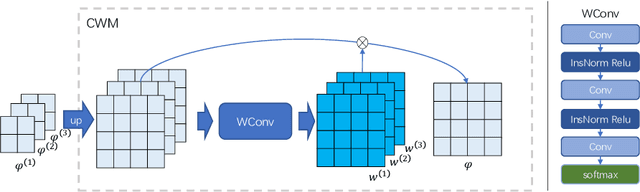
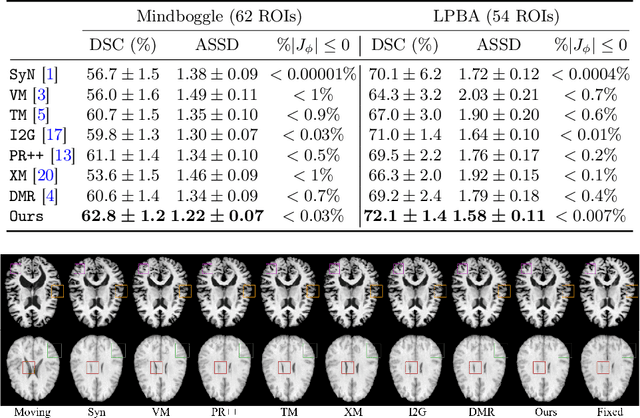
The Transformer structures have been widely used in computer vision and have recently made an impact in the area of medical image registration. However, the use of Transformer in most registration networks is straightforward. These networks often merely use the attention mechanism to boost the feature learning as the segmentation networks do, but do not sufficiently design to be adapted for the registration task. In this paper, we propose a novel motion decomposition Transformer (ModeT) to explicitly model multiple motion modalities by fully exploiting the intrinsic capability of the Transformer structure for deformation estimation. The proposed ModeT naturally transforms the multi-head neighborhood attention relationship into the multi-coordinate relationship to model multiple motion modes. Then the competitive weighting module (CWM) fuses multiple deformation sub-fields to generate the resulting deformation field. Extensive experiments on two public brain magnetic resonance imaging (MRI) datasets show that our method outperforms current state-of-the-art registration networks and Transformers, demonstrating the potential of our ModeT for the challenging non-rigid deformation estimation problem. The benchmarks and our code are publicly available at https://github.com/ZAX130/SmileCode.
Instructive Feature Enhancement for Dichotomous Medical Image Segmentation
Jun 06, 2023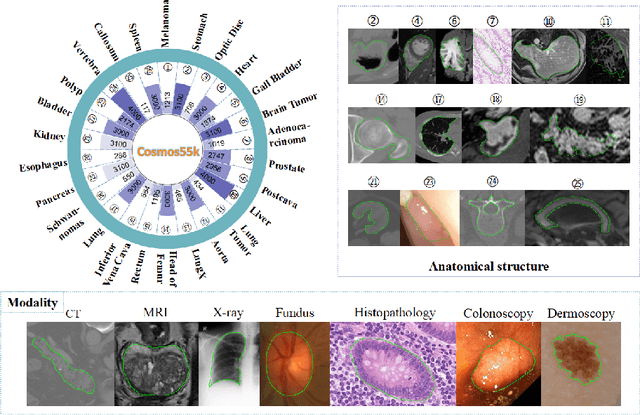
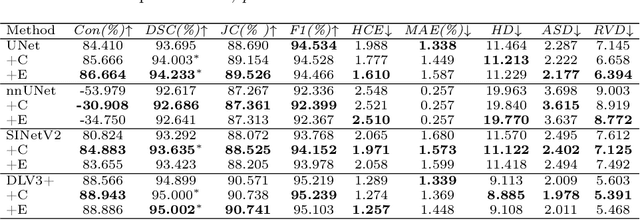
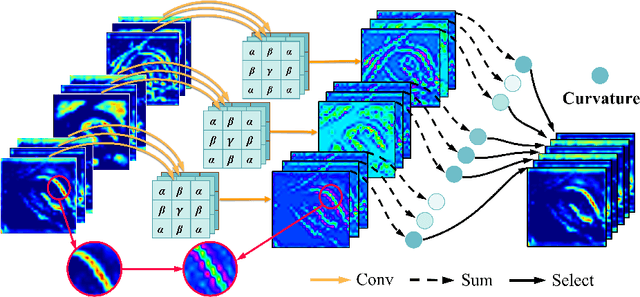
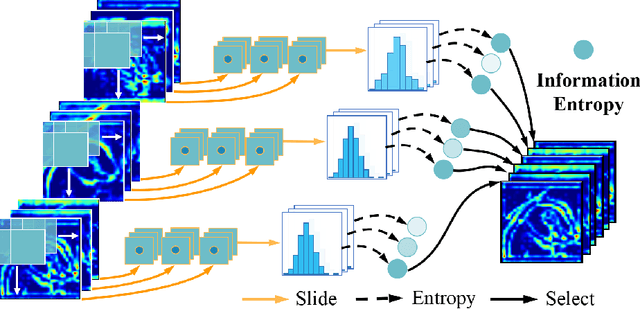
Deep neural networks have been widely applied in dichotomous medical image segmentation (DMIS) of many anatomical structures in several modalities, achieving promising performance. However, existing networks tend to struggle with task-specific, heavy and complex designs to improve accuracy. They made little instructions to which feature channels would be more beneficial for segmentation, and that may be why the performance and universality of these segmentation models are hindered. In this study, we propose an instructive feature enhancement approach, namely IFE, to adaptively select feature channels with rich texture cues and strong discriminability to enhance raw features based on local curvature or global information entropy criteria. Being plug-and-play and applicable for diverse DMIS tasks, IFE encourages the model to focus on texture-rich features which are especially important for the ambiguous and challenging boundary identification, simultaneously achieving simplicity, universality, and certain interpretability. To evaluate the proposed IFE, we constructed the first large-scale DMIS dataset Cosmos55k, which contains 55,023 images from 7 modalities and 26 anatomical structures. Extensive experiments show that IFE can improve the performance of classic segmentation networks across different anatomies and modalities with only slight modifications. Code is available at https://github.com/yezi-66/IFE
Weakly supervised segmentation of intracranial aneurysms using a 3D focal modulation UNet
Aug 06, 2023
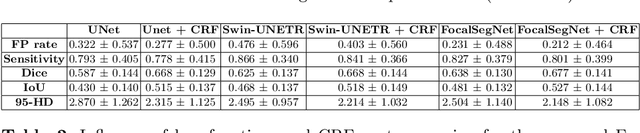
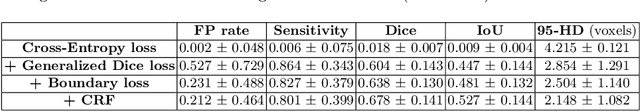
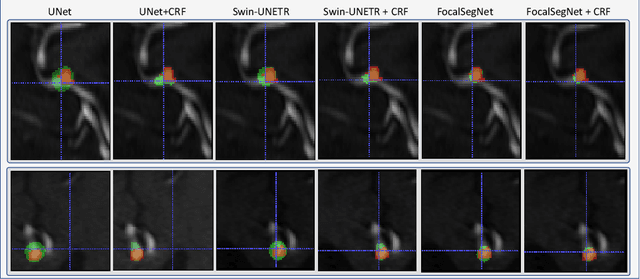
Accurate identification and quantification of unruptured intracranial aneurysms (UIAs) are essential for the risk assessment and treatment decisions of this cerebrovascular disorder. Current assessment based on 2D manual measures of aneurysms on 3D magnetic resonance angiography (MRA) is sub-optimal and time-consuming. Automatic 3D measures can significantly benefit the clinical workflow and treatment outcomes. However, one major issue in medical image segmentation is the need for large well-annotated data, which can be expensive to obtain. Techniques that mitigate the requirement, such as weakly supervised learning with coarse labels are highly desirable. In this paper, we leverage coarse labels of UIAs from time-of-flight MRAs to obtain refined UIAs segmentation using a novel 3D focal modulation UNet, called FocalSegNet and conditional random field (CRF) postprocessing, with a Dice score of 0.68 and 95% Hausdorff distance of 0.95 mm. We evaluated the performance of the proposed algorithms against the state-of-the-art 3D UNet and Swin-UNETR, and demonstrated the superiority of the proposed FocalSegNet and the benefit of focal modulation for the task.
Memory Encoding Model
Aug 02, 2023We explore a new class of brain encoding model by adding memory-related information as input. Memory is an essential brain mechanism that works alongside visual stimuli. During a vision-memory cognitive task, we found the non-visual brain is largely predictable using previously seen images. Our Memory Encoding Model (Mem) won the Algonauts 2023 visual brain competition even without model ensemble (single model score 66.8, ensemble score 70.8). Our ensemble model without memory input (61.4) can also stand a 3rd place. Furthermore, we observe periodic delayed brain response correlated to 6th-7th prior image, and hippocampus also showed correlated activity timed with this periodicity. We conjuncture that the periodic replay could be related to memory mechanism to enhance the working memory.
Embracing Compact and Robust Architectures for Multi-Exposure Image Fusion
May 20, 2023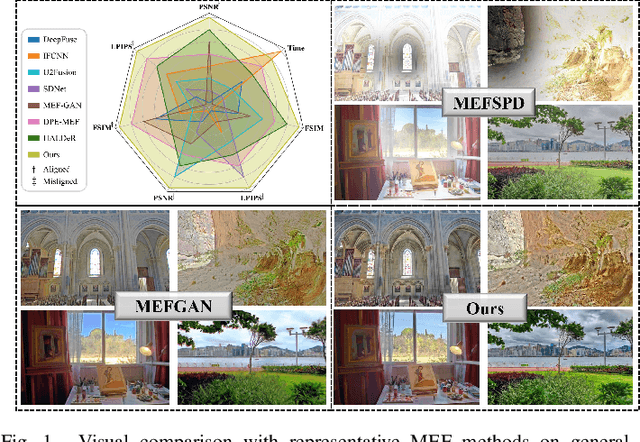
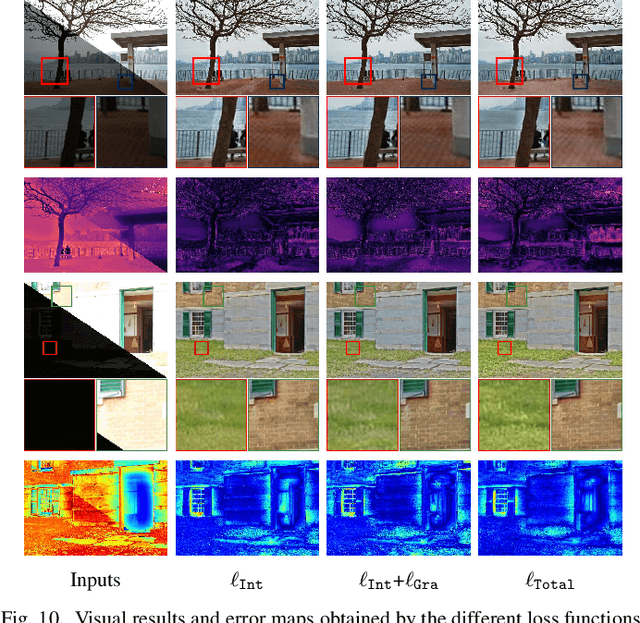
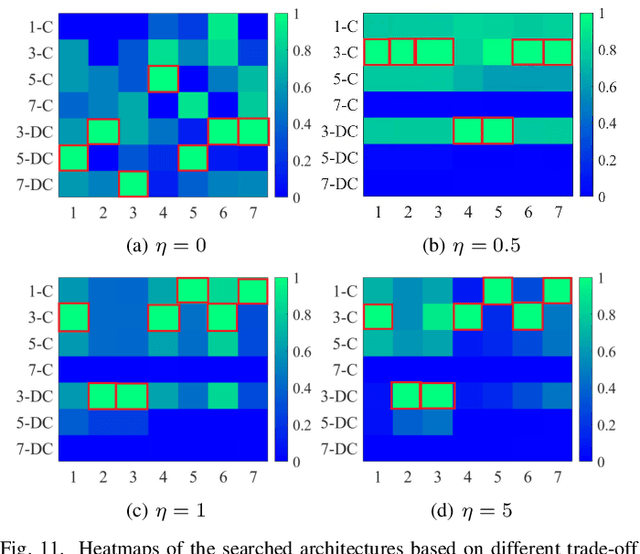
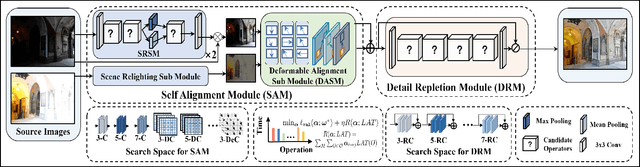
In recent years, deep learning-based methods have achieved remarkable progress in multi-exposure image fusion. However, existing methods rely on aligned image pairs, inevitably generating artifacts when faced with device shaking in real-world scenarios. Moreover, these learning-based methods are built on handcrafted architectures and operations by increasing network depth or width, neglecting different exposure characteristics. As a result, these direct cascaded architectures with redundant parameters fail to achieve highly effective inference time and lead to massive computation. To alleviate these issues, in this paper, we propose a search-based paradigm, involving self-alignment and detail repletion modules for robust multi-exposure image fusion. By utilizing scene relighting and deformable convolutions, the self-alignment module can accurately align images despite camera movement. Furthermore, by imposing a hardware-sensitive constraint, we introduce neural architecture search to discover compact and efficient networks, investigating effective feature representation for fusion. We realize the state-of-the-art performance in comparison to various competitive schemes, yielding a 4.02% and 29.34% improvement in PSNR for general and misaligned scenarios, respectively, while reducing inference time by 68.1%. The source code will be available at https://github.com/LiuZhu-CV/CRMEF.
SLPT: Selective Labeling Meets Prompt Tuning on Label-Limited Lesion Segmentation
Aug 09, 2023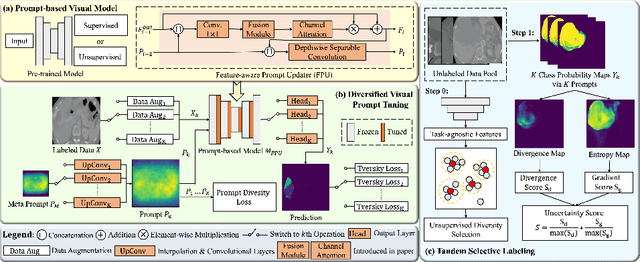
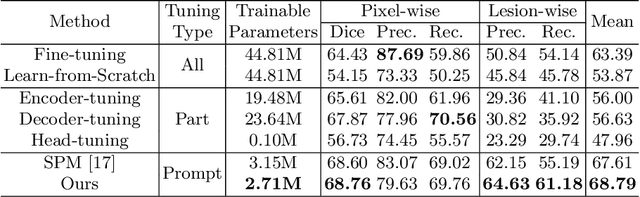
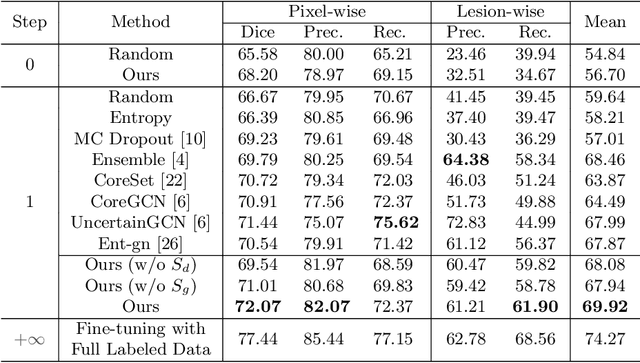
Medical image analysis using deep learning is often challenged by limited labeled data and high annotation costs. Fine-tuning the entire network in label-limited scenarios can lead to overfitting and suboptimal performance. Recently, prompt tuning has emerged as a more promising technique that introduces a few additional tunable parameters as prompts to a task-agnostic pre-trained model, and updates only these parameters using supervision from limited labeled data while keeping the pre-trained model unchanged. However, previous work has overlooked the importance of selective labeling in downstream tasks, which aims to select the most valuable downstream samples for annotation to achieve the best performance with minimum annotation cost. To address this, we propose a framework that combines selective labeling with prompt tuning (SLPT) to boost performance in limited labels. Specifically, we introduce a feature-aware prompt updater to guide prompt tuning and a TandEm Selective LAbeling (TESLA) strategy. TESLA includes unsupervised diversity selection and supervised selection using prompt-based uncertainty. In addition, we propose a diversified visual prompt tuning strategy to provide multi-prompt-based discrepant predictions for TESLA. We evaluate our method on liver tumor segmentation and achieve state-of-the-art performance, outperforming traditional fine-tuning with only 6% of tunable parameters, also achieving 94% of full-data performance by labeling only 5% of the data.
An automated pipeline for quantitative T2* fetal body MRI and segmentation at low field
Aug 09, 2023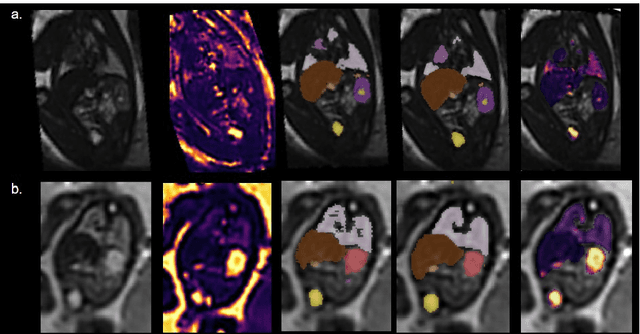
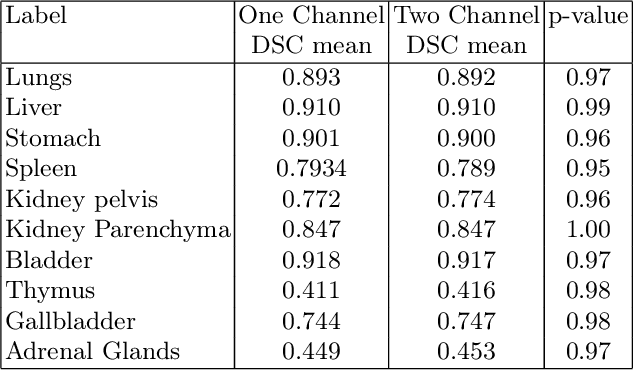
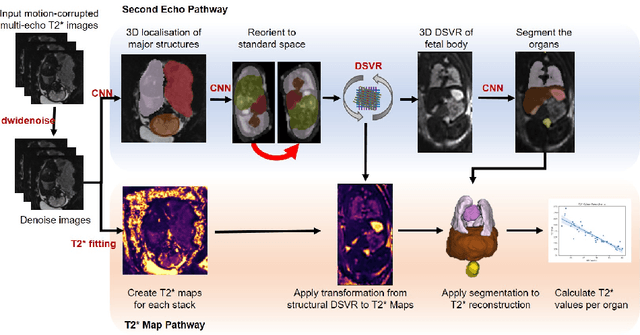
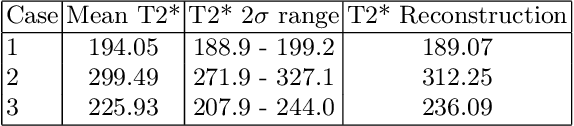
Fetal Magnetic Resonance Imaging at low field strengths is emerging as an exciting direction in perinatal health. Clinical low field (0.55T) scanners are beneficial for fetal imaging due to their reduced susceptibility-induced artefacts, increased T2* values, and wider bore (widening access for the increasingly obese pregnant population). However, the lack of standard automated image processing tools such as segmentation and reconstruction hampers wider clinical use. In this study, we introduce a semi-automatic pipeline using quantitative MRI for the fetal body at low field strength resulting in fast and detailed quantitative T2* relaxometry analysis of all major fetal body organs. Multi-echo dynamic sequences of the fetal body were acquired and reconstructed into a single high-resolution volume using deformable slice-to-volume reconstruction, generating both structural and quantitative T2* 3D volumes. A neural network trained using a semi-supervised approach was created to automatically segment these fetal body 3D volumes into ten different organs (resulting in dice values > 0.74 for 8 out of 10 organs). The T2* values revealed a strong relationship with GA in the lungs, liver, and kidney parenchyma (R^2>0.5). This pipeline was used successfully for a wide range of GAs (17-40 weeks), and is robust to motion artefacts. Low field fetal MRI can be used to perform advanced MRI analysis, and is a viable option for clinical scanning.
SegMatch: A semi-supervised learning method for surgical instrument segmentation
Aug 09, 2023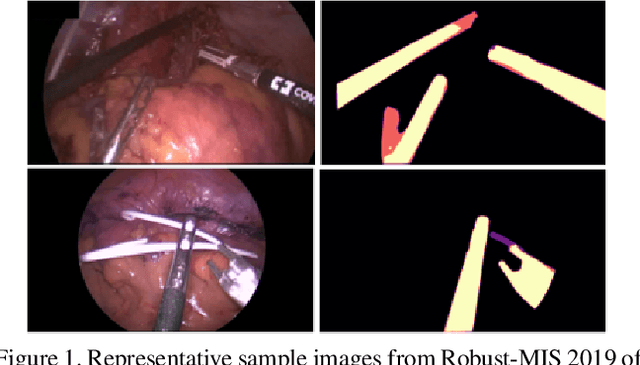
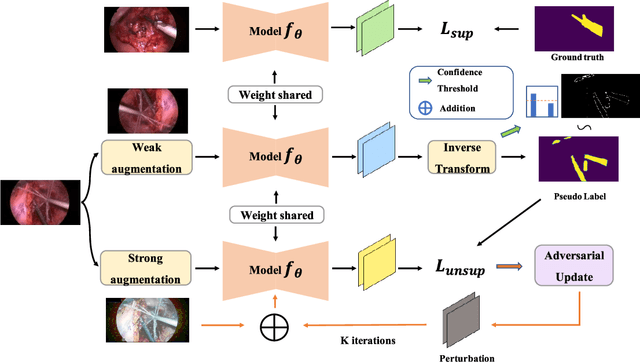
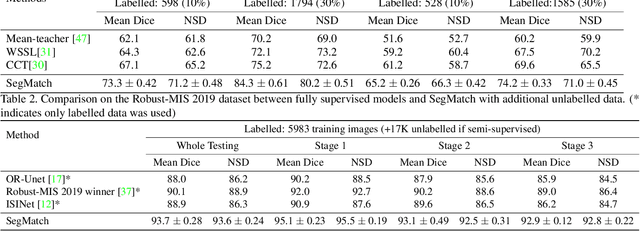
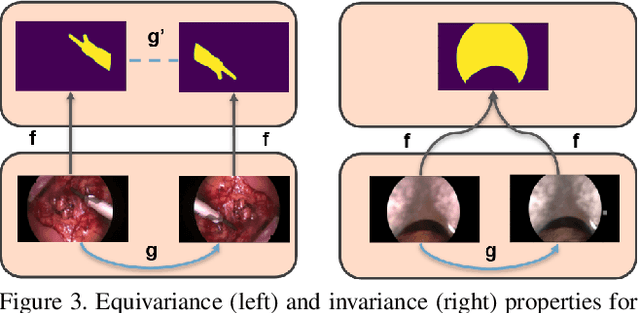
Surgical instrument segmentation is recognised as a key enabler to provide advanced surgical assistance and improve computer assisted interventions. In this work, we propose SegMatch, a semi supervised learning method to reduce the need for expensive annotation for laparoscopic and robotic surgical images. SegMatch builds on FixMatch, a widespread semi supervised classification pipeline combining consistency regularization and pseudo labelling, and adapts it for the purpose of segmentation. In our proposed SegMatch, the unlabelled images are weakly augmented and fed into the segmentation model to generate a pseudo-label to enforce the unsupervised loss against the output of the model for the adversarial augmented image on the pixels with a high confidence score. Our adaptation for segmentation tasks includes carefully considering the equivariance and invariance properties of the augmentation functions we rely on. To increase the relevance of our augmentations, we depart from using only handcrafted augmentations and introduce a trainable adversarial augmentation strategy. Our algorithm was evaluated on the MICCAI Instrument Segmentation Challenge datasets Robust-MIS 2019 and EndoVis 2017. Our results demonstrate that adding unlabelled data for training purposes allows us to surpass the performance of fully supervised approaches which are limited by the availability of training data in these challenges. SegMatch also outperforms a range of state-of-the-art semi-supervised learning semantic segmentation models in different labelled to unlabelled data ratios.
Learning Concise and Descriptive Attributes for Visual Recognition
Aug 07, 2023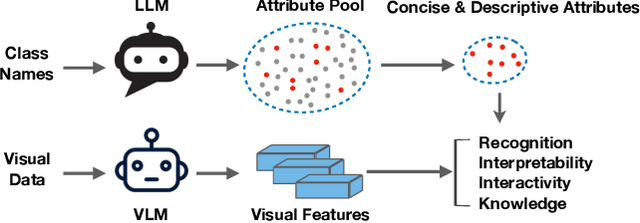
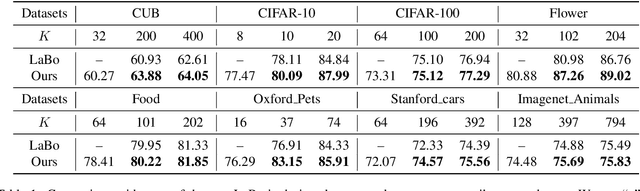
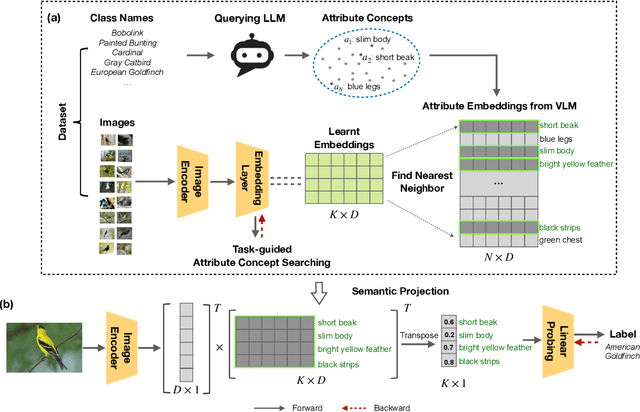
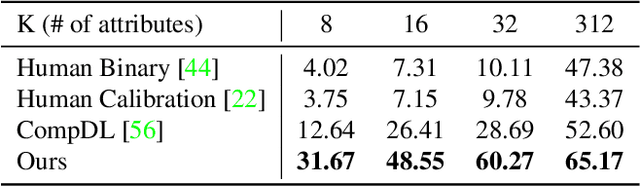
Recent advances in foundation models present new opportunities for interpretable visual recognition -- one can first query Large Language Models (LLMs) to obtain a set of attributes that describe each class, then apply vision-language models to classify images via these attributes. Pioneering work shows that querying thousands of attributes can achieve performance competitive with image features. However, our further investigation on 8 datasets reveals that LLM-generated attributes in a large quantity perform almost the same as random words. This surprising finding suggests that significant noise may be present in these attributes. We hypothesize that there exist subsets of attributes that can maintain the classification performance with much smaller sizes, and propose a novel learning-to-search method to discover those concise sets of attributes. As a result, on the CUB dataset, our method achieves performance close to that of massive LLM-generated attributes (e.g., 10k attributes for CUB), yet using only 32 attributes in total to distinguish 200 bird species. Furthermore, our new paradigm demonstrates several additional benefits: higher interpretability and interactivity for humans, and the ability to summarize knowledge for a recognition task.
Fixed Inter-Neuron Covariability Induces Adversarial Robustness
Aug 07, 2023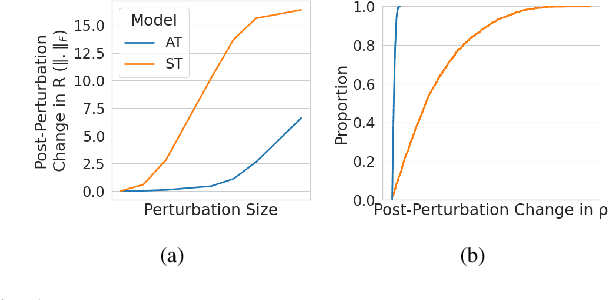
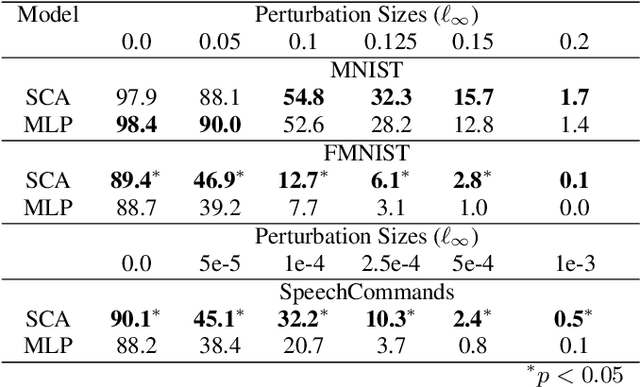
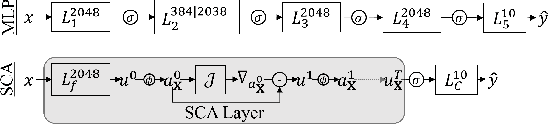
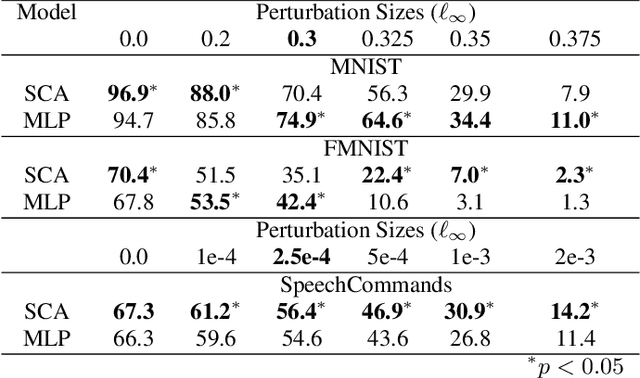
The vulnerability to adversarial perturbations is a major flaw of Deep Neural Networks (DNNs) that raises question about their reliability when in real-world scenarios. On the other hand, human perception, which DNNs are supposed to emulate, is highly robust to such perturbations, indicating that there may be certain features of the human perception that make it robust but are not represented in the current class of DNNs. One such feature is that the activity of biological neurons is correlated and the structure of this correlation tends to be rather rigid over long spans of times, even if it hampers performance and learning. We hypothesize that integrating such constraints on the activations of a DNN would improve its adversarial robustness, and, to test this hypothesis, we have developed the Self-Consistent Activation (SCA) layer, which comprises of neurons whose activations are consistent with each other, as they conform to a fixed, but learned, covariability pattern. When evaluated on image and sound recognition tasks, the models with a SCA layer achieved high accuracy, and exhibited significantly greater robustness than multi-layer perceptron models to state-of-the-art Auto-PGD adversarial attacks \textit{without being trained on adversarially perturbed data
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge