"Image": models, code, and papers
Nested Diffusion Processes for Anytime Image Generation
May 30, 2023
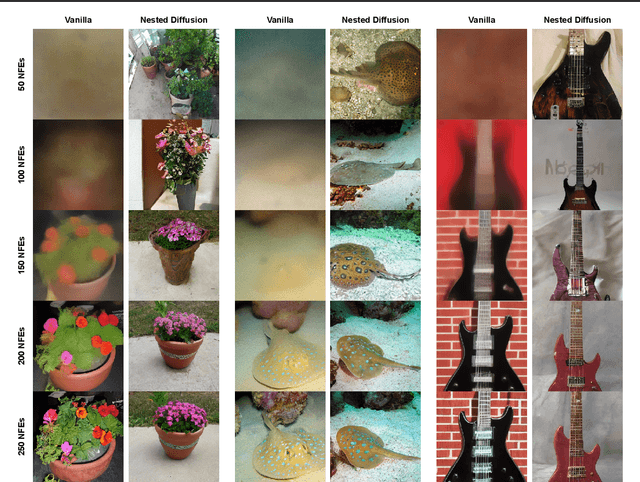
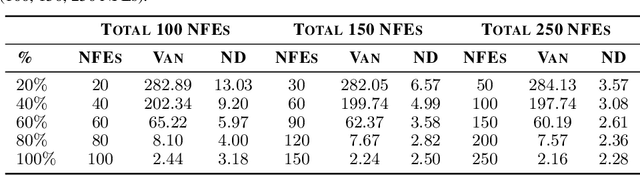
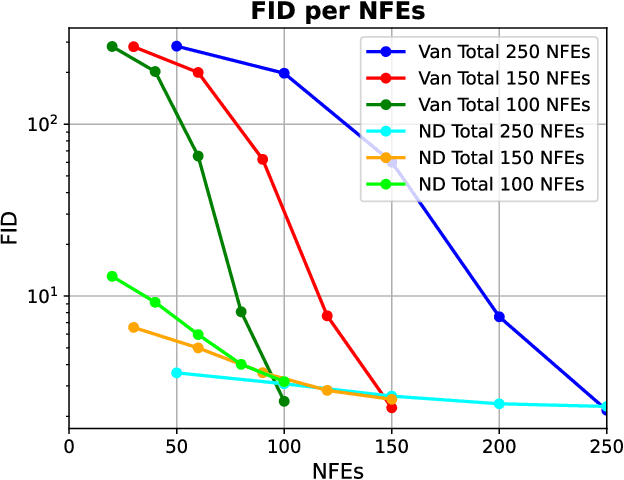

Diffusion models are the current state-of-the-art in image generation, synthesizing high-quality images by breaking down the generation process into many fine-grained denoising steps. Despite their good performance, diffusion models are computationally expensive, requiring many neural function evaluations (NFEs). In this work, we propose an anytime diffusion-based method that can generate viable images when stopped at arbitrary times before completion. Using existing pretrained diffusion models, we show that the generation scheme can be recomposed as two nested diffusion processes, enabling fast iterative refinement of a generated image. We use this Nested Diffusion approach to peek into the generation process and enable flexible scheduling based on the instantaneous preference of the user. In experiments on ImageNet and Stable Diffusion-based text-to-image generation, we show, both qualitatively and quantitatively, that our method's intermediate generation quality greatly exceeds that of the original diffusion model, while the final slow generation result remains comparable.
Pseudo Flow Consistency for Self-Supervised 6D Object Pose Estimation
Aug 19, 2023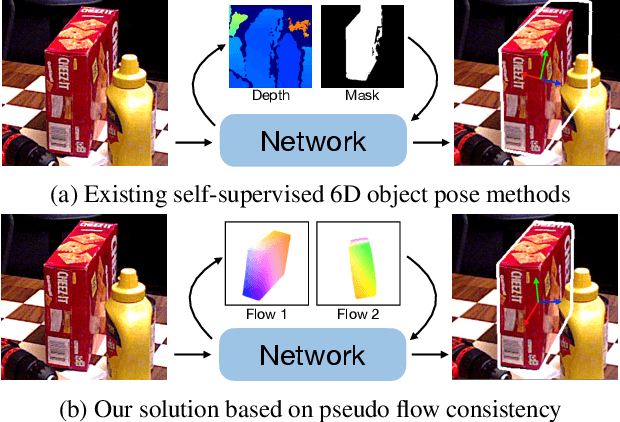
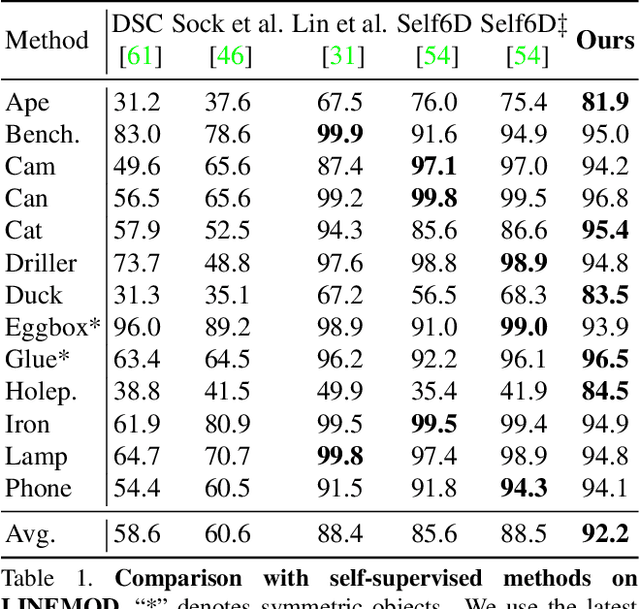
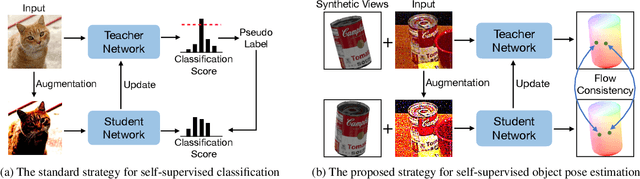
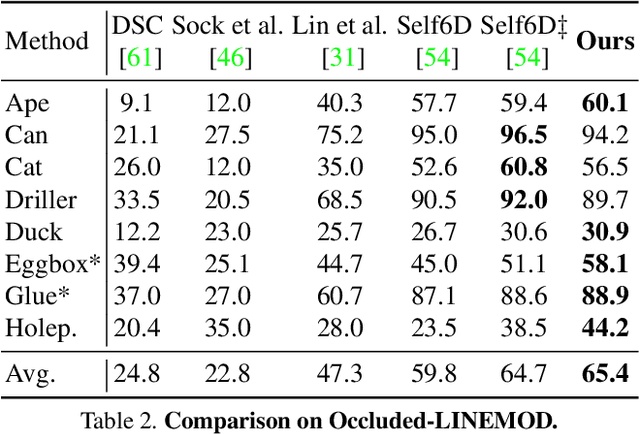
Most self-supervised 6D object pose estimation methods can only work with additional depth information or rely on the accurate annotation of 2D segmentation masks, limiting their application range. In this paper, we propose a 6D object pose estimation method that can be trained with pure RGB images without any auxiliary information. We first obtain a rough pose initialization from networks trained on synthetic images rendered from the target's 3D mesh. Then, we introduce a refinement strategy leveraging the geometry constraint in synthetic-to-real image pairs from multiple different views. We formulate this geometry constraint as pixel-level flow consistency between the training images with dynamically generated pseudo labels. We evaluate our method on three challenging datasets and demonstrate that it outperforms state-of-the-art self-supervised methods significantly, with neither 2D annotations nor additional depth images.
Generative Approach for Probabilistic Human Mesh Recovery using Diffusion Models
Aug 05, 2023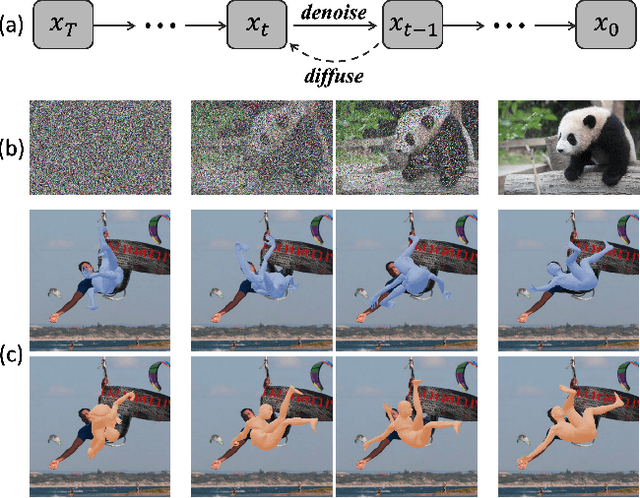
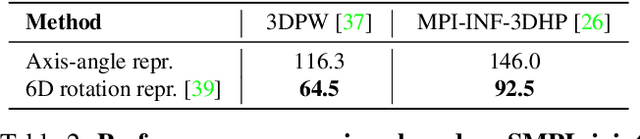
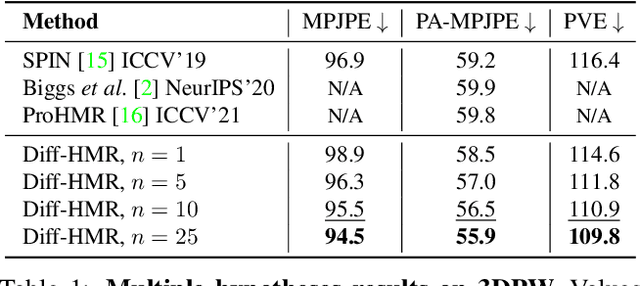
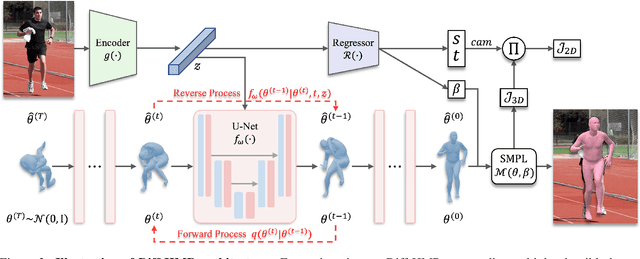

This work focuses on the problem of reconstructing a 3D human body mesh from a given 2D image. Despite the inherent ambiguity of the task of human mesh recovery, most existing works have adopted a method of regressing a single output. In contrast, we propose a generative approach framework, called "Diffusion-based Human Mesh Recovery (Diff-HMR)" that takes advantage of the denoising diffusion process to account for multiple plausible outcomes. During the training phase, the SMPL parameters are diffused from ground-truth parameters to random distribution, and Diff-HMR learns the reverse process of this diffusion. In the inference phase, the model progressively refines the given random SMPL parameters into the corresponding parameters that align with the input image. Diff-HMR, being a generative approach, is capable of generating diverse results for the same input image as the input noise varies. We conduct validation experiments, and the results demonstrate that the proposed framework effectively models the inherent ambiguity of the task of human mesh recovery in a probabilistic manner. The code is available at https://github.com/hanbyel0105/Diff-HMR
MedAugment: Universal Automatic Data Augmentation Plug-in for Medical Image Analysis
Jun 30, 2023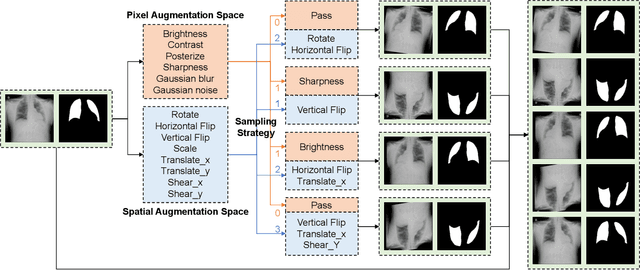
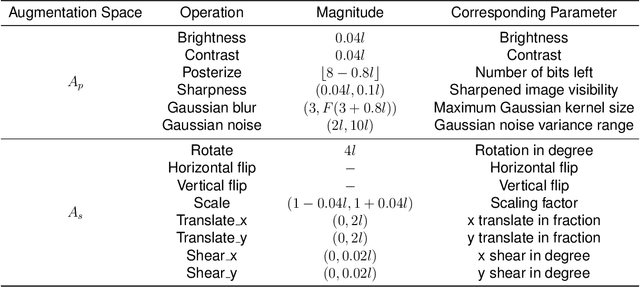
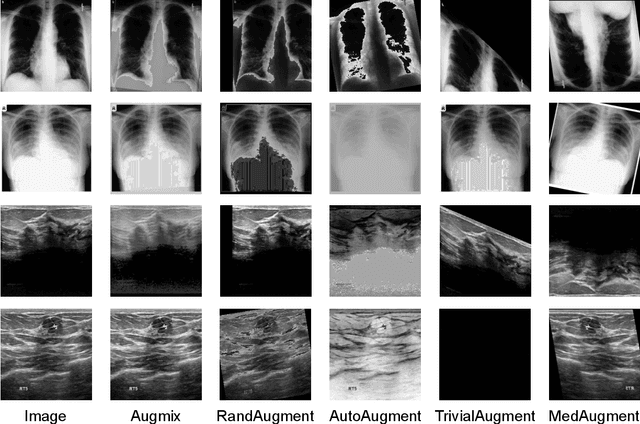
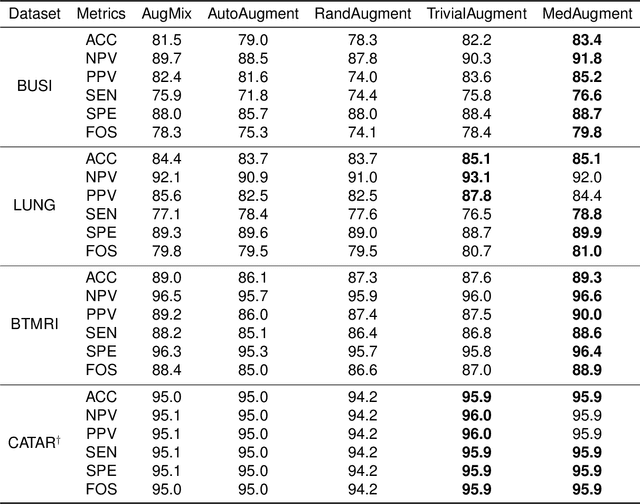
Data Augmentation (DA) technique has been widely implemented in the computer vision field to relieve the data shortage, while the DA in Medical Image Analysis (MIA) is still mostly experience-driven. Here, we develop a plug-and-use DA method, named MedAugment, to introduce the automatic DA argumentation to the MIA field. To settle the difference between natural images and medical images, we divide the augmentation space into pixel augmentation space and spatial augmentation space. A novel operation sampling strategy is also proposed when sampling DA operations from the spaces. To demonstrate the performance and universality of MedAugment, we implement extensive experiments on four classification datasets and three segmentation datasets. The results show that our MedAugment outperforms most state-of-the-art DA methods. This work shows that the plug-and-use MedAugment may benefit the MIA community. Code is available at https://github.com/NUS-Tim/MedAugment_Pytorch.
Single-Image-Based Deep Learning for Segmentation of Early Esophageal Cancer Lesions
Jun 09, 2023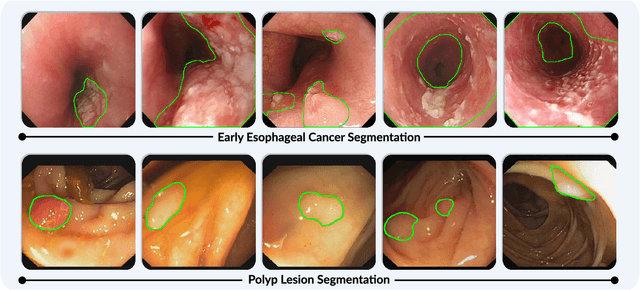
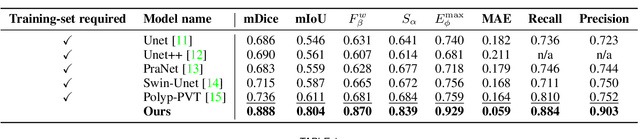
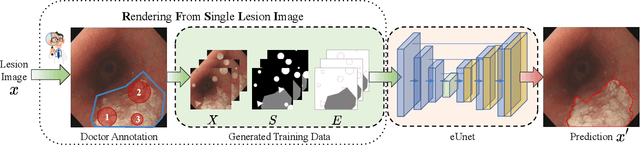
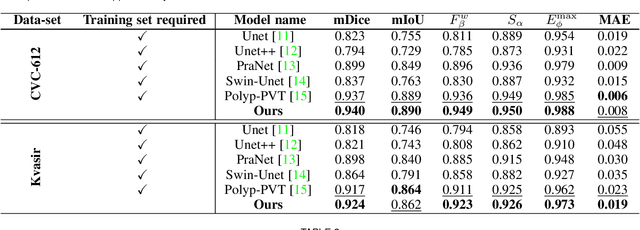
Accurate segmentation of lesions is crucial for diagnosis and treatment of early esophageal cancer (EEC). However, neither traditional nor deep learning-based methods up to today can meet the clinical requirements, with the mean Dice score - the most important metric in medical image analysis - hardly exceeding 0.75. In this paper, we present a novel deep learning approach for segmenting EEC lesions. Our approach stands out for its uniqueness, as it relies solely on a single image coming from one patient, forming the so-called "You-Only-Have-One" (YOHO) framework. On one hand, this "one-image-one-network" learning ensures complete patient privacy as it does not use any images from other patients as the training data. On the other hand, it avoids nearly all generalization-related problems since each trained network is applied only to the input image itself. In particular, we can push the training to "over-fitting" as much as possible to increase the segmentation accuracy. Our technical details include an interaction with clinical physicians to utilize their expertise, a geometry-based rendering of a single lesion image to generate the training set (the \emph{biggest} novelty), and an edge-enhanced UNet. We have evaluated YOHO over an EEC data-set created by ourselves and achieved a mean Dice score of 0.888, which represents a significant advance toward clinical applications.
A Mini-Batch Quasi-Newton Proximal Method for Constrained Total-Variation Nonlinear Image Reconstruction
Jul 05, 2023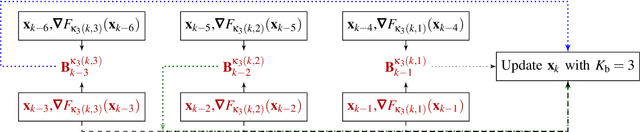
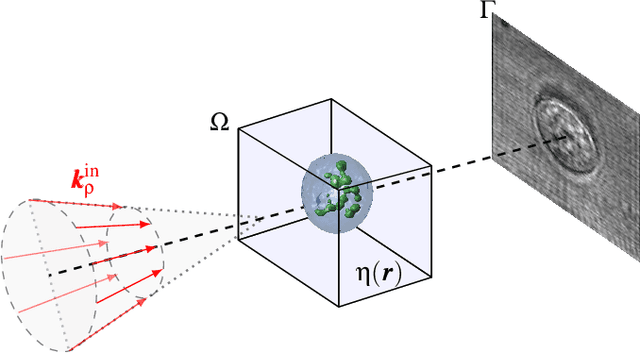
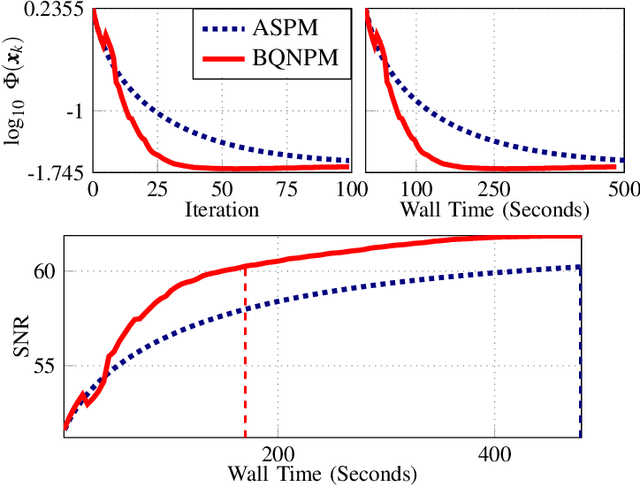
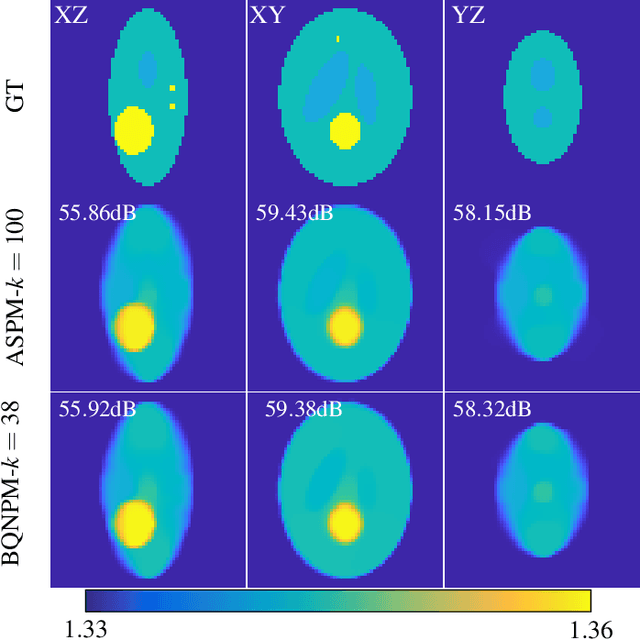
Over the years, computational imaging with accurate nonlinear physical models has drawn considerable interest due to its ability to achieve high-quality reconstructions. However, such nonlinear models are computationally demanding. A popular choice for solving the corresponding inverse problems is accelerated stochastic proximal methods (ASPMs), with the caveat that each iteration is expensive. To overcome this issue, we propose a mini-batch quasi-Newton proximal method (BQNPM) tailored to image-reconstruction problems with total-variation regularization. It involves an efficient approach that computes a weighted proximal mapping at a cost similar to that of the proximal mapping in ASPMs. However, BQNPM requires fewer iterations than ASPMs to converge. We assess the performance of BQNPM on three-dimensional inverse-scattering problems with linear and nonlinear physical models. Our results on simulated and real data show the effectiveness and efficiency of BQNPM,
Classification Committee for Active Deep Object Detection
Aug 16, 2023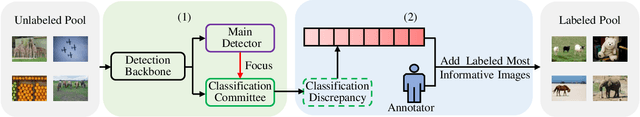
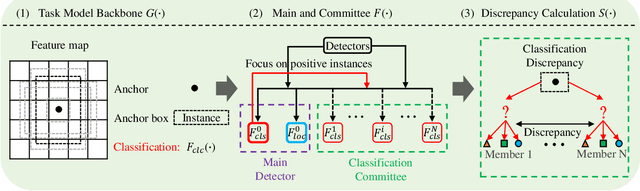
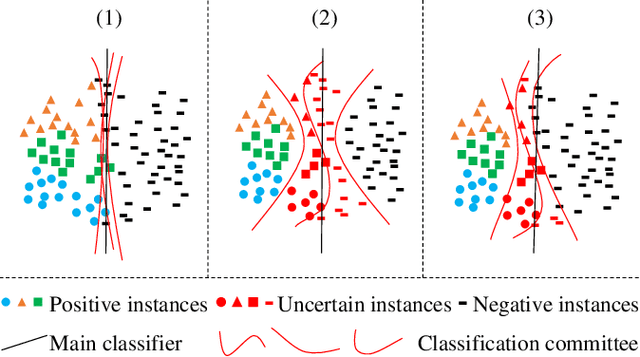
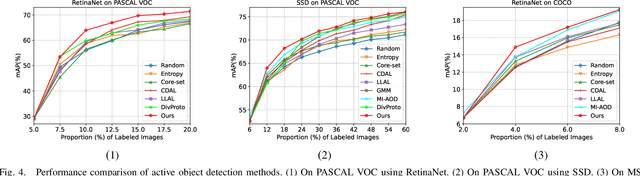
In object detection, the cost of labeling is much high because it needs not only to confirm the categories of multiple objects in an image but also to accurately determine the bounding boxes of each object. Thus, integrating active learning into object detection will raise pretty positive significance. In this paper, we propose a classification committee for active deep object detection method by introducing a discrepancy mechanism of multiple classifiers for samples' selection when training object detectors. The model contains a main detector and a classification committee. The main detector denotes the target object detector trained from a labeled pool composed of the selected informative images. The role of the classification committee is to select the most informative images according to their uncertainty values from the view of classification, which is expected to focus more on the discrepancy and representative of instances. Specifically, they compute the uncertainty for a specified instance within the image by measuring its discrepancy output by the committee pre-trained via the proposed Maximum Classifiers Discrepancy Group Loss (MCDGL). The most informative images are finally determined by selecting the ones with many high-uncertainty instances. Besides, to mitigate the impact of interference instances, we design a Focus on Positive Instances Loss (FPIL) to make the committee the ability to automatically focus on the representative instances as well as precisely encode their discrepancies for the same instance. Experiments are conducted on Pascal VOC and COCO datasets versus some popular object detectors. And results show that our method outperforms the state-of-the-art active learning methods, which verifies the effectiveness of the proposed method.
TransTIC: Transferring Transformer-based Image Compression from Human Visualization to Machine Perception
Jun 08, 2023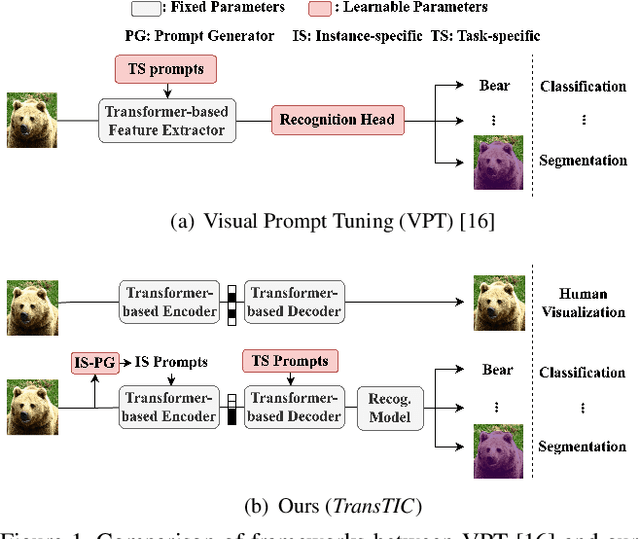
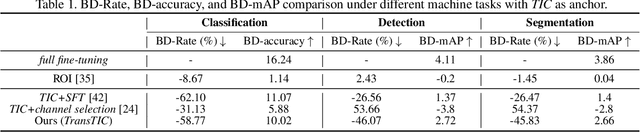
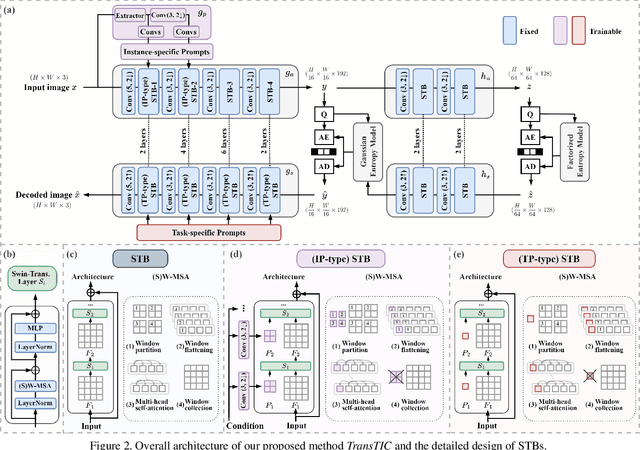
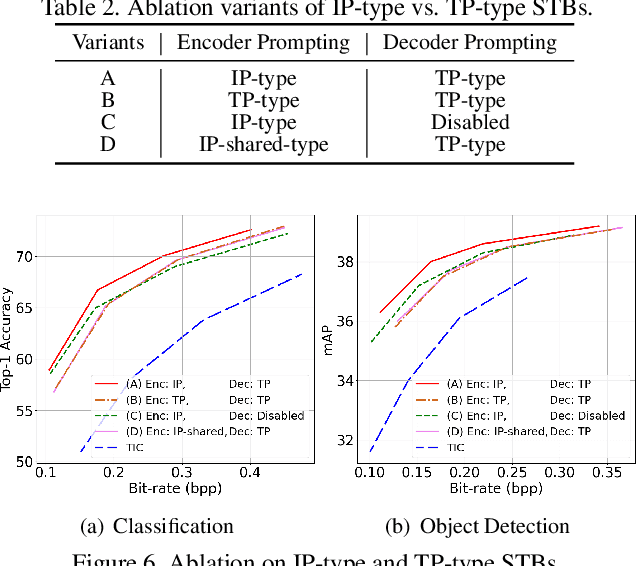
This work aims for transferring a Transformer-based image compression codec from human vision to machine perception without fine-tuning the codec. We propose a transferable Transformer-based image compression framework, termed TransTIC. Inspired by visual prompt tuning, we propose an instance-specific prompt generator to inject instance-specific prompts to the encoder and task-specific prompts to the decoder. Extensive experiments show that our proposed method is capable of transferring the codec to various machine tasks and outshining the competing methods significantly. To our best knowledge, this work is the first attempt to utilize prompting on the low-level image compression task.
Rethinking Exemplars for Continual Semantic Segmentation in Endoscopy Scenes: Entropy-based Mini-Batch Pseudo-Replay
Aug 27, 2023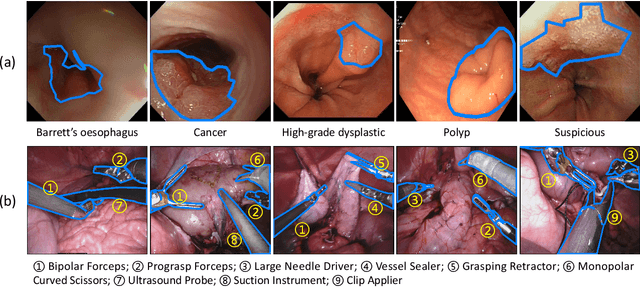
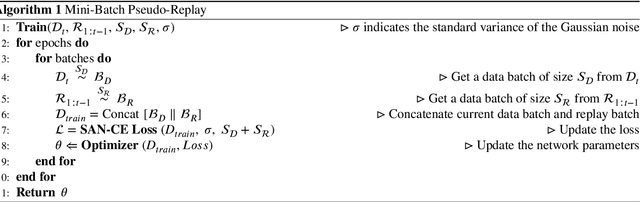

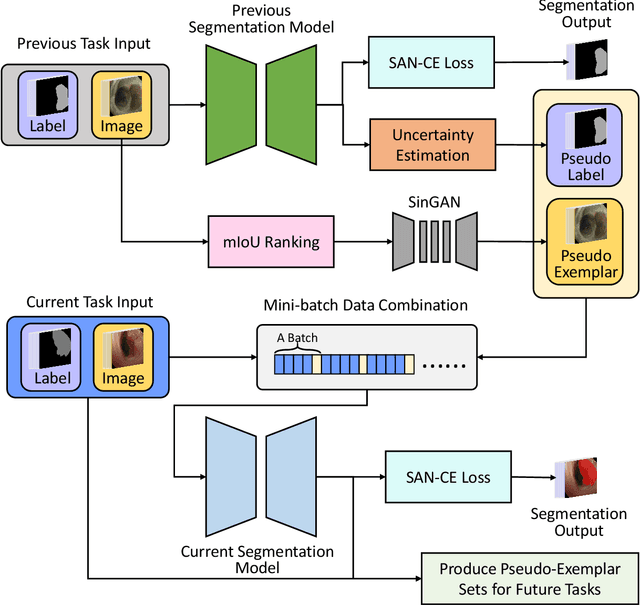
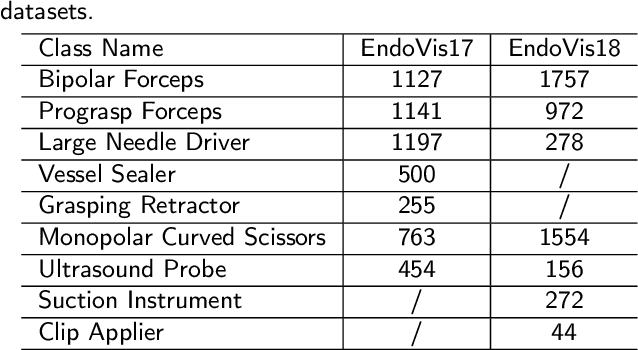
Endoscopy is a widely used technique for the early detection of diseases or robotic-assisted minimally invasive surgery (RMIS). Numerous deep learning (DL)-based research works have been developed for automated diagnosis or processing of endoscopic view. However, existing DL models may suffer from catastrophic forgetting. When new target classes are introduced over time or cross institutions, the performance of old classes may suffer severe degradation. More seriously, data privacy and storage issues may lead to the unavailability of old data when updating the model. Therefore, it is necessary to develop a continual learning (CL) methodology to solve the problem of catastrophic forgetting in endoscopic image segmentation. To tackle this, we propose a Endoscopy Continual Semantic Segmentation (EndoCSS) framework that does not involve the storage and privacy issues of exemplar data. The framework includes a mini-batch pseudo-replay (MB-PR) mechanism and a self-adaptive noisy cross-entropy (SAN-CE) loss. The MB-PR strategy circumvents privacy and storage issues by generating pseudo-replay images through a generative model. Meanwhile, the MB-PR strategy can also correct the model deviation to the replay data and current training data, which is aroused by the significant difference in the amount of current and replay images. Therefore, the model can perform effective representation learning on both new and old tasks. SAN-CE loss can help model fitting by adjusting the model's output logits, and also improve the robustness of training. Extensive continual semantic segmentation (CSS) experiments on public datasets demonstrate that our method can robustly and effectively address the catastrophic forgetting brought by class increment in endoscopy scenes. The results show that our framework holds excellent potential for real-world deployment in a streaming learning manner.
Cryo-forum: A framework for orientation recovery with uncertainty measure with the application in cryo-EM image analysis
Jul 19, 2023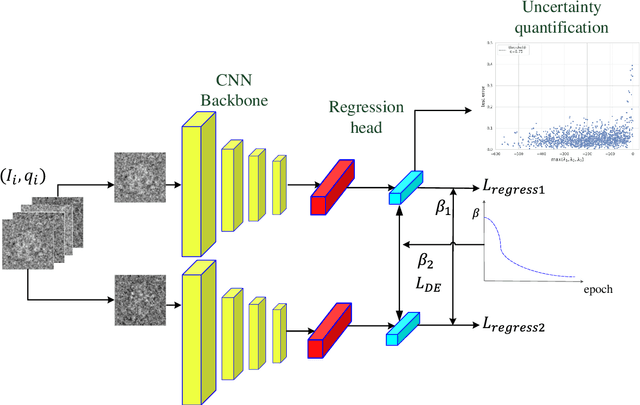
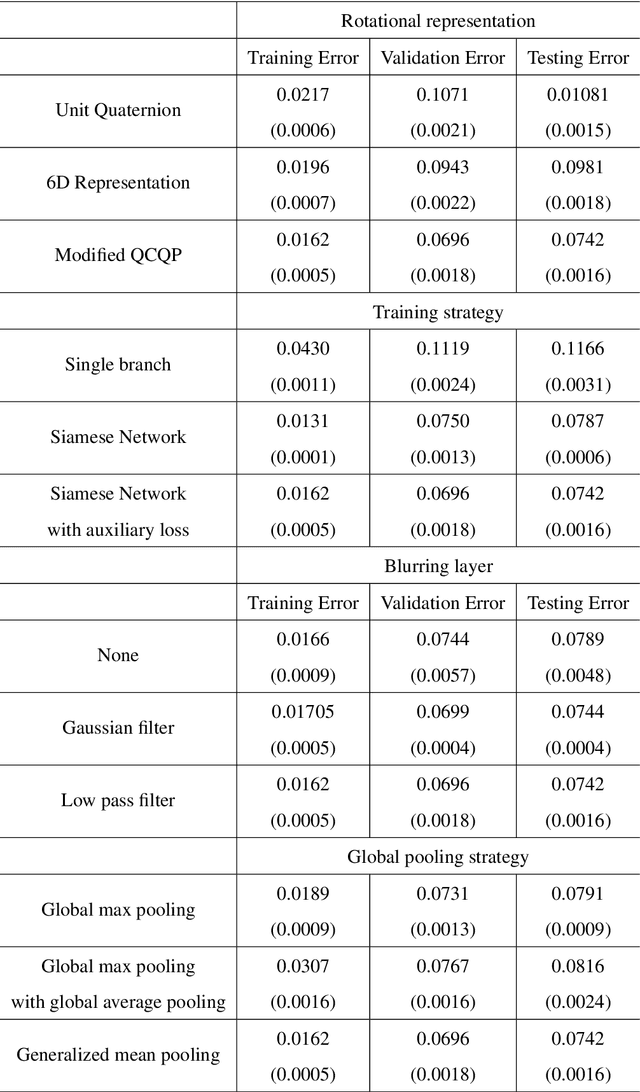
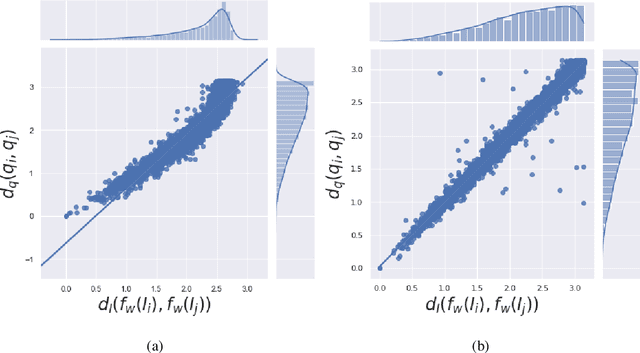
In single-particle cryo-electron microscopy (cryo-EM), the efficient determination of orientation parameters for 2D projection images poses a significant challenge yet is crucial for reconstructing 3D structures. This task is complicated by the high noise levels present in the cryo-EM datasets, which often include outliers, necessitating several time-consuming 2D clean-up processes. Recently, solutions based on deep learning have emerged, offering a more streamlined approach to the traditionally laborious task of orientation estimation. These solutions often employ amortized inference, eliminating the need to estimate parameters individually for each image. However, these methods frequently overlook the presence of outliers and may not adequately concentrate on the components used within the network. This paper introduces a novel approach that uses a 10-dimensional feature vector to represent the orientation and applies a Quadratically-Constrained Quadratic Program to derive the predicted orientation as a unit quaternion, supplemented by an uncertainty metric. Furthermore, we propose a unique loss function that considers the pairwise distances between orientations, thereby enhancing the accuracy of our method. Finally, we also comprehensively evaluate the design choices involved in constructing the encoder network, a topic that has not received sufficient attention in the literature. Our numerical analysis demonstrates that our methodology effectively recovers orientations from 2D cryo-EM images in an end-to-end manner. Importantly, the inclusion of uncertainty quantification allows for direct clean-up of the dataset at the 3D level. Lastly, we package our proposed methods into a user-friendly software suite named cryo-forum, designed for easy accessibility by the developers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge