"Image": models, code, and papers
Deepbet: Fast brain extraction of T1-weighted MRI using Convolutional Neural Networks
Aug 14, 2023
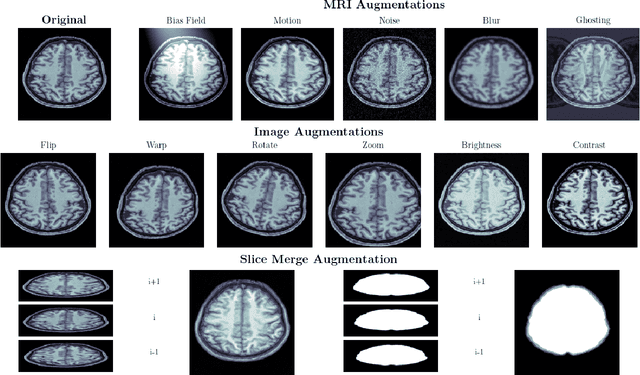
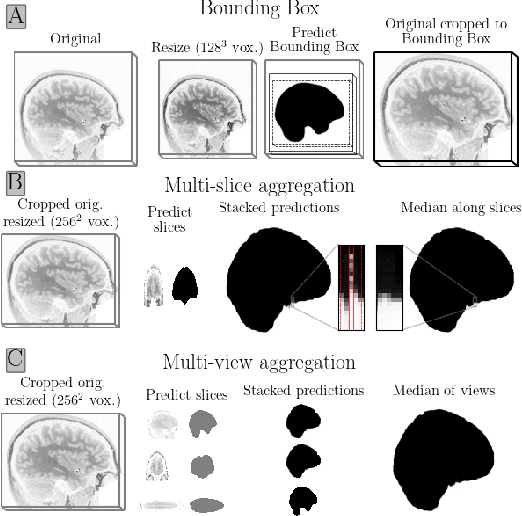
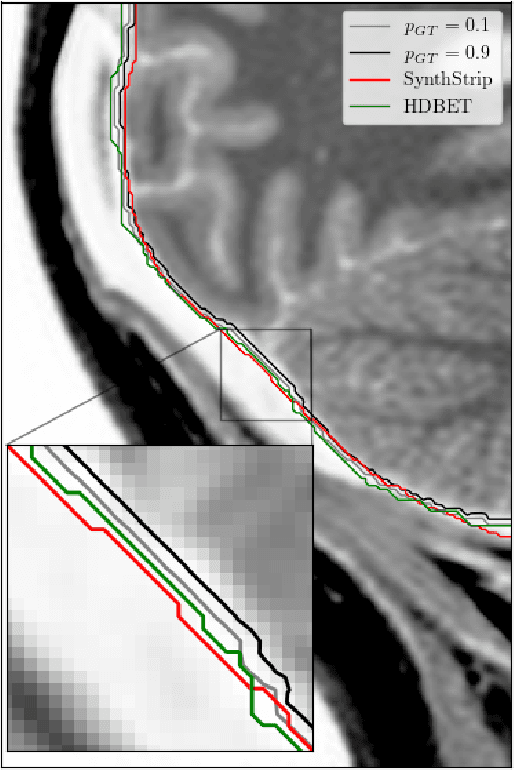
Brain extraction in magnetic resonance imaging (MRI) data is an important segmentation step in many neuroimaging preprocessing pipelines. Image segmentation is one of the research fields in which deep learning had the biggest impact in recent years enabling high precision segmentation with minimal compute. Consequently, traditional brain extraction methods are now being replaced by deep learning-based methods. Here, we used a unique dataset comprising 568 T1-weighted (T1w) MR images from 191 different studies in combination with cutting edge deep learning methods to build a fast, high-precision brain extraction tool called deepbet. deepbet uses LinkNet, a modern UNet architecture, in a two stage prediction process. This increases its segmentation performance, setting a novel state-of-the-art performance during cross-validation with a median Dice score (DSC) of 99.0% on unseen datasets, outperforming current state of the art models (DSC = 97.8% and DSC = 97.9%). While current methods are more sensitive to outliers, resulting in Dice scores as low as 76.5%, deepbet manages to achieve a Dice score of > 96.9% for all samples. Finally, our model accelerates brain extraction by a factor of ~10 compared to current methods, enabling the processing of one image in ~2 seconds on low level hardware.
A One Stop 3D Target Reconstruction and multilevel Segmentation Method
Aug 14, 2023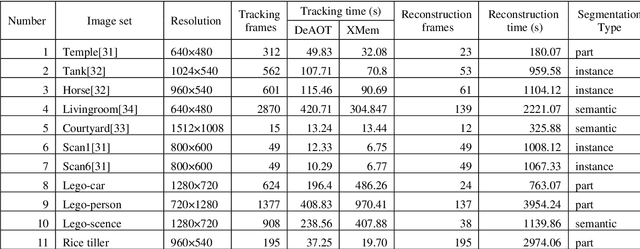
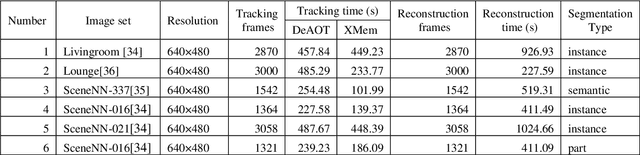
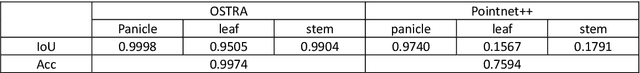
3D object reconstruction and multilevel segmentation are fundamental to computer vision research. Existing algorithms usually perform 3D scene reconstruction and target objects segmentation independently, and the performance is not fully guaranteed due to the challenge of the 3D segmentation. Here we propose an open-source one stop 3D target reconstruction and multilevel segmentation framework (OSTRA), which performs segmentation on 2D images, tracks multiple instances with segmentation labels in the image sequence, and then reconstructs labelled 3D objects or multiple parts with Multi-View Stereo (MVS) or RGBD-based 3D reconstruction methods. We extend object tracking and 3D reconstruction algorithms to support continuous segmentation labels to leverage the advances in the 2D image segmentation, especially the Segment-Anything Model (SAM) which uses the pretrained neural network without additional training for new scenes, for 3D object segmentation. OSTRA supports most popular 3D object models including point cloud, mesh and voxel, and achieves high performance for semantic segmentation, instance segmentation and part segmentation on several 3D datasets. It even surpasses the manual segmentation in scenes with complex structures and occlusions. Our method opens up a new avenue for reconstructing 3D targets embedded with rich multi-scale segmentation information in complex scenes. OSTRA is available from https://github.com/ganlab/OSTRA.
Polyp-SAM++: Can A Text Guided SAM Perform Better for Polyp Segmentation?
Aug 12, 2023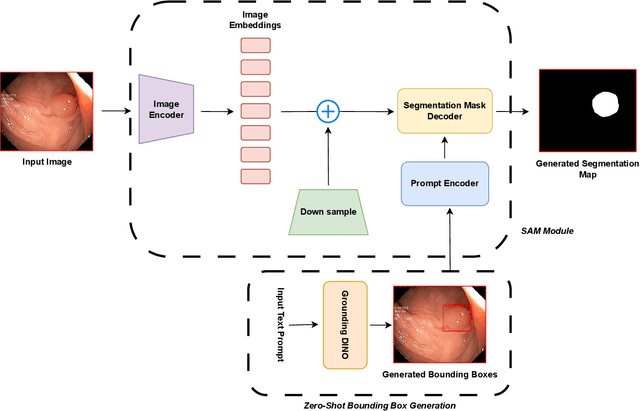
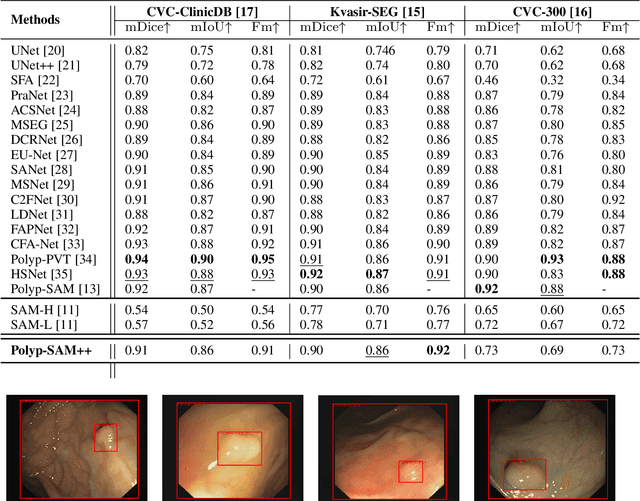
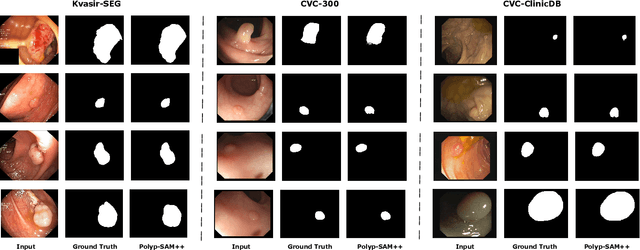
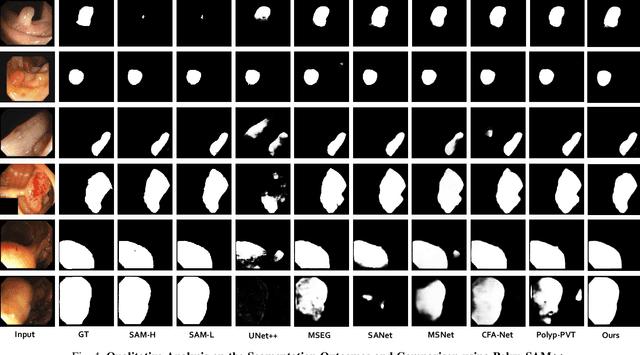
Meta recently released SAM (Segment Anything Model) which is a general-purpose segmentation model. SAM has shown promising results in a wide variety of segmentation tasks including medical image segmentation. In the field of medical image segmentation, polyp segmentation holds a position of high importance, thus creating a model which is robust and precise is quite challenging. Polyp segmentation is a fundamental task to ensure better diagnosis and cure of colorectal cancer. As such in this study, we will see how Polyp-SAM++, a text prompt-aided SAM, can better utilize a SAM using text prompting for robust and more precise polyp segmentation. We will evaluate the performance of a text-guided SAM on the polyp segmentation task on benchmark datasets. We will also compare the results of text-guided SAM vs unprompted SAM. With this study, we hope to advance the field of polyp segmentation and inspire more, intriguing research. The code and other details will be made publically available soon at https://github.com/RisabBiswas/Polyp-SAM++.
Enhancement of the Prefiltered Rotationally Invariant Non-local PCA Algorithm for MRI
Aug 27, 2023Magnetic resonance imaging (MRI) is a non-invasive medical imaging technique offering high-resolution 3D images and valuable insights into human tissue conditions. Even at present, the refinement of denoising methods for MRI remains a crucial concern for improving the quality of the images. This study aims to enhance the prefiltered rotationally invariant non-local principal component analysis (PRI-NL-PCA) algorithm. This paper relaxed the original restriction, using the particle swarm optimization and traversal method to determine the optimal parameters of the algorithm. This paper also combined the component filters of the original algorithm and picked the most suitable combination as the new collaborative algorithm. It was found that the original algorithm has already achieved the best possible outcome, apart from a few threshold parameters that need to be adjusted. The effective way to further enhance the performance is to attach only one NL-PCA filter before and after the pre-filtered rotationally invariant non-local mean (PRI-NLM) filter. Although the performance of the new collaborative algorithm is still a little short of advanced deep learning methods, it shows that the algorithm based on PCA denoising is indeed feasible. It requires only a few parameters to be adjusted, and it is conceivable that they can be determined directly from the image, granting it a strong general capacity for various body parts, and it merits further exploration. An auxiliary tool was also extracted from the new algorithm, encouraging further combination of it with other state-of-the-art methods to further improve their denoising performance.
Self-supervised Noise2noise Method Utilizing Corrupted Images with a Modular Network for LDCT Denoising
Aug 13, 2023Deep learning is a very promising technique for low-dose computed tomography (LDCT) image denoising. However, traditional deep learning methods require paired noisy and clean datasets, which are often difficult to obtain. This paper proposes a new method for performing LDCT image denoising with only LDCT data, which means that normal-dose CT (NDCT) is not needed. We adopt a combination including the self-supervised noise2noise model and the noisy-as-clean strategy. First, we add a second yet similar type of noise to LDCT images multiple times. Note that we use LDCT images based on the noisy-as-clean strategy for corruption instead of NDCT images. Then, the noise2noise model is executed with only the secondary corrupted images for training. We select a modular U-Net structure from several candidates with shared parameters to perform the task, which increases the receptive field without increasing the parameter size. The experimental results obtained on the Mayo LDCT dataset show the effectiveness of the proposed method compared with that of state-of-the-art deep learning methods. The developed code is available at https://github.com/XYuan01/Self-supervised-Noise2Noise-for-LDCT.
Exploring Diverse In-Context Configurations for Image Captioning
May 26, 2023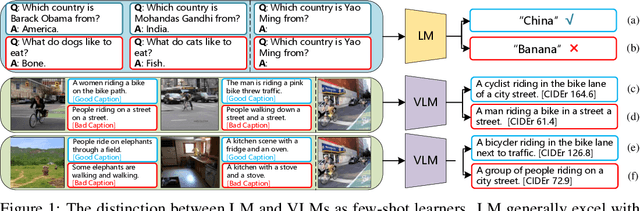
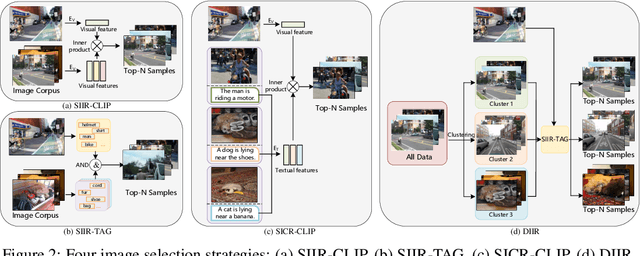
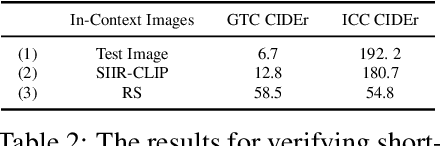
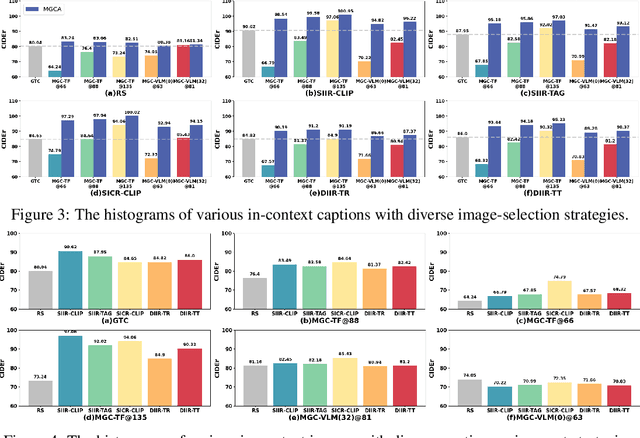
After discovering that Language Models (LMs) can be good in-context few-shot learners, numerous strategies have been proposed to optimize in-context sequence configurations. Recently, researchers in Vision-Language (VL) domains also develop their few-shot learners, while they only use the simplest way, i.e., randomly sampling, to configure in-context image-text pairs. In order to explore the effects of varying configurations on VL in-context learning, we devised four strategies for image selection and four for caption assignment to configure in-context image-text pairs for image captioning. Here Image Captioning is used as the case study since it can be seen as the visually-conditioned LM. Our comprehensive experiments yield two counter-intuitive but valuable insights, highlighting the distinct characteristics of VL in-context learning due to multi-modal synergy, as compared to the NLP case.
On the characteristics of natural hydraulic dampers: An image-based approach to study the fluid flow behaviour inside the human meniscal tissue
Jul 24, 2023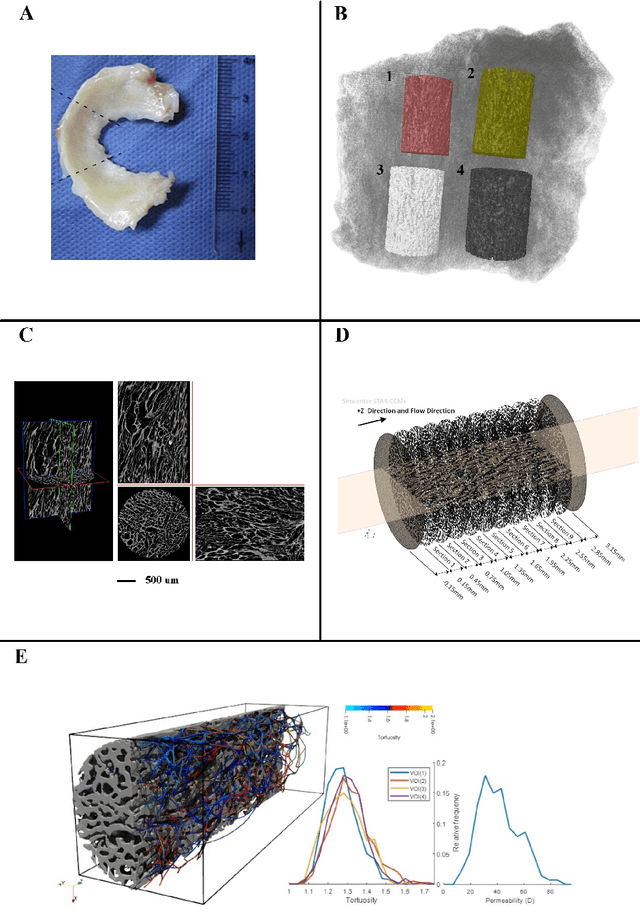
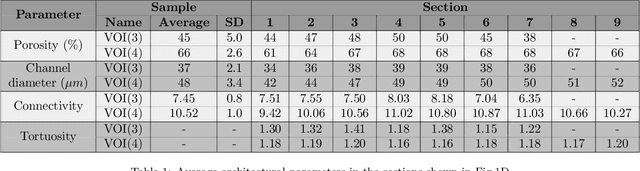
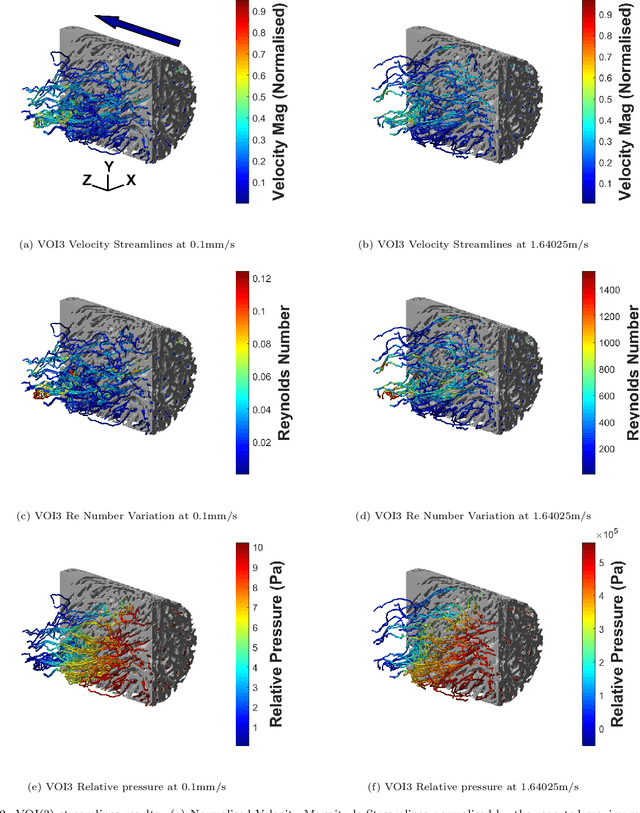
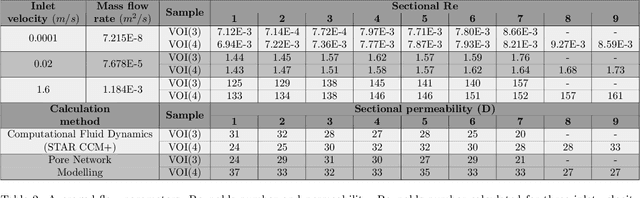
The meniscal tissue is a layered material with varying properties influenced by collagen content and arrangement. Understanding the relationship between structure and properties is crucial for disease management, treatment development, and biomaterial design. The internal layer of the meniscus is softer and more deformable than the outer layers, thanks to interconnected collagen channels that guide fluid flow. To investigate these relationships, we propose a novel approach that combines Computational Fluid Dynamics (CFD) with Image Analysis (CFD-IA). We analyze fluid flow in the internal architecture of the human meniscus across a range of inlet velocities (0.1mm/s to 1.6m/s) using high-resolution 3D micro-computed tomography scans. Statistical correlations are observed between architectural parameters (tortuosity, connectivity, porosity, pore size) and fluid flow parameters (Re number distribution, permeability). Some channels exhibit Re values of 1400 at an inlet velocity of 1.6m/s, and a transition from Darcy's regime to a non-Darcian regime occurs around an inlet velocity of 0.02m/s. Location-dependent permeability ranges from 20-32 Darcy. Regression modelling reveals a strong correlation between fluid velocity and tortuosity at high inlet velocities, as well as with channel diameter at low inlet velocities. At higher inlet velocities, flow paths deviate more from the preferential direction, resulting in a decrease in the concentration parameter by an average of 0.4. This research provides valuable insights into the fluid flow behaviour within the meniscus and its structural influences.
Putting Humans in the Image Captioning Loop
Jun 06, 2023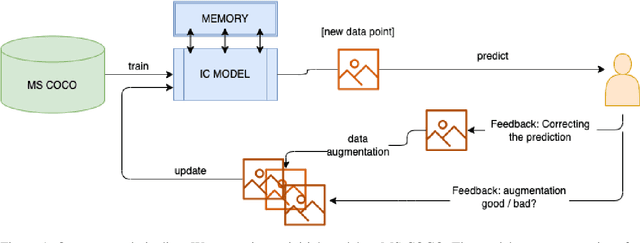
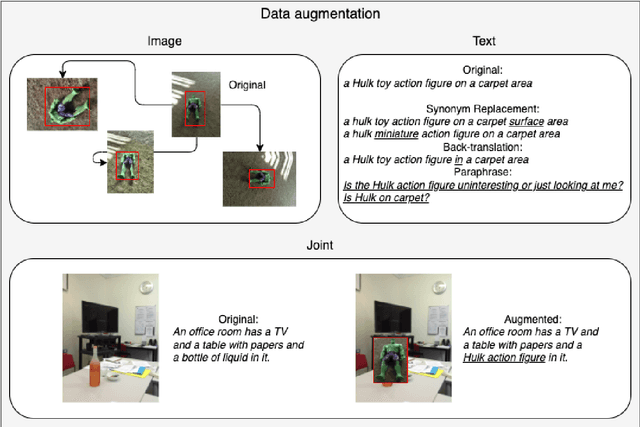
Image Captioning (IC) models can highly benefit from human feedback in the training process, especially in cases where data is limited. We present work-in-progress on adapting an IC system to integrate human feedback, with the goal to make it easily adaptable to user-specific data. Our approach builds on a base IC model pre-trained on the MS COCO dataset, which generates captions for unseen images. The user will then be able to offer feedback on the image and the generated/predicted caption, which will be augmented to create additional training instances for the adaptation of the model. The additional instances are integrated into the model using step-wise updates, and a sparse memory replay component is used to avoid catastrophic forgetting. We hope that this approach, while leading to improved results, will also result in customizable IC models.
Cross-Spatial Pixel Integration and Cross-Stage Feature Fusion Based Transformer Network for Remote Sensing Image Super-Resolution
Jul 06, 2023
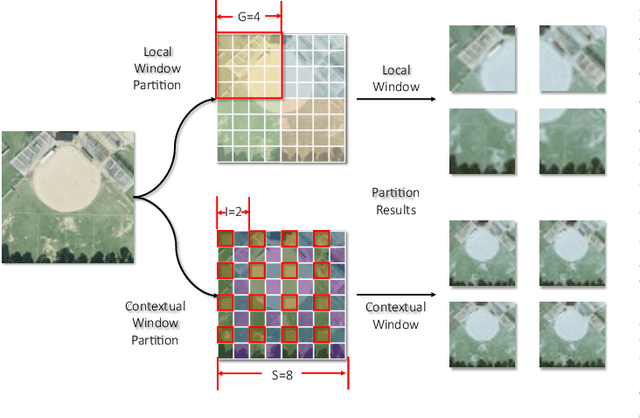
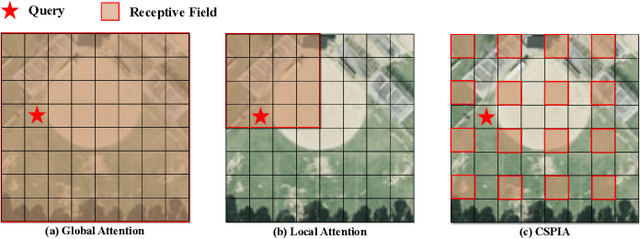
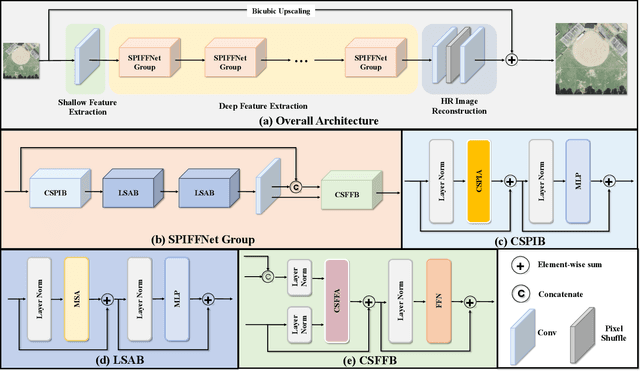
Remote sensing image super-resolution (RSISR) plays a vital role in enhancing spatial detials and improving the quality of satellite imagery. Recently, Transformer-based models have shown competitive performance in RSISR. To mitigate the quadratic computational complexity resulting from global self-attention, various methods constrain attention to a local window, enhancing its efficiency. Consequently, the receptive fields in a single attention layer are inadequate, leading to insufficient context modeling. Furthermore, while most transform-based approaches reuse shallow features through skip connections, relying solely on these connections treats shallow and deep features equally, impeding the model's ability to characterize them. To address these issues, we propose a novel transformer architecture called Cross-Spatial Pixel Integration and Cross-Stage Feature Fusion Based Transformer Network (SPIFFNet) for RSISR. Our proposed model effectively enhances global cognition and understanding of the entire image, facilitating efficient integration of features cross-stages. The model incorporates cross-spatial pixel integration attention (CSPIA) to introduce contextual information into a local window, while cross-stage feature fusion attention (CSFFA) adaptively fuses features from the previous stage to improve feature expression in line with the requirements of the current stage. We conducted comprehensive experiments on multiple benchmark datasets, demonstrating the superior performance of our proposed SPIFFNet in terms of both quantitative metrics and visual quality when compared to state-of-the-art methods.
HistoColAi: An Open-Source Web Platform for Collaborative Digital Histology Image Annotation with AI-Driven Predictive Integration
Jul 11, 2023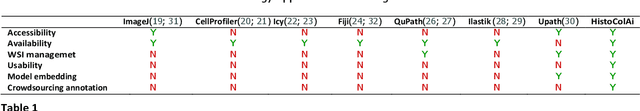
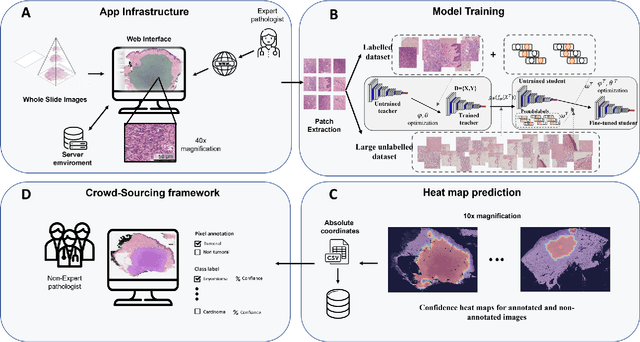
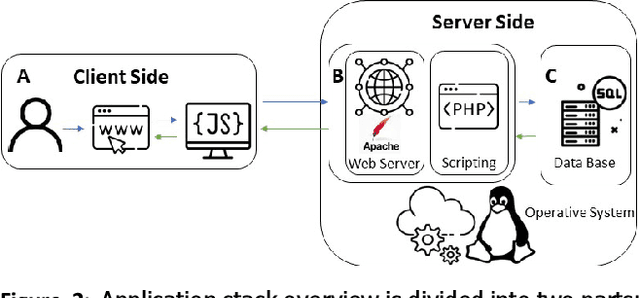
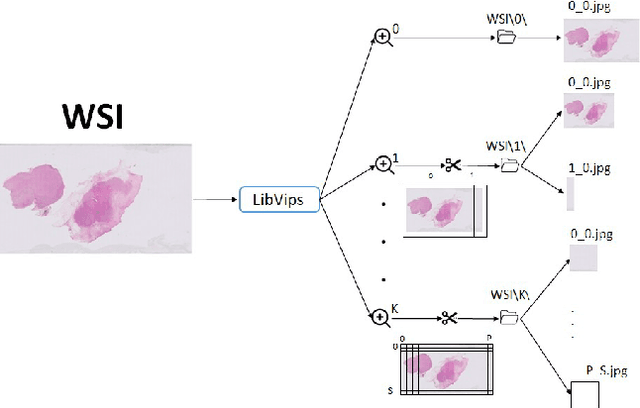
Digital pathology has become a standard in the pathology workflow due to its many benefits. These include the level of detail of the whole slide images generated and the potential immediate sharing of cases between hospitals. Recent advances in deep learning-based methods for image analysis make them of potential aid in digital pathology. However, a major limitation in developing computer-aided diagnostic systems for pathology is the lack of an intuitive and open web application for data annotation. This paper proposes a web service that efficiently provides a tool to visualize and annotate digitized histological images. In addition, to show and validate the tool, in this paper we include a use case centered on the diagnosis of spindle cell skin neoplasm for multiple annotators. A usability study of the tool is also presented, showing the feasibility of the developed tool.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge