"Image": models, code, and papers
Distributionally Robust Classification on a Data Budget
Aug 07, 2023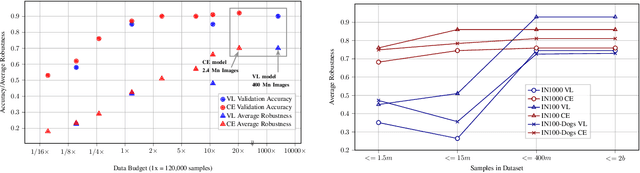
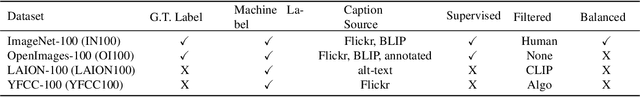


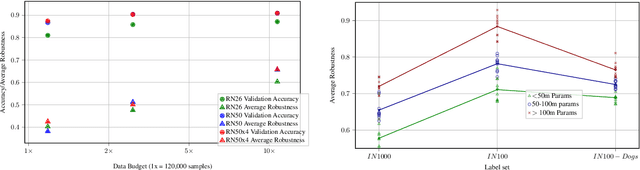
Real world uses of deep learning require predictable model behavior under distribution shifts. Models such as CLIP show emergent natural distributional robustness comparable to humans, but may require hundreds of millions of training samples. Can we train robust learners in a domain where data is limited? To rigorously address this question, we introduce JANuS (Joint Annotations and Names Set), a collection of four new training datasets with images, labels, and corresponding captions, and perform a series of carefully controlled investigations of factors contributing to robustness in image classification, then compare those results to findings derived from a large-scale meta-analysis. Using this approach, we show that standard ResNet-50 trained with the cross-entropy loss on 2.4 million image samples can attain comparable robustness to a CLIP ResNet-50 trained on 400 million samples. To our knowledge, this is the first result showing (near) state-of-the-art distributional robustness on limited data budgets. Our dataset is available at \url{https://huggingface.co/datasets/penfever/JANuS_dataset}, and the code used to reproduce our experiments can be found at \url{https://github.com/penfever/vlhub/}.
PUG: Photorealistic and Semantically Controllable Synthetic Data for Representation Learning
Aug 08, 2023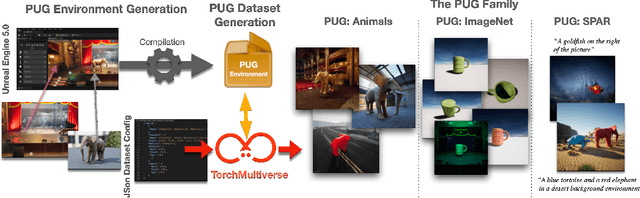
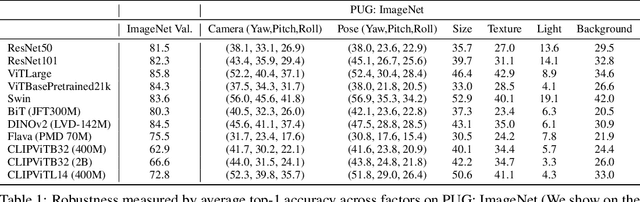

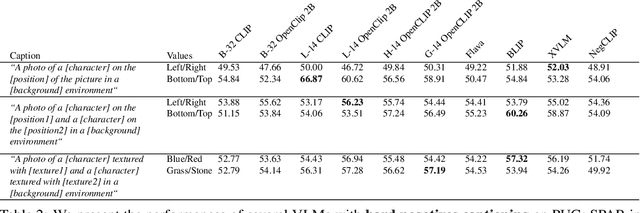
Synthetic image datasets offer unmatched advantages for designing and evaluating deep neural networks: they make it possible to (i) render as many data samples as needed, (ii) precisely control each scene and yield granular ground truth labels (and captions), (iii) precisely control distribution shifts between training and testing to isolate variables of interest for sound experimentation. Despite such promise, the use of synthetic image data is still limited -- and often played down -- mainly due to their lack of realism. Most works therefore rely on datasets of real images, which have often been scraped from public images on the internet, and may have issues with regards to privacy, bias, and copyright, while offering little control over how objects precisely appear. In this work, we present a path to democratize the use of photorealistic synthetic data: we develop a new generation of interactive environments for representation learning research, that offer both controllability and realism. We use the Unreal Engine, a powerful game engine well known in the entertainment industry, to produce PUG (Photorealistic Unreal Graphics) environments and datasets for representation learning. In this paper, we demonstrate the potential of PUG to enable more rigorous evaluations of vision models.
Bridging the Gap: Exploring the Capabilities of Bridge-Architectures for Complex Visual Reasoning Tasks
Jul 31, 2023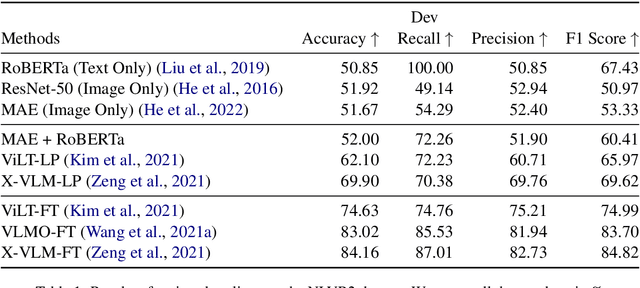
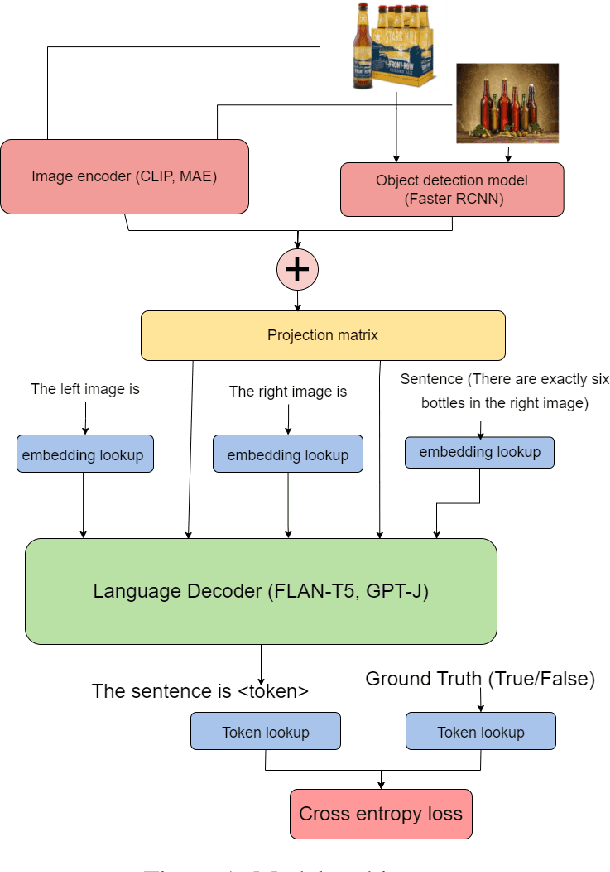
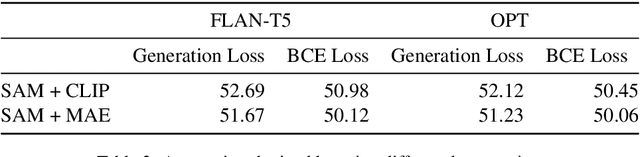
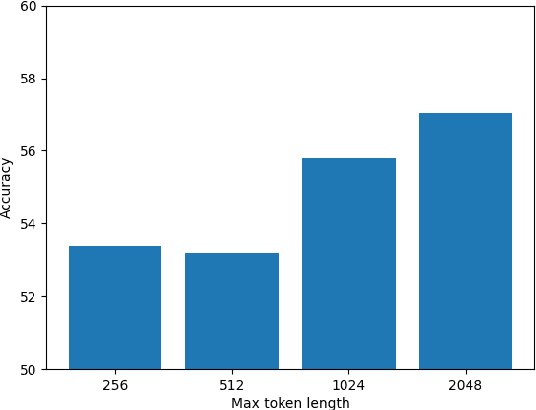
In recent times there has been a surge of multi-modal architectures based on Large Language Models, which leverage the zero shot generation capabilities of LLMs and project image embeddings into the text space and then use the auto-regressive capacity to solve tasks such as VQA, captioning, and image retrieval. We name these architectures as "bridge-architectures" as they project from the image space to the text space. These models deviate from the traditional recipe of training transformer based multi-modal models, which involve using large-scale pre-training and complex multi-modal interactions through co or cross attention. However, the capabilities of bridge architectures have not been tested on complex visual reasoning tasks which require fine grained analysis about the image. In this project, we investigate the performance of these bridge-architectures on the NLVR2 dataset, and compare it to state-of-the-art transformer based architectures. We first extend the traditional bridge architectures for the NLVR2 dataset, by adding object level features to faciliate fine-grained object reasoning. Our analysis shows that adding object level features to bridge architectures does not help, and that pre-training on multi-modal data is key for good performance on complex reasoning tasks such as NLVR2. We also demonstrate some initial results on a recently bridge-architecture, LLaVA, in the zero shot setting and analyze its performance.
A Deep Learning Approach for Virtual Contrast Enhancement in Contrast Enhanced Spectral Mammography
Aug 03, 2023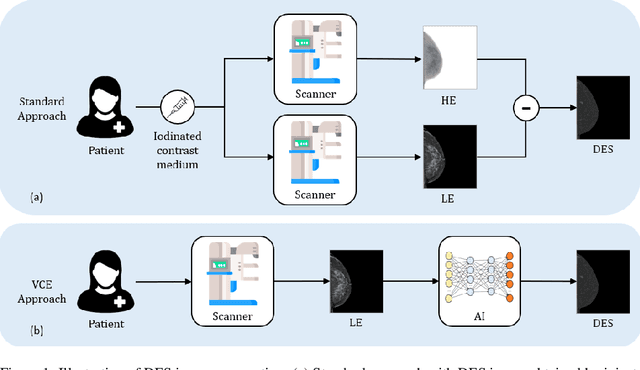
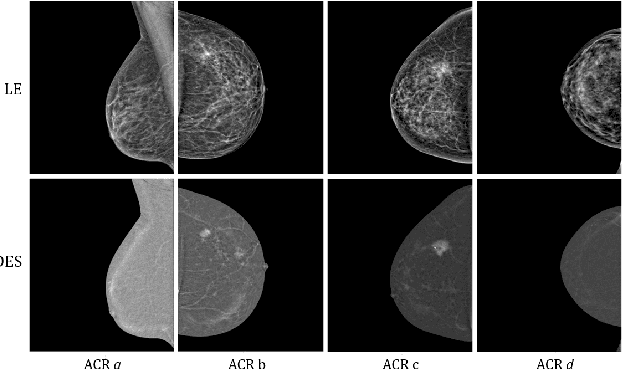
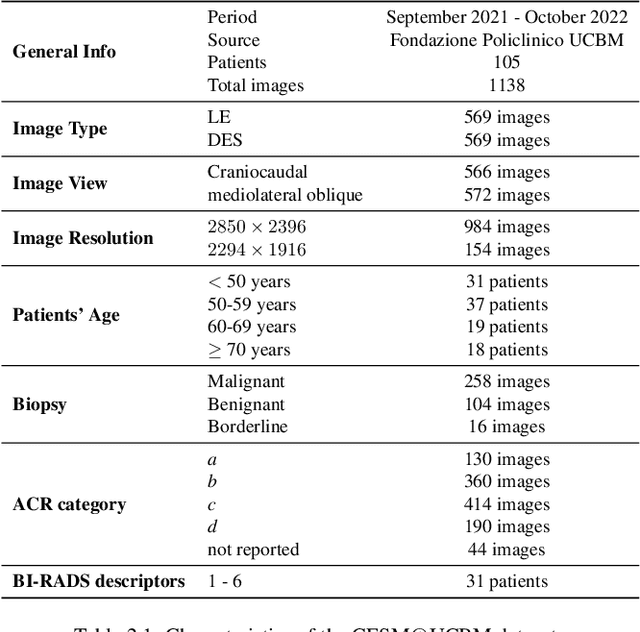
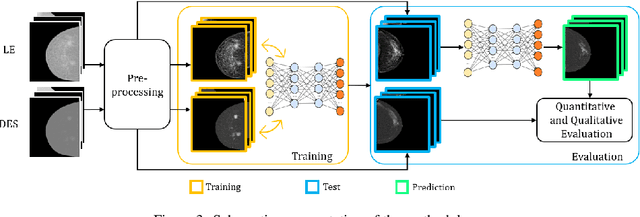
Contrast Enhanced Spectral Mammography (CESM) is a dual-energy mammographic imaging technique that first needs intravenously administration of an iodinated contrast medium; then, it collects both a low-energy image, comparable to standard mammography, and a high-energy image. The two scans are then combined to get a recombined image showing contrast enhancement. Despite CESM diagnostic advantages for breast cancer diagnosis, the use of contrast medium can cause side effects, and CESM also beams patients with a higher radiation dose compared to standard mammography. To address these limitations this work proposes to use deep generative models for virtual contrast enhancement on CESM, aiming to make the CESM contrast-free as well as to reduce the radiation dose. Our deep networks, consisting of an autoencoder and two Generative Adversarial Networks, the Pix2Pix, and the CycleGAN, generate synthetic recombined images solely from low-energy images. We perform an extensive quantitative and qualitative analysis of the model's performance, also exploiting radiologists' assessments, on a novel CESM dataset that includes 1138 images that, as a further contribution of this work, we make publicly available. The results show that CycleGAN is the most promising deep network to generate synthetic recombined images, highlighting the potential of artificial intelligence techniques for virtual contrast enhancement in this field.
CartiMorph: a framework for automated knee articular cartilage morphometrics
Aug 03, 2023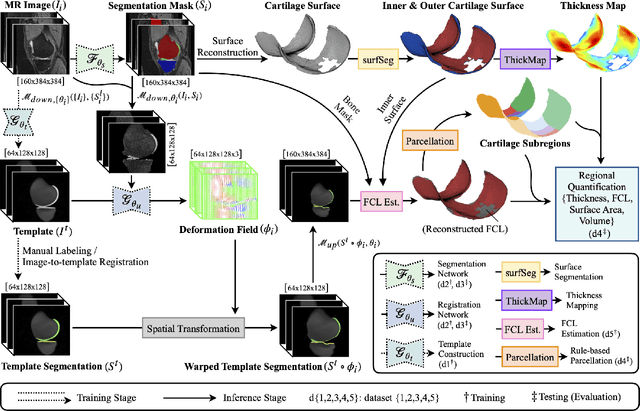
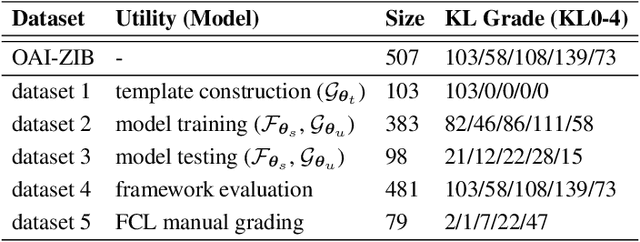
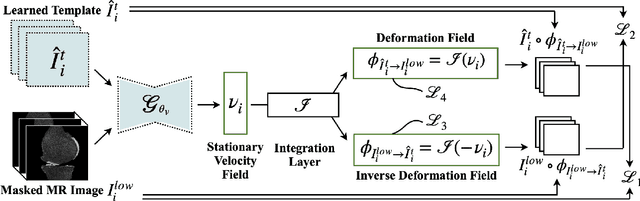
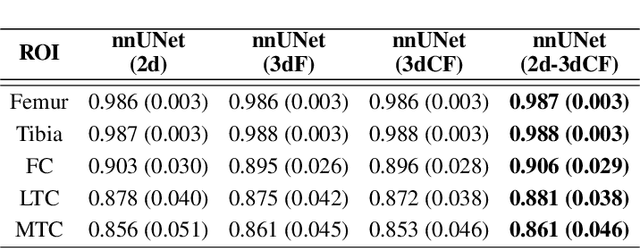
We introduce CartiMorph, a framework for automated knee articular cartilage morphometrics. It takes an image as input and generates quantitative metrics for cartilage subregions, including the percentage of full-thickness cartilage loss (FCL), mean thickness, surface area, and volume. CartiMorph leverages the power of deep learning models for hierarchical image feature representation. Deep learning models were trained and validated for tissue segmentation, template construction, and template-to-image registration. We established methods for surface-normal-based cartilage thickness mapping, FCL estimation, and rule-based cartilage parcellation. Our cartilage thickness map showed less error in thin and peripheral regions. We evaluated the effectiveness of the adopted segmentation model by comparing the quantitative metrics obtained from model segmentation and those from manual segmentation. The root-mean-squared deviation of the FCL measurements was less than 8%, and strong correlations were observed for the mean thickness (Pearson's correlation coefficient $\rho \in [0.82,0.97]$), surface area ($\rho \in [0.82,0.98]$) and volume ($\rho \in [0.89,0.98]$) measurements. We compared our FCL measurements with those from a previous study and found that our measurements deviated less from the ground truths. We observed superior performance of the proposed rule-based cartilage parcellation method compared with the atlas-based approach. CartiMorph has the potential to promote imaging biomarkers discovery for knee osteoarthritis.
A Visual Quality Assessment Method for Raster Images in Scanned Document
Jul 25, 2023
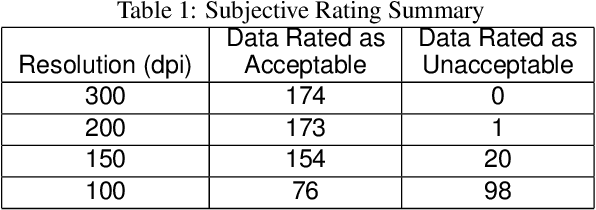
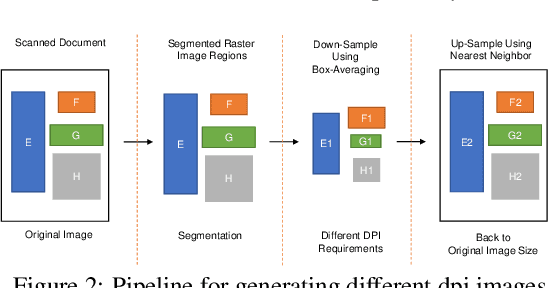
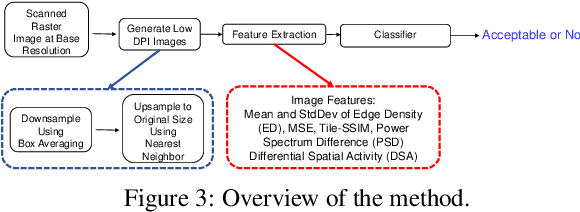
Image quality assessment (IQA) is an active research area in the field of image processing. Most prior works focus on visual quality of natural images captured by cameras. In this paper, we explore visual quality of scanned documents, focusing on raster image areas. Different from many existing works which aim to estimate a visual quality score, we propose a machine learning based classification method to determine whether the visual quality of a scanned raster image at a given resolution setting is acceptable. We conduct a psychophysical study to determine the acceptability at different image resolutions based on human subject ratings and use them as the ground truth to train our machine learning model. However, this dataset is unbalanced as most images were rated as visually acceptable. To address the data imbalance problem, we introduce several noise models to simulate the degradation of image quality during the scanning process. Our results show that by including augmented data in training, we can significantly improve the performance of the classifier to determine whether the visual quality of raster images in a scanned document is acceptable or not for a given resolution setting.
Transformer-based Annotation Bias-aware Medical Image Segmentation
Jun 02, 2023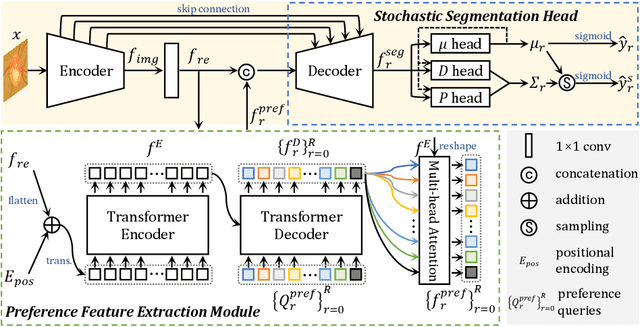
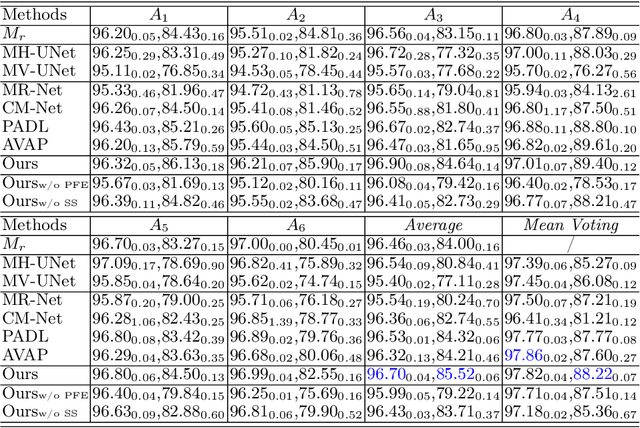
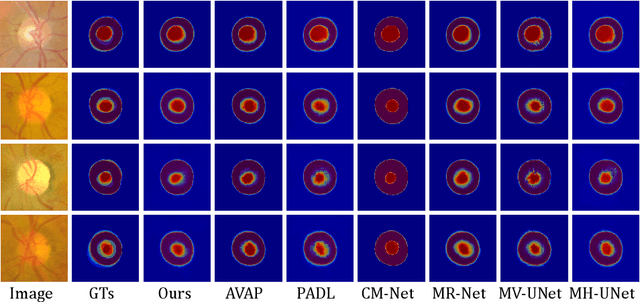
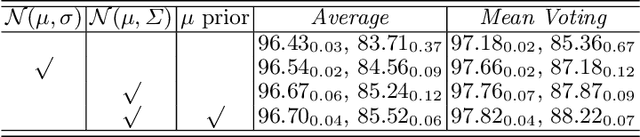
Manual medical image segmentation is subjective and suffers from annotator-related bias, which can be mimicked or amplified by deep learning methods. Recently, researchers have suggested that such bias is the combination of the annotator preference and stochastic error, which are modeled by convolution blocks located after decoder and pixel-wise independent Gaussian distribution, respectively. It is unlikely that convolution blocks can effectively model the varying degrees of preference at the full resolution level. Additionally, the independent pixel-wise Gaussian distribution disregards pixel correlations, leading to a discontinuous boundary. This paper proposes a Transformer-based Annotation Bias-aware (TAB) medical image segmentation model, which tackles the annotator-related bias via modeling annotator preference and stochastic errors. TAB employs the Transformer with learnable queries to extract the different preference-focused features. This enables TAB to produce segmentation with various preferences simultaneously using a single segmentation head. Moreover, TAB takes the multivariant normal distribution assumption that models pixel correlations, and learns the annotation distribution to disentangle the stochastic error. We evaluated our TAB on an OD/OC segmentation benchmark annotated by six annotators. Our results suggest that TAB outperforms existing medical image segmentation models which take into account the annotator-related bias.
A Novel Cross-Perturbation for Single Domain Generalization
Aug 02, 2023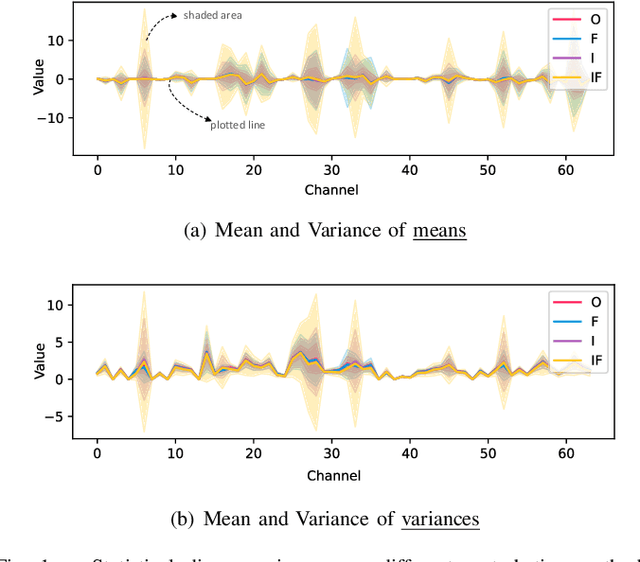
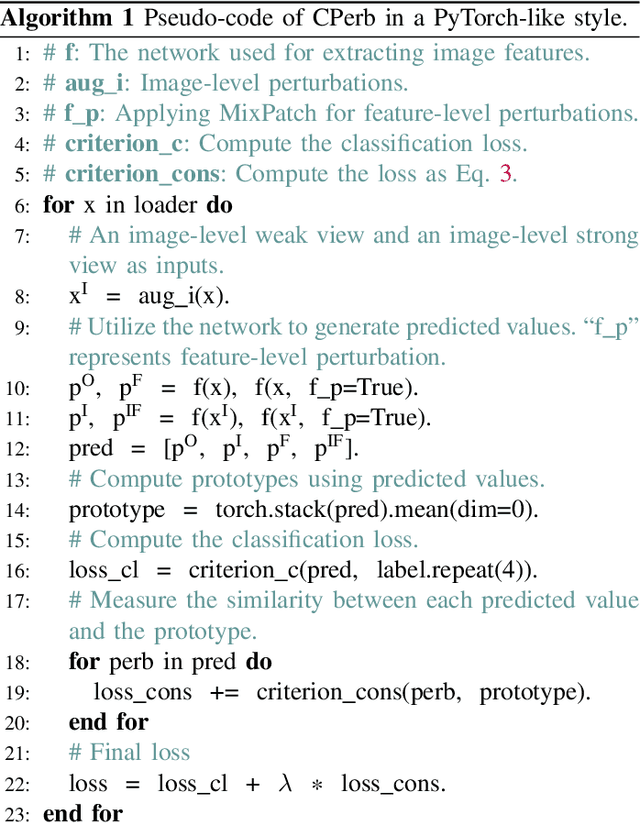
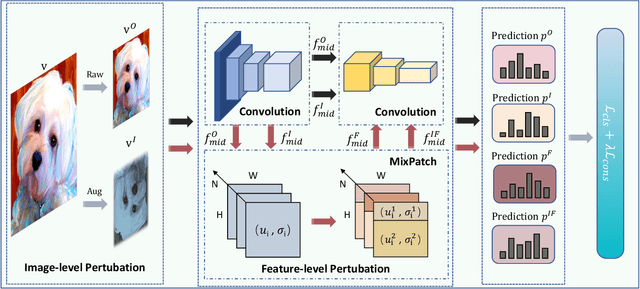
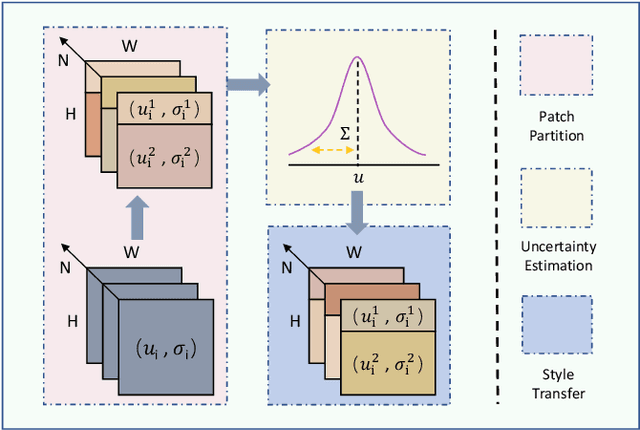

Single domain generalization aims to enhance the ability of the model to generalize to unknown domains when trained on a single source domain. However, the limited diversity in the training data hampers the learning of domain-invariant features, resulting in compromised generalization performance. To address this, data perturbation (augmentation) has emerged as a crucial method to increase data diversity. Nevertheless, existing perturbation methods often focus on either image-level or feature-level perturbations independently, neglecting their synergistic effects. To overcome these limitations, we propose CPerb, a simple yet effective cross-perturbation method. Specifically, CPerb utilizes both horizontal and vertical operations. Horizontally, it applies image-level and feature-level perturbations to enhance the diversity of the training data, mitigating the issue of limited diversity in single-source domains. Vertically, it introduces multi-route perturbation to learn domain-invariant features from different perspectives of samples with the same semantic category, thereby enhancing the generalization capability of the model. Additionally, we propose MixPatch, a novel feature-level perturbation method that exploits local image style information to further diversify the training data. Extensive experiments on various benchmark datasets validate the effectiveness of our method.
AutoLTS: Automating Cycling Stress Assessment via Contrastive Learning and Spatial Post-processing
Aug 15, 2023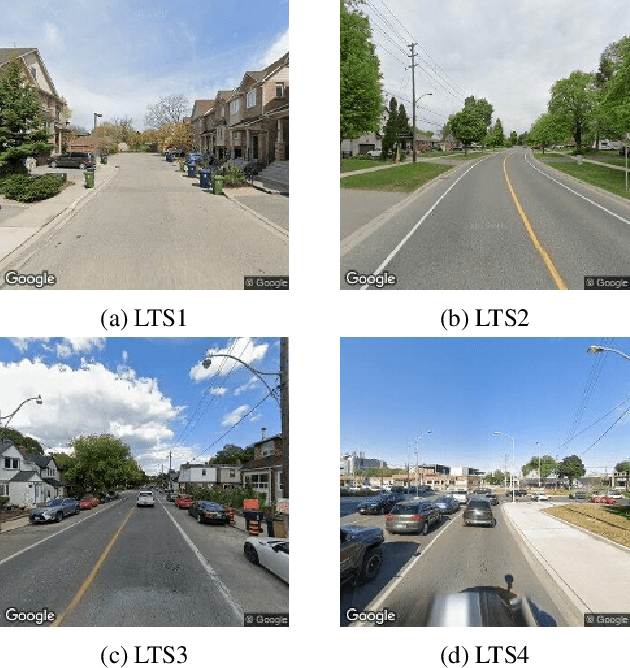
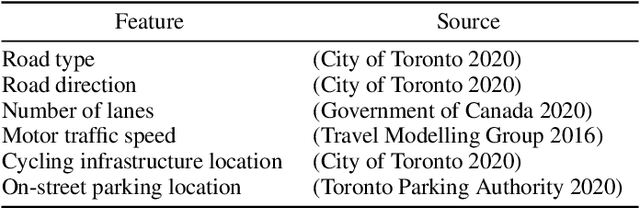

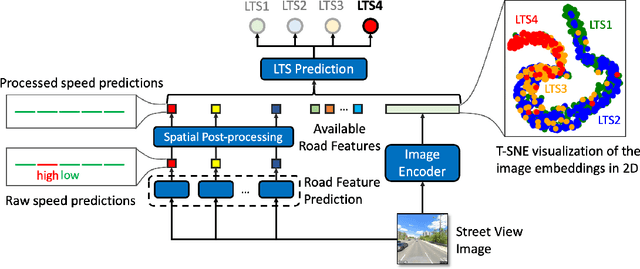
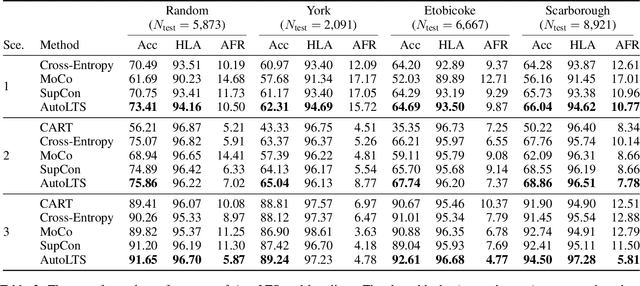
Cycling stress assessment, which quantifies cyclists' perceived stress imposed by the built environment and motor traffics, increasingly informs cycling infrastructure planning and cycling route recommendation. However, currently calculating cycling stress is slow and data-intensive, which hinders its broader application. In this paper, We propose a deep learning framework to support accurate, fast, and large-scale cycling stress assessments for urban road networks based on street-view images. Our framework features i) a contrastive learning approach that leverages the ordinal relationship among cycling stress labels, and ii) a post-processing technique that enforces spatial smoothness into our predictions. On a dataset of 39,153 road segments collected in Toronto, Canada, our results demonstrate the effectiveness of our deep learning framework and the value of using image data for cycling stress assessment in the absence of high-quality road geometry and motor traffic data.
Classification of White Blood Cells Using Machine and Deep Learning Models: A Systematic Review
Aug 21, 2023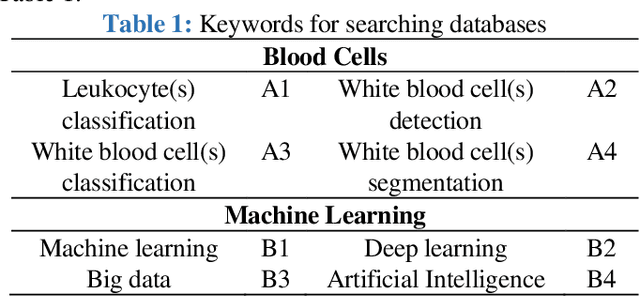
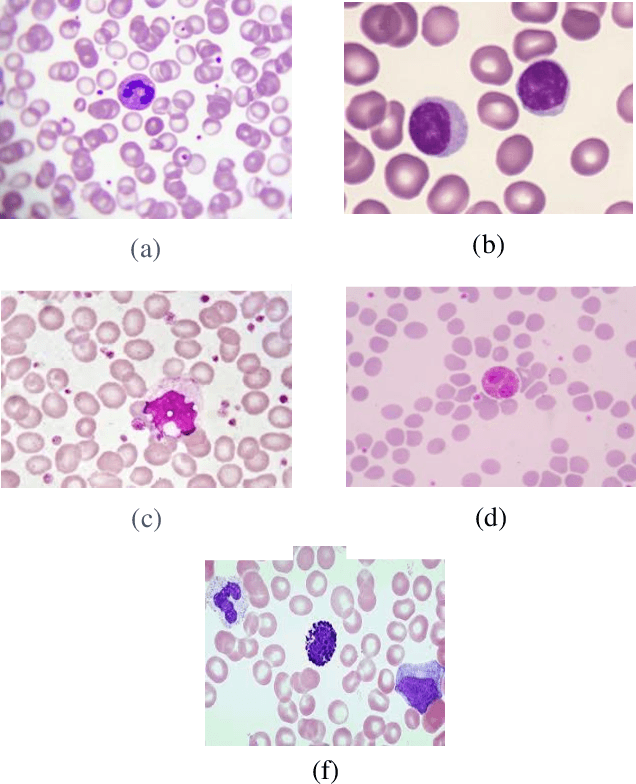

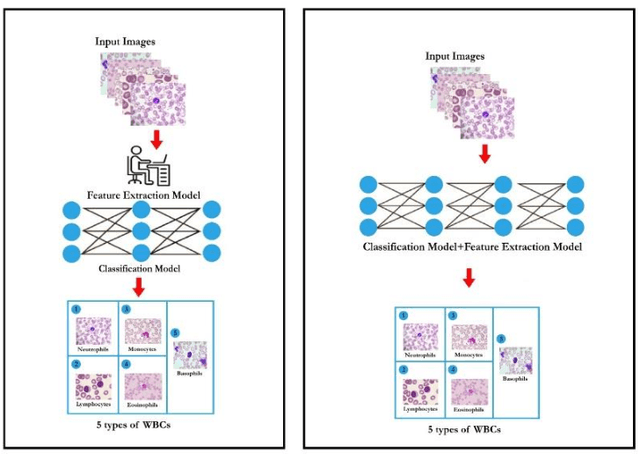
Machine learning (ML) and deep learning (DL) models have been employed to significantly improve analyses of medical imagery, with these approaches used to enhance the accuracy of prediction and classification. Model predictions and classifications assist diagnoses of various cancers and tumors. This review presents an in-depth analysis of modern techniques applied within the domain of medical image analysis for white blood cell classification. The methodologies that use blood smear images, magnetic resonance imaging (MRI), X-rays, and similar medical imaging domains are identified and discussed, with a detailed analysis of ML/DL techniques applied to the classification of white blood cells (WBCs) representing the primary focus of the review. The data utilized in this research has been extracted from a collection of 136 primary papers that were published between the years 2006 and 2023. The most widely used techniques and best-performing white blood cell classification methods are identified. While the use of ML and DL for white blood cell classification has concurrently increased and improved in recent year, significant challenges remain - 1) Availability of appropriate datasets remain the primary challenge, and may be resolved using data augmentation techniques. 2) Medical training of researchers is recommended to improve current understanding of white blood cell structure and subsequent selection of appropriate classification models. 3) Advanced DL networks including Generative Adversarial Networks, R-CNN, Fast R-CNN, and faster R-CNN will likely be increasingly employed to supplement or replace current techniques.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge