"Image To Image Translation": models, code, and papers
CycleGAN with three different unpaired datasets
Aug 12, 2022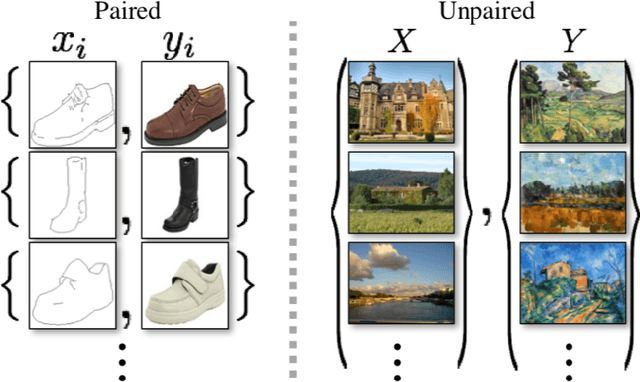
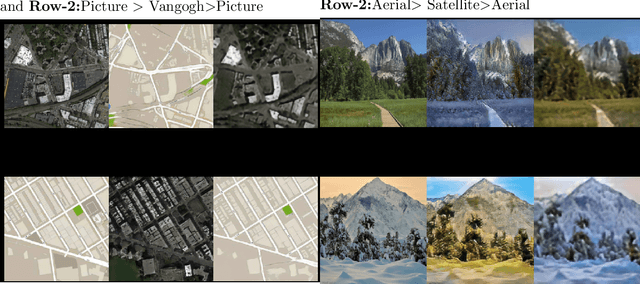
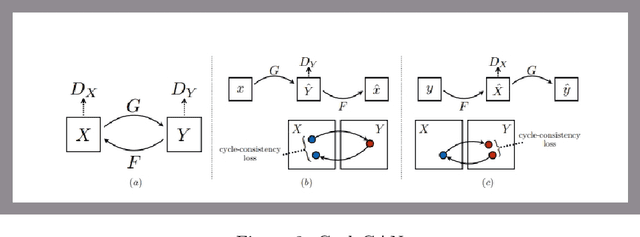


The original publication Unpaired Image-to-Image Translation using Cycle-Consistent Adversarial Networks served as the inspiration for this implementation project. Researchers developed a novel method for doing image-to-image translations using an unpaired dataset in the original study. Despite the fact that the pix2pix models findings are good, the matched dataset is frequently not available. In the absence of paired data, cycleGAN can therefore get over this issue by converting images to images. In order to lessen the difference between the images, they implemented cycle consistency loss.I evaluated CycleGAN with three different datasets, and this paper briefly discusses the findings and conclusions.
Digital twins of physical printing-imaging channel
Oct 28, 2022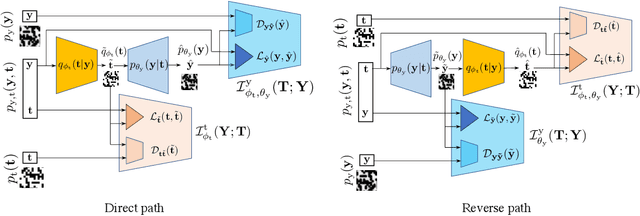
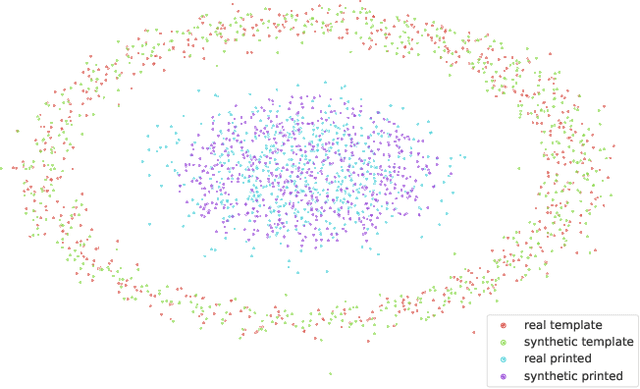
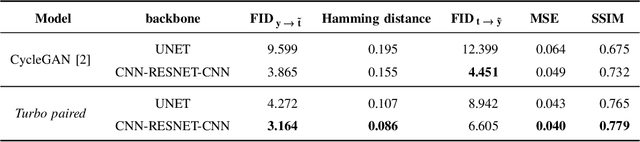
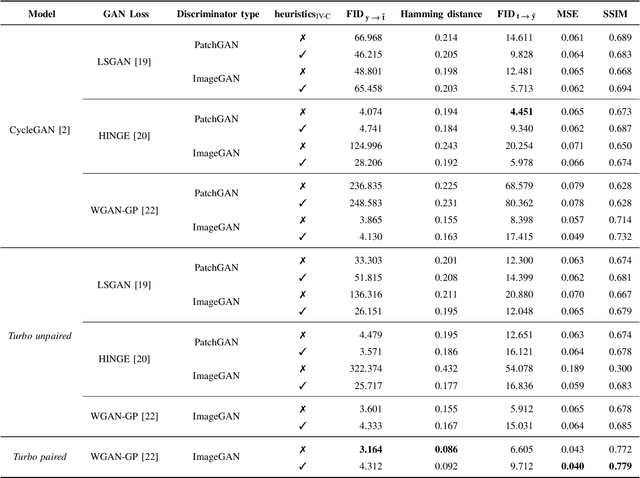
In this paper, we address the problem of modeling a printing-imaging channel built on a machine learning approach a.k.a. digital twin for anti-counterfeiting applications based on copy detection patterns (CDP). The digital twin is formulated on an information-theoretic framework called Turbo that uses variational approximations of mutual information developed for both encoder and decoder in a two-directional information passage. The proposed model generalizes several state-of-the-art architectures such as adversarial autoencoder (AAE), CycleGAN and adversarial latent space autoencoder (ALAE). This model can be applied to any type of printing and imaging and it only requires training data consisting of digital templates or artworks that are sent to a printing device and data acquired by an imaging device. Moreover, these data can be paired, unpaired or hybrid paired-unpaired which makes the proposed architecture very flexible and scalable to many practical setups. We demonstrate the impact of various architectural factors, metrics and discriminators on the overall system performance in the task of generation/prediction of printed CDP from their digital counterparts and vice versa. We also compare the proposed system with several state-of-the-art methods used for image-to-image translation applications.
Exploring Patch-wise Semantic Relation for Contrastive Learning in Image-to-Image Translation Tasks
Mar 03, 2022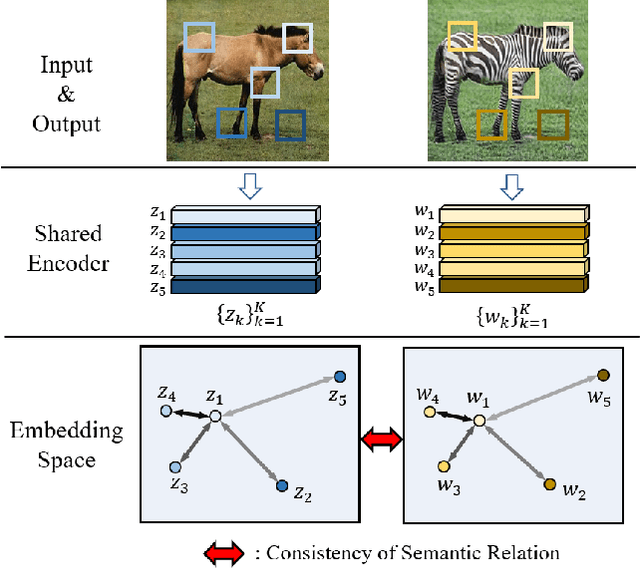
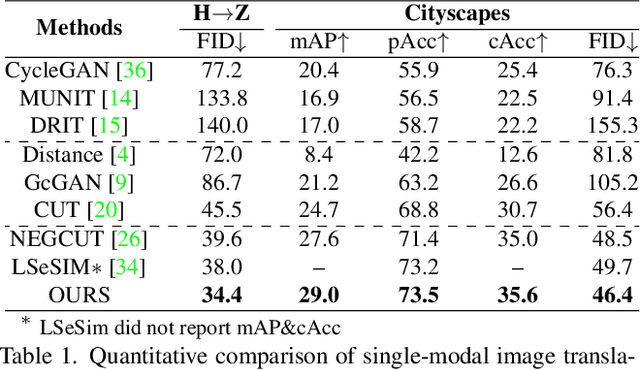
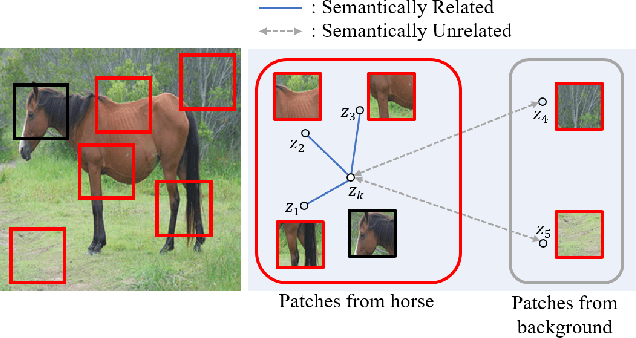
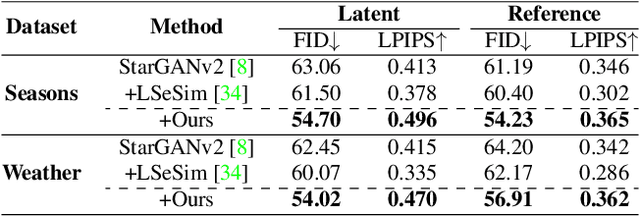
Recently, contrastive learning-based image translation methods have been proposed, which contrasts different spatial locations to enhance the spatial correspondence. However, the methods often ignore the diverse semantic relation within the images. To address this, here we propose a novel semantic relation consistency (SRC) regularization along with the decoupled contrastive learning, which utilize the diverse semantics by focusing on the heterogeneous semantics between the image patches of a single image. To further improve the performance, we present a hard negative mining by exploiting the semantic relation. We verified our method for three tasks: single-modal and multi-modal image translations, and GAN compression task for image translation. Experimental results confirmed the state-of-art performance of our method in all the three tasks.
XDGAN: Multi-Modal 3D Shape Generation in 2D Space
Oct 06, 2022
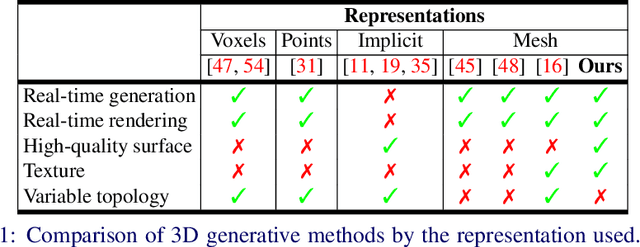

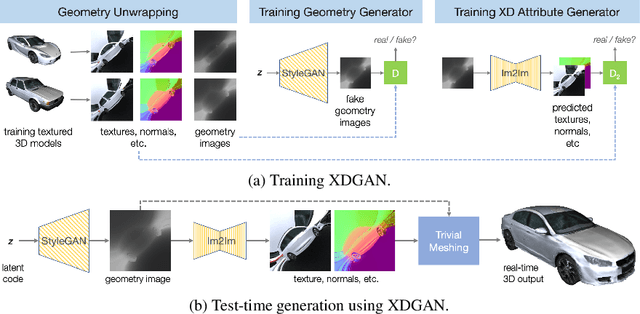
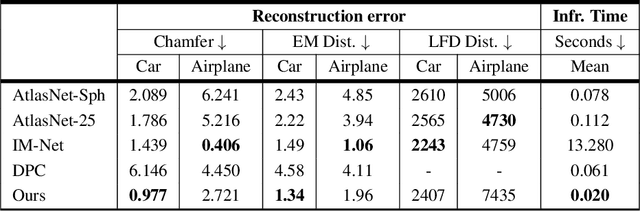
Generative models for 2D images has recently seen tremendous progress in quality, resolution and speed as a result of the efficiency of 2D convolutional architectures. However it is difficult to extend this progress into the 3D domain since most current 3D representations rely on custom network components. This paper addresses a central question: Is it possible to directly leverage 2D image generative models to generate 3D shapes instead? To answer this, we propose XDGAN, an effective and fast method for applying 2D image GAN architectures to the generation of 3D object geometry combined with additional surface attributes, like color textures and normals. Specifically, we propose a novel method to convert 3D shapes into compact 1-channel geometry images and leverage StyleGAN3 and image-to-image translation networks to generate 3D objects in 2D space. The generated geometry images are quick to convert to 3D meshes, enabling real-time 3D object synthesis, visualization and interactive editing. Moreover, the use of standard 2D architectures can help bring more 2D advances into the 3D realm. We show both quantitatively and qualitatively that our method is highly effective at various tasks such as 3D shape generation, single view reconstruction and shape manipulation, while being significantly faster and more flexible compared to recent 3D generative models.
What can we learn about a generated image corrupting its latent representation?
Oct 12, 2022Generative adversarial networks (GANs) offer an effective solution to the image-to-image translation problem, thereby allowing for new possibilities in medical imaging. They can translate images from one imaging modality to another at a low cost. For unpaired datasets, they rely mostly on cycle loss. Despite its effectiveness in learning the underlying data distribution, it can lead to a discrepancy between input and output data. The purpose of this work is to investigate the hypothesis that we can predict image quality based on its latent representation in the GANs bottleneck. We achieve this by corrupting the latent representation with noise and generating multiple outputs. The degree of differences between them is interpreted as the strength of the representation: the more robust the latent representation, the fewer changes in the output image the corruption causes. Our results demonstrate that our proposed method has the ability to i) predict uncertain parts of synthesized images, and ii) identify samples that may not be reliable for downstream tasks, e.g., liver segmentation task.
Counterfactual Explanation and Instance-Generation using Cycle-Consistent Generative Adversarial Networks
Jan 21, 2023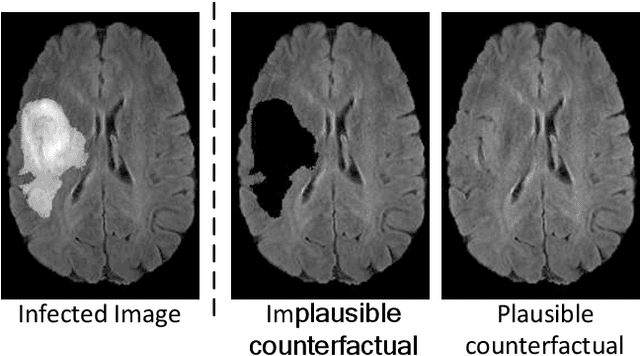
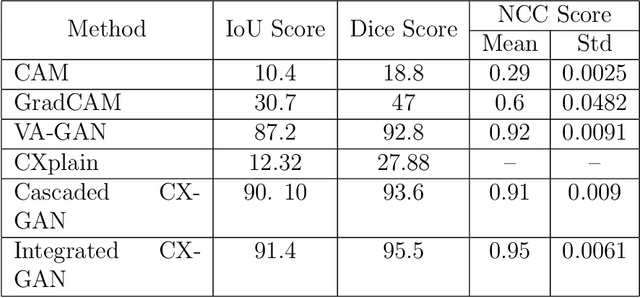
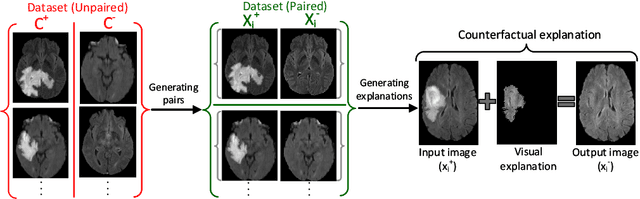
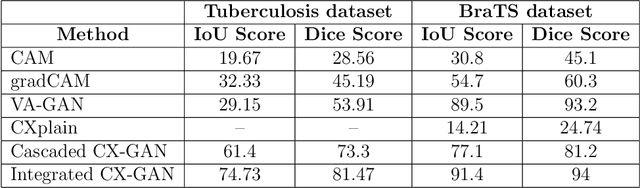
The image-based diagnosis is now a vital aspect of modern automation assisted diagnosis. To enable models to produce pixel-level diagnosis, pixel-level ground-truth labels are essentially required. However, since it is often not straight forward to obtain the labels in many application domains such as in medical image, classification-based approaches have become the de facto standard to perform the diagnosis. Though they can identify class-salient regions, they may not be useful for diagnosis where capturing all of the evidences is important requirement. Alternatively, a counterfactual explanation (CX) aims at providing explanations using a casual reasoning process of form "If X has not happend, Y would not heppend". Existing CX approaches, however, use classifier to explain features that can change its predictions. Thus, they can only explain class-salient features, rather than entire object of interest. This hence motivates us to propose a novel CX strategy that is not reliant on image classification. This work is inspired from the recent developments in generative adversarial networks (GANs) based image-to-image domain translation, and leverages to translate an abnormal image to counterpart normal image (i.e. counterfactual instance CI) to find discrepancy maps between the two. Since it is generally not possible to obtain abnormal and normal image pairs, we leverage Cycle-Consistency principle (a.k.a CycleGAN) to perform the translation in unsupervised way. We formulate CX in terms of a discrepancy map that, when added from the abnormal image, will make it indistinguishable from the CI. We evaluate our method on three datasets including a synthetic, tuberculosis and BraTS dataset. All these experiments confirm the supremacy of propose method in generating accurate CX and CI.
Shapes2Toon: Generating Cartoon Characters from Simple Geometric Shapes
Nov 03, 2022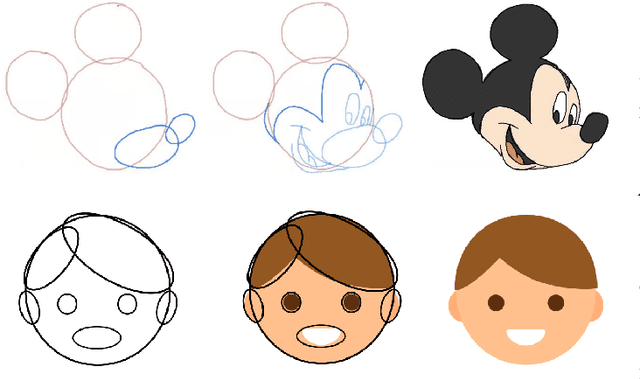
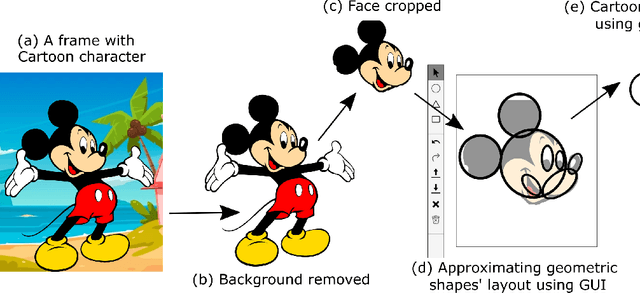
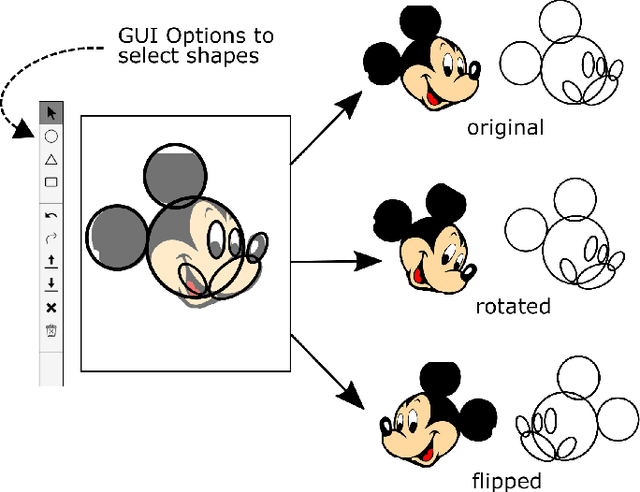
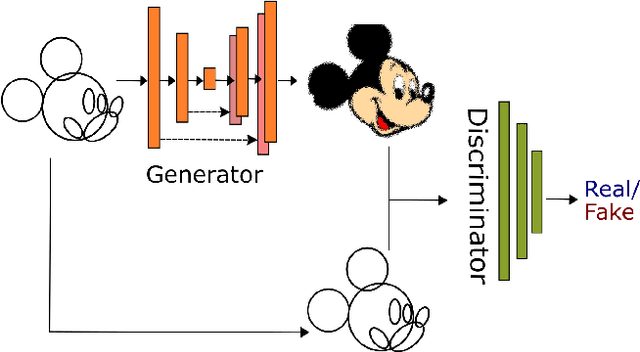
Cartoons are an important part of our entertainment culture. Though drawing a cartoon is not for everyone, creating it using an arrangement of basic geometric primitives that approximates that character is a fairly frequent technique in art. The key motivation behind this technique is that human bodies - as well as cartoon figures - can be split down into various basic geometric primitives. Numerous tutorials are available that demonstrate how to draw figures using an appropriate arrangement of fundamental shapes, thus assisting us in creating cartoon characters. This technique is very beneficial for children in terms of teaching them how to draw cartoons. In this paper, we develop a tool - shape2toon - that aims to automate this approach by utilizing a generative adversarial network which combines geometric primitives (i.e. circles) and generate a cartoon figure (i.e. Mickey Mouse) depending on the given approximation. For this purpose, we created a dataset of geometrically represented cartoon characters. We apply an image-to-image translation technique on our dataset and report the results in this paper. The experimental results show that our system can generate cartoon characters from input layout of geometric shapes. In addition, we demonstrate a web-based tool as a practical implication of our work.
CoMoGAN: continuous model-guided image-to-image translation
Apr 08, 2021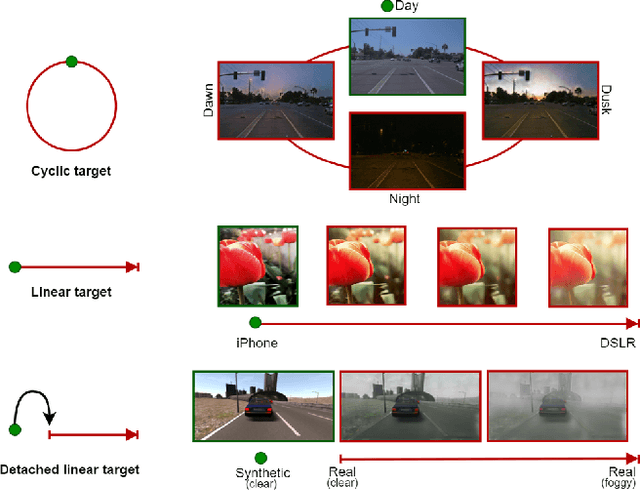

CoMoGAN is a continuous GAN relying on the unsupervised reorganization of the target data on a functional manifold. To that matter, we introduce a new Functional Instance Normalization layer and residual mechanism, which together disentangle image content from position on target manifold. We rely on naive physics-inspired models to guide the training while allowing private model/translations features. CoMoGAN can be used with any GAN backbone and allows new types of image translation, such as cyclic image translation like timelapse generation, or detached linear translation. On all datasets, it outperforms the literature. Our code is available at http://github.com/cv-rits/CoMoGAN .
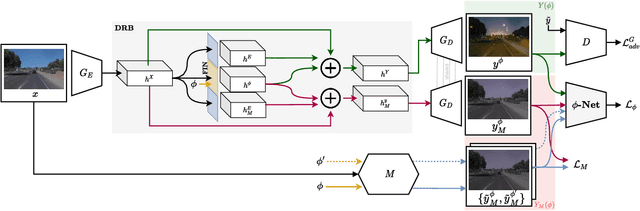
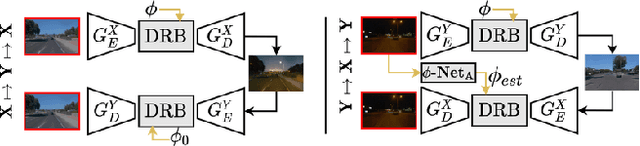
Deceiving Image-to-Image Translation Networks for Autonomous Driving with Adversarial Perturbations
Jan 06, 2020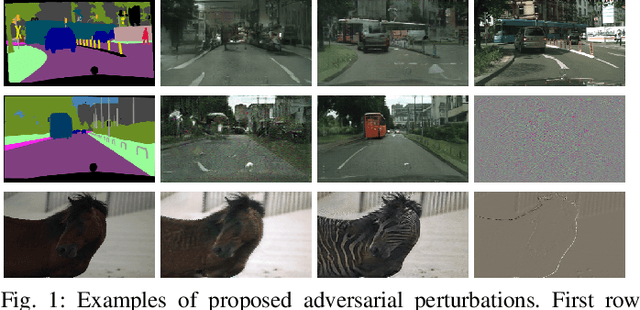
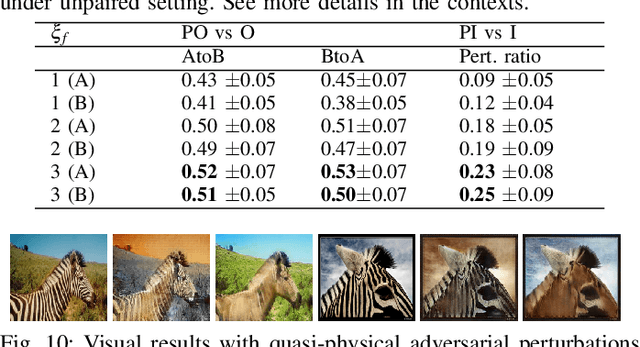
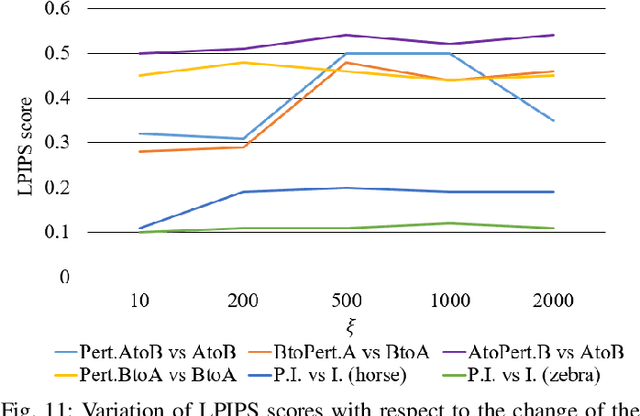
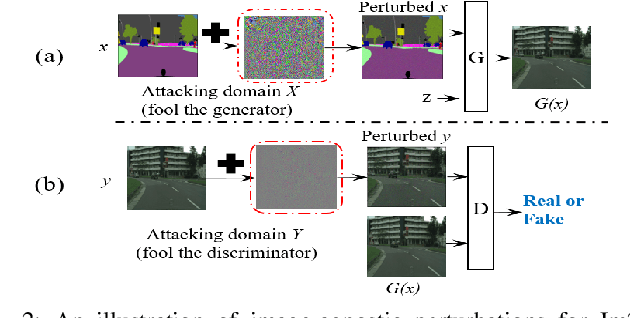
Deep neural networks (DNNs) have achieved impressive performance on handling computer vision problems, however, it has been found that DNNs are vulnerable to adversarial examples. For such reason, adversarial perturbations have been recently studied in several respects. However, most previous works have focused on image classification tasks, and it has never been studied regarding adversarial perturbations on Image-to-image (Im2Im) translation tasks, showing great success in handling paired and/or unpaired mapping problems in the field of autonomous driving and robotics. This paper examines different types of adversarial perturbations that can fool Im2Im frameworks for autonomous driving purpose. We propose both quasi-physical and digital adversarial perturbations that can make Im2Im models yield unexpected results. We then empirically analyze these perturbations and show that they generalize well under both paired for image synthesis and unpaired settings for style transfer. We also validate that there exist some perturbation thresholds over which the Im2Im mapping is disrupted or impossible. The existence of these perturbations reveals that there exist crucial weaknesses in Im2Im models. Lastly, we show that our methods illustrate how perturbations affect the quality of outputs, pioneering the improvement of the robustness of current SOTA networks for autonomous driving.
Diabetic foot ulcers monitoring by employing super resolution and noise reduction deep learning techniques
Sep 20, 2022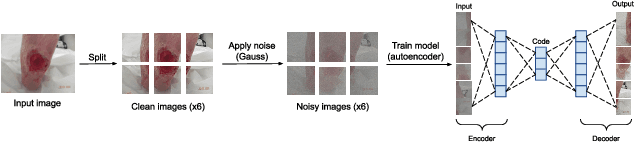
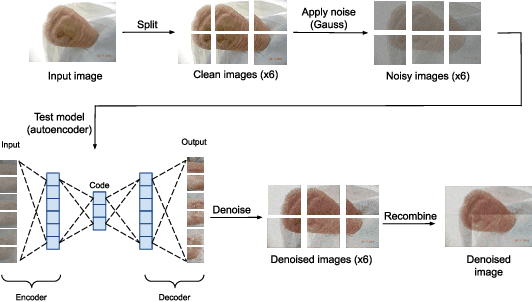
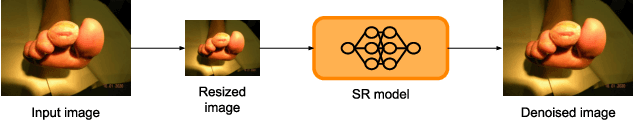
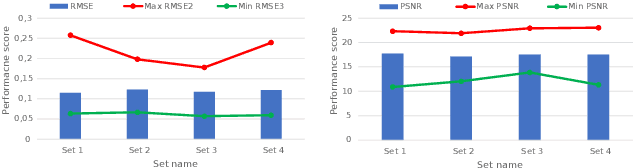
Diabetic foot ulcers (DFUs) constitute a serious complication for people with diabetes. The care of DFU patients can be substantially improved through self-management, in order to achieve early-diagnosis, ulcer prevention, and complications management in existing ulcers. In this paper, we investigate two categories of image-to-image translation techniques (ItITT), which will support decision making and monitoring of diabetic foot ulcers: noise reduction and super-resolution. In the former case, we investigated the capabilities on noise removal, for convolutional neural network stacked-autoencoders (CNN-SAE). CNN-SAE was tested on RGB images, induced with Gaussian noise. The latter scenario involves the deployment of four deep learning super-resolution models. The performance of all models, for both scenarios, was evaluated in terms of execution time and perceived quality. Results indicate that applied techniques consist a viable and easy to implement alternative that should be used by any system designed for DFU monitoring.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge