"Image": models, code, and papers
MiDaS v3.1 -- A Model Zoo for Robust Monocular Relative Depth Estimation
Jul 26, 2023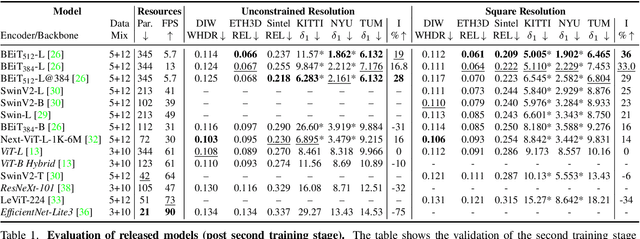
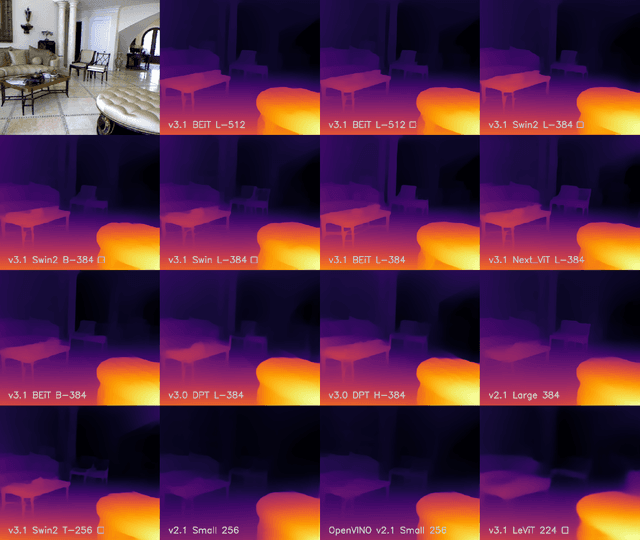
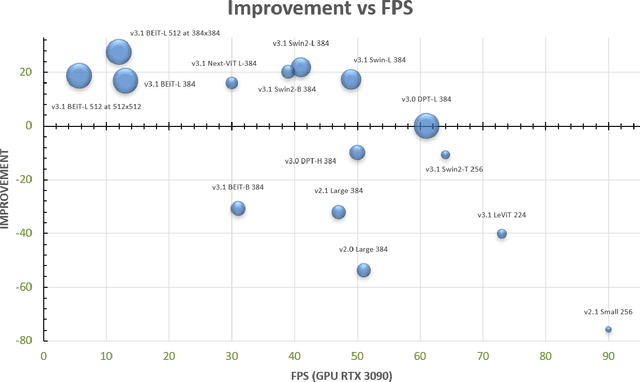
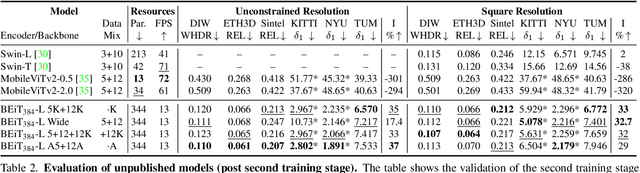

We release MiDaS v3.1 for monocular depth estimation, offering a variety of new models based on different encoder backbones. This release is motivated by the success of transformers in computer vision, with a large variety of pretrained vision transformers now available. We explore how using the most promising vision transformers as image encoders impacts depth estimation quality and runtime of the MiDaS architecture. Our investigation also includes recent convolutional approaches that achieve comparable quality to vision transformers in image classification tasks. While the previous release MiDaS v3.0 solely leverages the vanilla vision transformer ViT, MiDaS v3.1 offers additional models based on BEiT, Swin, SwinV2, Next-ViT and LeViT. These models offer different performance-runtime tradeoffs. The best model improves the depth estimation quality by 28% while efficient models enable downstream tasks requiring high frame rates. We also describe the general process for integrating new backbones. A video summarizing the work can be found at https://youtu.be/UjaeNNFf9sE and the code is available at https://github.com/isl-org/MiDaS.
SegMatch: A semi-supervised learning method for surgical instrument segmentation
Aug 09, 2023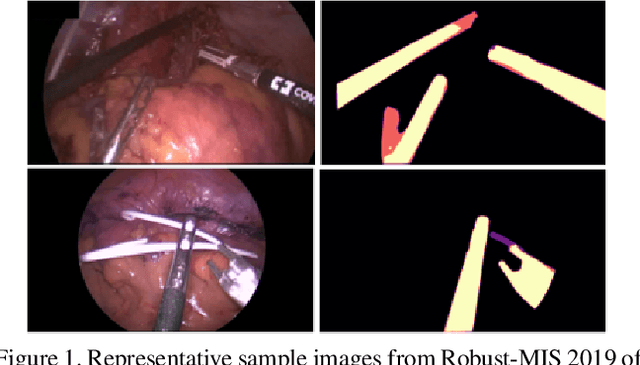
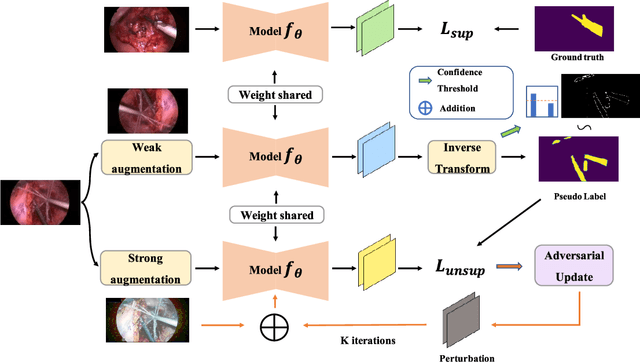
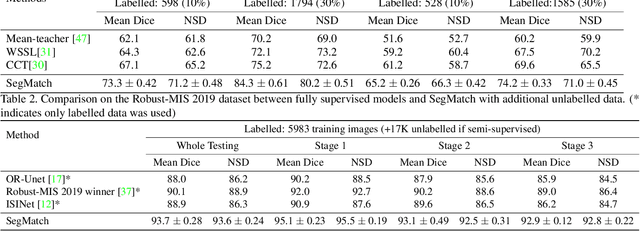
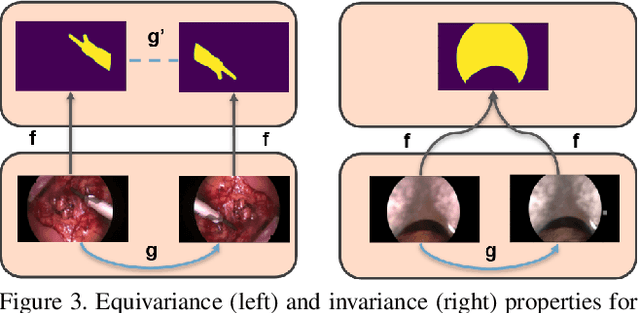
Surgical instrument segmentation is recognised as a key enabler to provide advanced surgical assistance and improve computer assisted interventions. In this work, we propose SegMatch, a semi supervised learning method to reduce the need for expensive annotation for laparoscopic and robotic surgical images. SegMatch builds on FixMatch, a widespread semi supervised classification pipeline combining consistency regularization and pseudo labelling, and adapts it for the purpose of segmentation. In our proposed SegMatch, the unlabelled images are weakly augmented and fed into the segmentation model to generate a pseudo-label to enforce the unsupervised loss against the output of the model for the adversarial augmented image on the pixels with a high confidence score. Our adaptation for segmentation tasks includes carefully considering the equivariance and invariance properties of the augmentation functions we rely on. To increase the relevance of our augmentations, we depart from using only handcrafted augmentations and introduce a trainable adversarial augmentation strategy. Our algorithm was evaluated on the MICCAI Instrument Segmentation Challenge datasets Robust-MIS 2019 and EndoVis 2017. Our results demonstrate that adding unlabelled data for training purposes allows us to surpass the performance of fully supervised approaches which are limited by the availability of training data in these challenges. SegMatch also outperforms a range of state-of-the-art semi-supervised learning semantic segmentation models in different labelled to unlabelled data ratios.
SLPT: Selective Labeling Meets Prompt Tuning on Label-Limited Lesion Segmentation
Aug 09, 2023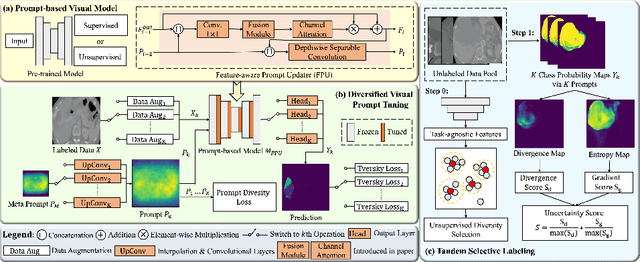
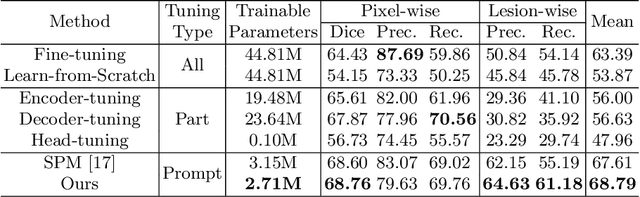
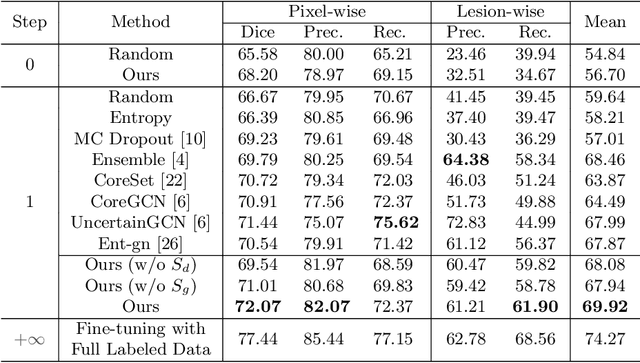
Medical image analysis using deep learning is often challenged by limited labeled data and high annotation costs. Fine-tuning the entire network in label-limited scenarios can lead to overfitting and suboptimal performance. Recently, prompt tuning has emerged as a more promising technique that introduces a few additional tunable parameters as prompts to a task-agnostic pre-trained model, and updates only these parameters using supervision from limited labeled data while keeping the pre-trained model unchanged. However, previous work has overlooked the importance of selective labeling in downstream tasks, which aims to select the most valuable downstream samples for annotation to achieve the best performance with minimum annotation cost. To address this, we propose a framework that combines selective labeling with prompt tuning (SLPT) to boost performance in limited labels. Specifically, we introduce a feature-aware prompt updater to guide prompt tuning and a TandEm Selective LAbeling (TESLA) strategy. TESLA includes unsupervised diversity selection and supervised selection using prompt-based uncertainty. In addition, we propose a diversified visual prompt tuning strategy to provide multi-prompt-based discrepant predictions for TESLA. We evaluate our method on liver tumor segmentation and achieve state-of-the-art performance, outperforming traditional fine-tuning with only 6% of tunable parameters, also achieving 94% of full-data performance by labeling only 5% of the data.
An automated pipeline for quantitative T2* fetal body MRI and segmentation at low field
Aug 09, 2023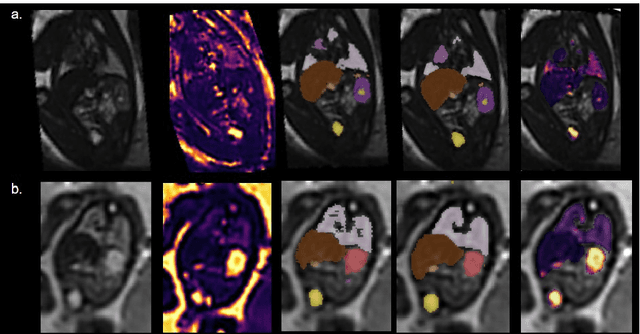
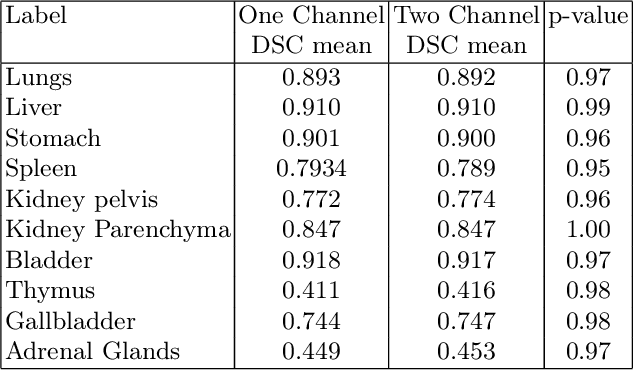
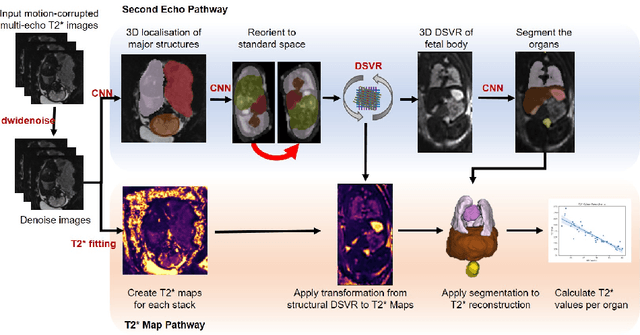
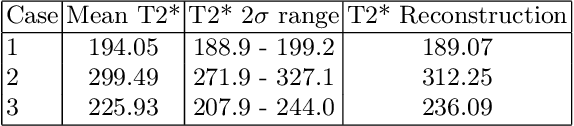
Fetal Magnetic Resonance Imaging at low field strengths is emerging as an exciting direction in perinatal health. Clinical low field (0.55T) scanners are beneficial for fetal imaging due to their reduced susceptibility-induced artefacts, increased T2* values, and wider bore (widening access for the increasingly obese pregnant population). However, the lack of standard automated image processing tools such as segmentation and reconstruction hampers wider clinical use. In this study, we introduce a semi-automatic pipeline using quantitative MRI for the fetal body at low field strength resulting in fast and detailed quantitative T2* relaxometry analysis of all major fetal body organs. Multi-echo dynamic sequences of the fetal body were acquired and reconstructed into a single high-resolution volume using deformable slice-to-volume reconstruction, generating both structural and quantitative T2* 3D volumes. A neural network trained using a semi-supervised approach was created to automatically segment these fetal body 3D volumes into ten different organs (resulting in dice values > 0.74 for 8 out of 10 organs). The T2* values revealed a strong relationship with GA in the lungs, liver, and kidney parenchyma (R^2>0.5). This pipeline was used successfully for a wide range of GAs (17-40 weeks), and is robust to motion artefacts. Low field fetal MRI can be used to perform advanced MRI analysis, and is a viable option for clinical scanning.
LLMScore: Unveiling the Power of Large Language Models in Text-to-Image Synthesis Evaluation
May 18, 2023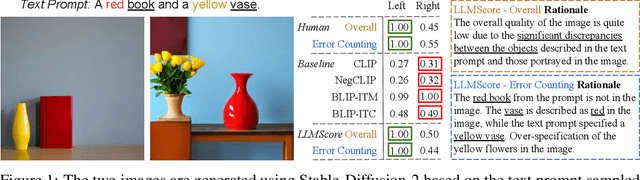
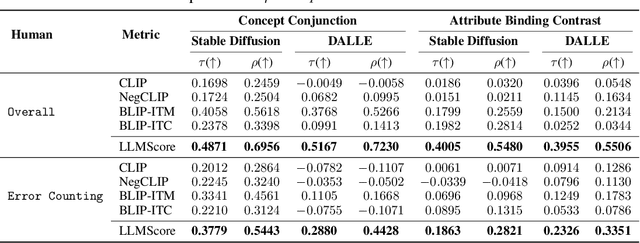
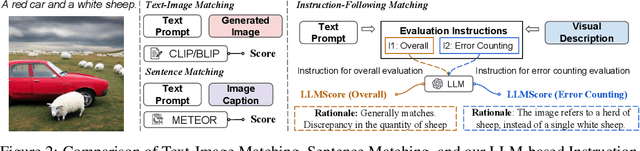
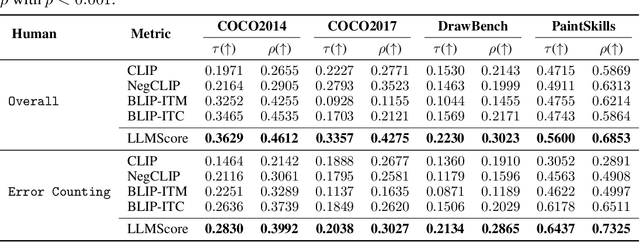
Existing automatic evaluation on text-to-image synthesis can only provide an image-text matching score, without considering the object-level compositionality, which results in poor correlation with human judgments. In this work, we propose LLMScore, a new framework that offers evaluation scores with multi-granularity compositionality. LLMScore leverages the large language models (LLMs) to evaluate text-to-image models. Initially, it transforms the image into image-level and object-level visual descriptions. Then an evaluation instruction is fed into the LLMs to measure the alignment between the synthesized image and the text, ultimately generating a score accompanied by a rationale. Our substantial analysis reveals the highest correlation of LLMScore with human judgments on a wide range of datasets (Attribute Binding Contrast, Concept Conjunction, MSCOCO, DrawBench, PaintSkills). Notably, our LLMScore achieves Kendall's tau correlation with human evaluations that is 58.8% and 31.2% higher than the commonly-used text-image matching metrics CLIP and BLIP, respectively.
Interactive Segmentation for Diverse Gesture Types Without Context
Jul 20, 2023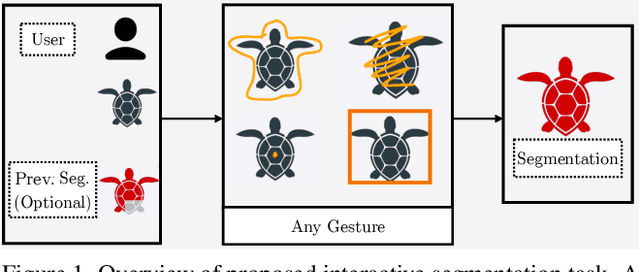
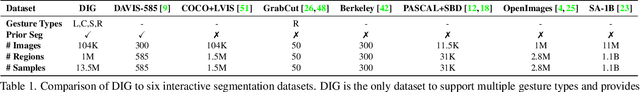
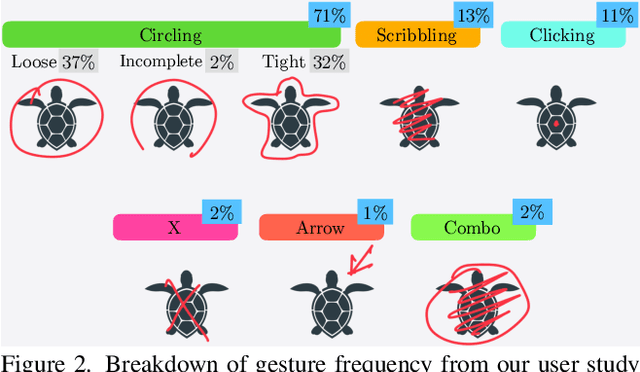
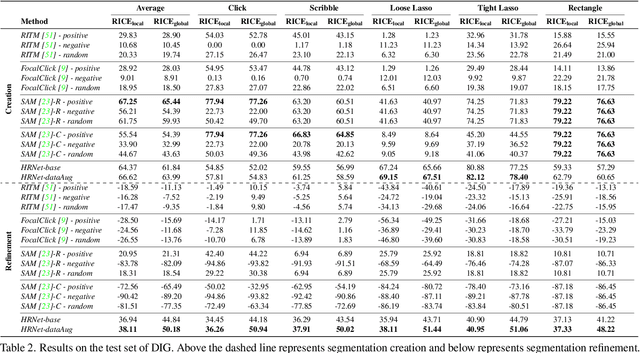
Interactive segmentation entails a human marking an image to guide how a model either creates or edits a segmentation. Our work addresses limitations of existing methods: they either only support one gesture type for marking an image (e.g., either clicks or scribbles) or require knowledge of the gesture type being employed, and require specifying whether marked regions should be included versus excluded in the final segmentation. We instead propose a simplified interactive segmentation task where a user only must mark an image, where the input can be of any gesture type without specifying the gesture type. We support this new task by introducing the first interactive segmentation dataset with multiple gesture types as well as a new evaluation metric capable of holistically evaluating interactive segmentation algorithms. We then analyze numerous interactive segmentation algorithms, including ones adapted for our novel task. While we observe promising performance overall, we also highlight areas for future improvement. To facilitate further extensions of this work, we publicly share our new dataset at https://github.com/joshmyersdean/dig.
Weakly supervised segmentation of intracranial aneurysms using a 3D focal modulation UNet
Aug 06, 2023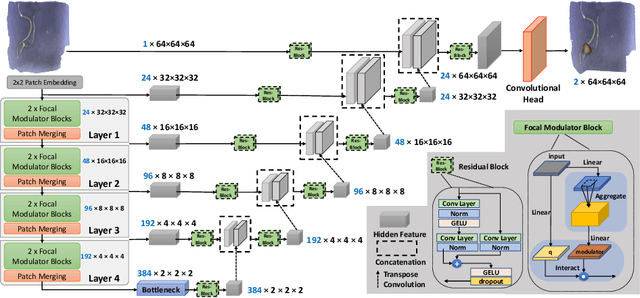
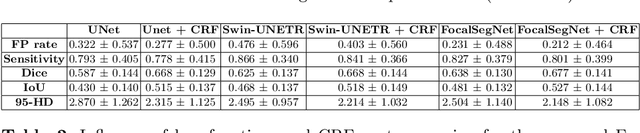
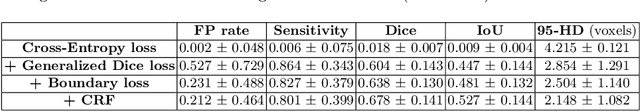
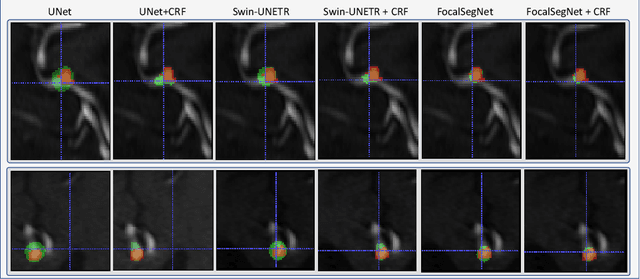
Accurate identification and quantification of unruptured intracranial aneurysms (UIAs) are essential for the risk assessment and treatment decisions of this cerebrovascular disorder. Current assessment based on 2D manual measures of aneurysms on 3D magnetic resonance angiography (MRA) is sub-optimal and time-consuming. Automatic 3D measures can significantly benefit the clinical workflow and treatment outcomes. However, one major issue in medical image segmentation is the need for large well-annotated data, which can be expensive to obtain. Techniques that mitigate the requirement, such as weakly supervised learning with coarse labels are highly desirable. In this paper, we leverage coarse labels of UIAs from time-of-flight MRAs to obtain refined UIAs segmentation using a novel 3D focal modulation UNet, called FocalSegNet and conditional random field (CRF) postprocessing, with a Dice score of 0.68 and 95% Hausdorff distance of 0.95 mm. We evaluated the performance of the proposed algorithms against the state-of-the-art 3D UNet and Swin-UNETR, and demonstrated the superiority of the proposed FocalSegNet and the benefit of focal modulation for the task.
Fact-Checking of AI-Generated Reports
Jul 27, 2023With advances in generative artificial intelligence (AI), it is now possible to produce realistic-looking automated reports for preliminary reads of radiology images. This can expedite clinical workflows, improve accuracy and reduce overall costs. However, it is also well-known that such models often hallucinate, leading to false findings in the generated reports. In this paper, we propose a new method of fact-checking of AI-generated reports using their associated images. Specifically, the developed examiner differentiates real and fake sentences in reports by learning the association between an image and sentences describing real or potentially fake findings. To train such an examiner, we first created a new dataset of fake reports by perturbing the findings in the original ground truth radiology reports associated with images. Text encodings of real and fake sentences drawn from these reports are then paired with image encodings to learn the mapping to real/fake labels. The utility of such an examiner is demonstrated for verifying automatically generated reports by detecting and removing fake sentences. Future generative AI approaches can use the resulting tool to validate their reports leading to a more responsible use of AI in expediting clinical workflows.
OmniDataComposer: A Unified Data Structure for Multimodal Data Fusion and Infinite Data Generation
Aug 08, 2023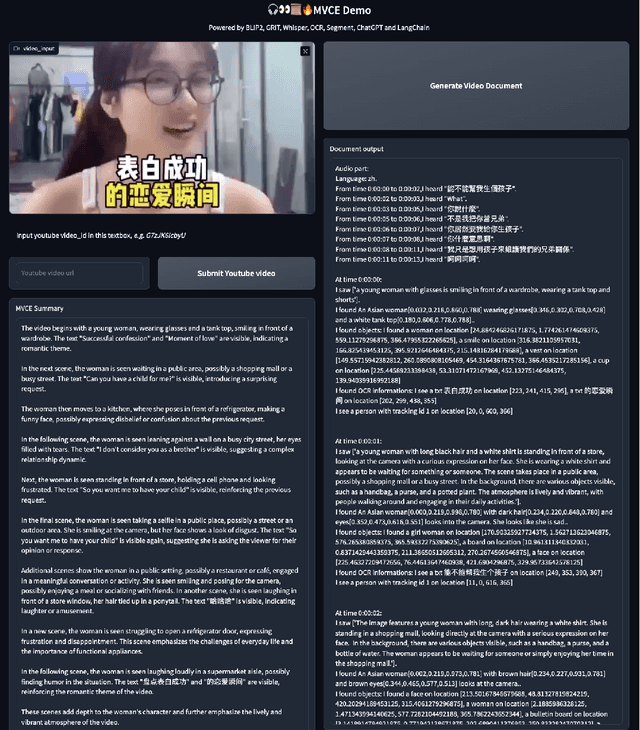
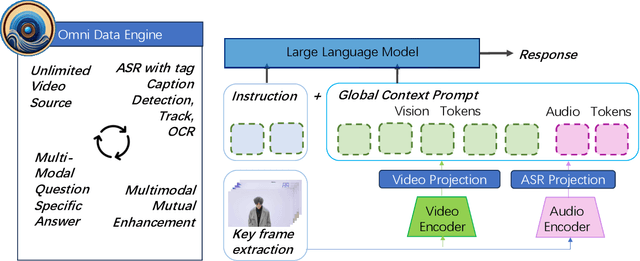

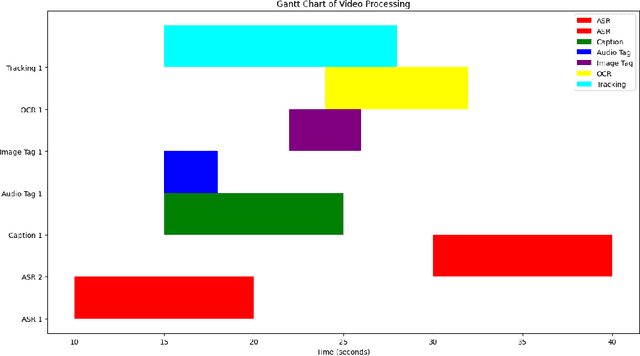
This paper presents OmniDataComposer, an innovative approach for multimodal data fusion and unlimited data generation with an intent to refine and uncomplicate interplay among diverse data modalities. Coming to the core breakthrough, it introduces a cohesive data structure proficient in processing and merging multimodal data inputs, which include video, audio, and text. Our crafted algorithm leverages advancements across multiple operations such as video/image caption extraction, dense caption extraction, Automatic Speech Recognition (ASR), Optical Character Recognition (OCR), Recognize Anything Model(RAM), and object tracking. OmniDataComposer is capable of identifying over 6400 categories of objects, substantially broadening the spectrum of visual information. It amalgamates these diverse modalities, promoting reciprocal enhancement among modalities and facilitating cross-modal data correction. \textbf{The final output metamorphoses each video input into an elaborate sequential document}, virtually transmuting videos into thorough narratives, making them easier to be processed by large language models. Future prospects include optimizing datasets for each modality to encourage unlimited data generation. This robust base will offer priceless insights to models like ChatGPT, enabling them to create higher quality datasets for video captioning and easing question-answering tasks based on video content. OmniDataComposer inaugurates a new stage in multimodal learning, imparting enormous potential for augmenting AI's understanding and generation of complex, real-world data.
Application for White Spot Syndrome Virus (WSSV) Monitoring using Edge Machine Learning
Aug 08, 2023The aquaculture industry, strongly reliant on shrimp exports, faces challenges due to viral infections like the White Spot Syndrome Virus (WSSV) that severely impact output yields. In this context, computer vision can play a significant role in identifying features not immediately evident to skilled or untrained eyes, potentially reducing the time required to report WSSV infections. In this study, the challenge of limited data for WSSV recognition was addressed. A mobile application dedicated to data collection and monitoring was developed to facilitate the creation of an image dataset to train a WSSV recognition model and improve country-wide disease surveillance. The study also includes a thorough analysis of WSSV recognition to address the challenge of imbalanced learning and on-device inference. The models explored, MobileNetV3-Small and EfficientNetV2-B0, gained an F1-Score of 0.72 and 0.99 respectively. The saliency heatmaps of both models were also observed to uncover the "black-box" nature of these models and to gain insight as to what features in the images are most important in making a prediction. These results highlight the effectiveness and limitations of using models designed for resource-constrained devices and balancing their performance in accurately recognizing WSSV, providing valuable information and direction in the use of computer vision in this domain.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge