EMDS-7: Environmental Microorganism Image Dataset Seventh Version for Multiple Object Detection Evaluation
Oct 28, 2021Hechen Yang, Chen Li, Xin Zhao, Bencheng Cai, Jiawei Zhang, Pingli Ma, Peng Zhao, Ao Chen, Tao Jiang, Hongzan Sun, Yueyang Teng, Shouliang Qi, Tao Jiang, Marcin Grzegorzek
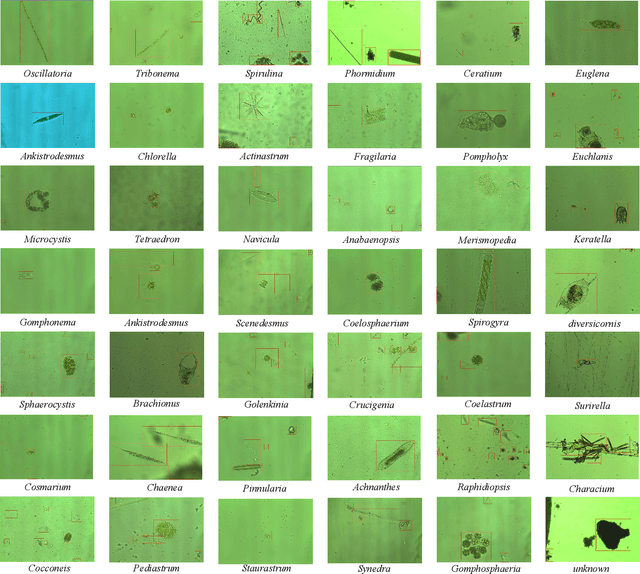
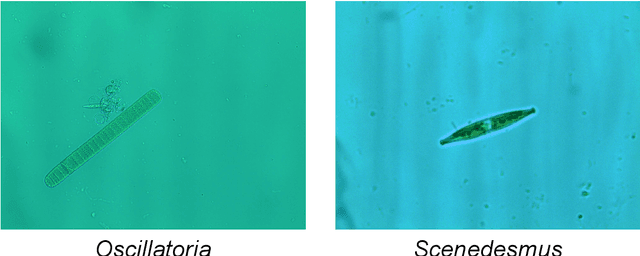

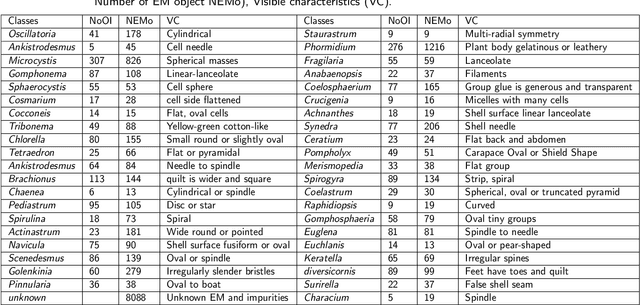

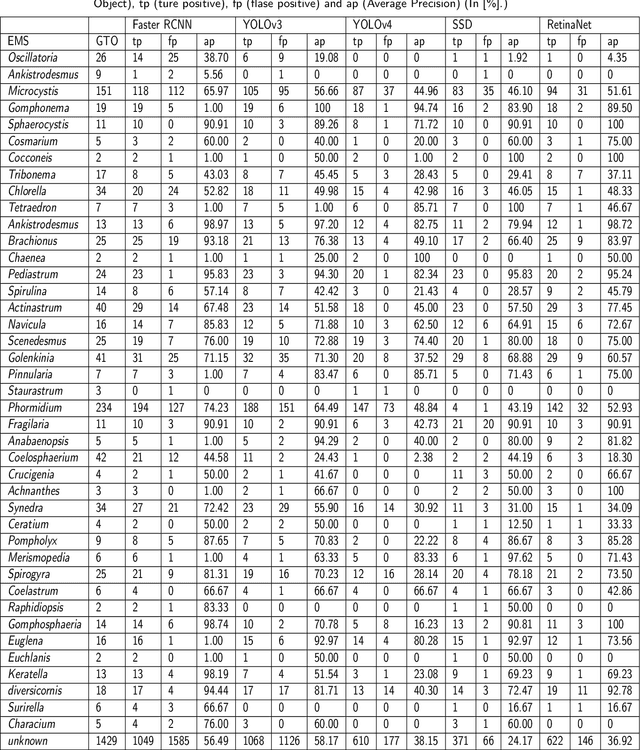
The Environmental Microorganism Image Dataset Seventh Version (EMDS-7) is a microscopic image data set, including the original Environmental Microorganism images (EMs) and the corresponding object labeling files in ".XML" format file. The EMDS-7 data set consists of 41 types of EMs, which has a total of 2365 images and 13216 labeled objects. The EMDS-7 database mainly focuses on the object detection. In order to prove the effectiveness of EMDS-7, we select the most commonly used deep learning methods (Faster-RCNN, YOLOv3, YOLOv4, SSD and RetinaNet) and evaluation indices for testing and evaluation. EMDS-7 is freely published for non-commercial purpose at: https://figshare.com/articles/dataset/EMDS-7_DataSet/16869571
Unsupervised Domain Adaptation for Retinal Vessel Segmentation with Adversarial Learning and Transfer Normalization
Aug 04, 2021Wei Feng, Lie Ju, Lin Wang, Kaimin Song, Xin Wang, Xin Zhao, Qingyi Tao, Zongyuan Ge
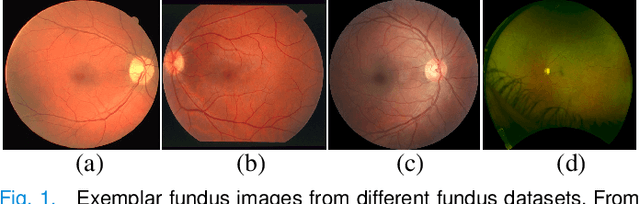
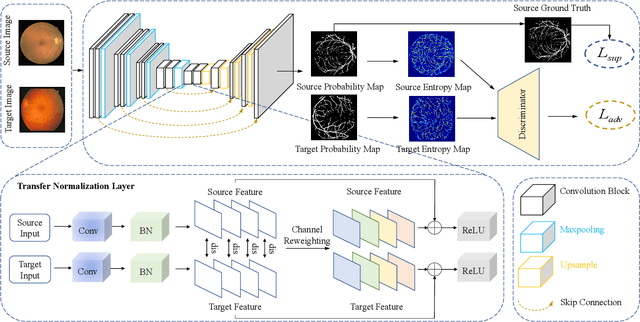
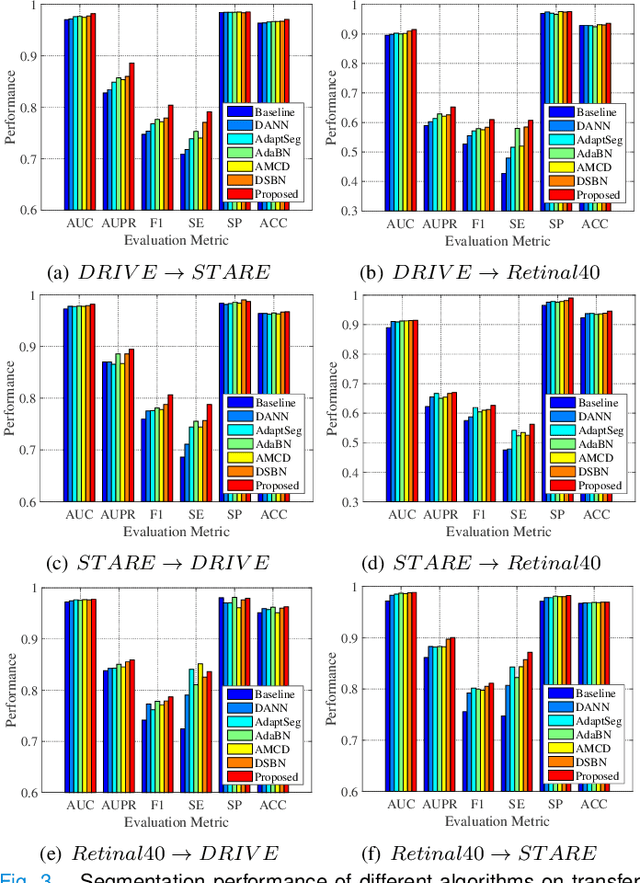

Retinal vessel segmentation plays a key role in computer-aided screening, diagnosis, and treatment of various cardiovascular and ophthalmic diseases. Recently, deep learning-based retinal vessel segmentation algorithms have achieved remarkable performance. However, due to the domain shift problem, the performance of these algorithms often degrades when they are applied to new data that is different from the training data. Manually labeling new data for each test domain is often a time-consuming and laborious task. In this work, we explore unsupervised domain adaptation in retinal vessel segmentation by using entropy-based adversarial learning and transfer normalization layer to train a segmentation network, which generalizes well across domains and requires no annotation of the target domain. Specifically, first, an entropy-based adversarial learning strategy is developed to reduce the distribution discrepancy between the source and target domains while also achieving the objective of entropy minimization on the target domain. In addition, a new transfer normalization layer is proposed to further boost the transferability of the deep network. It normalizes the features of each domain separately to compensate for the domain distribution gap. Besides, it also adaptively selects those feature channels that are more transferable between domains, thus further enhancing the generalization performance of the network. We conducted extensive experiments on three regular fundus image datasets and an ultra-widefield fundus image dataset, and the results show that our approach yields significant performance gains compared to other state-of-the-art methods.
A Comparison for Patch-level Classification of Deep Learning Methods on Transparent Environmental Microorganism Images: from Convolutional Neural Networks to Visual Transformers
Jul 21, 2021Hechen Yang, Chen Li, Jinghua Zhang, Peng Zhao, Ao Chen, Xin Zhao, Tao Jiang, Marcin Grzegorzek
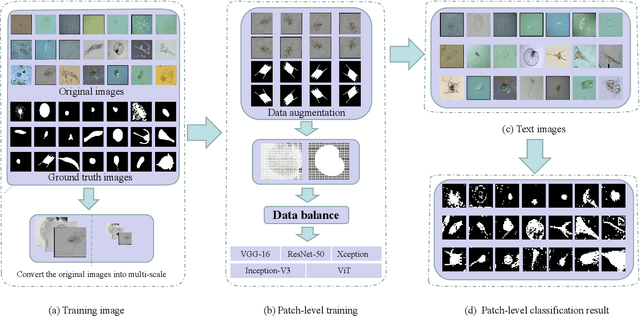
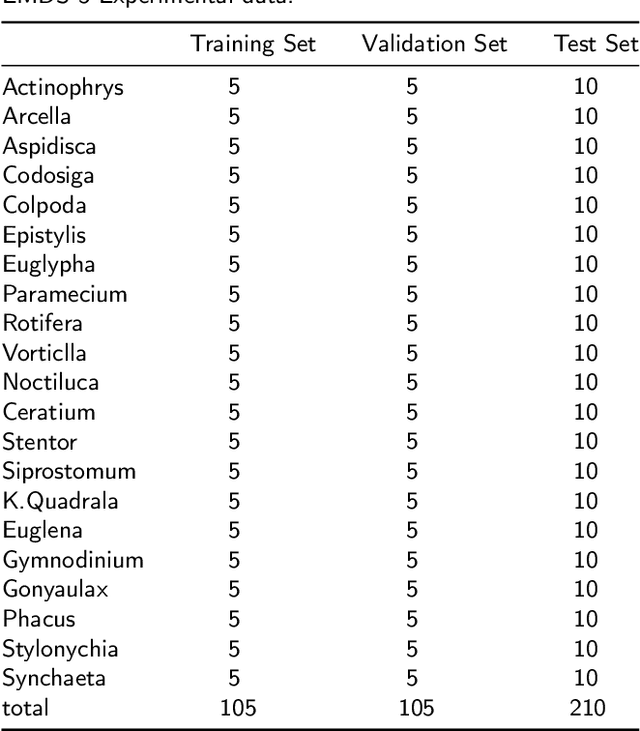
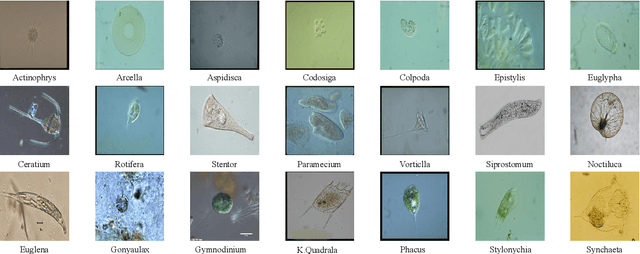
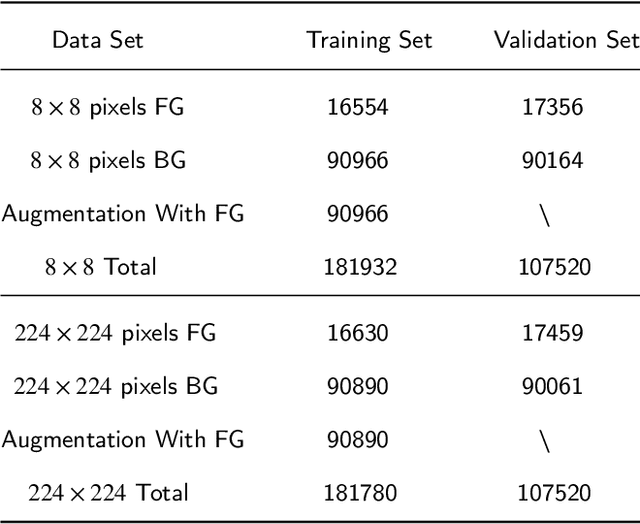
Nowadays, analysis of Transparent Environmental Microorganism Images (T-EM images) in the field of computer vision has gradually become a new and interesting spot. This paper compares different deep learning classification performance for the problem that T-EM images are challenging to analyze. We crop the T-EM images into 8 * 8 and 224 * 224 pixel patches in the same proportion and then divide the two different pixel patches into foreground and background according to ground truth. We also use four convolutional neural networks and a novel ViT network model to compare the foreground and background classification experiments. We conclude that ViT performs the worst in classifying 8 * 8 pixel patches, but it outperforms most convolutional neural networks in classifying 224 * 224 pixel patches.
Medical Matting: A New Perspective on Medical Segmentation with Uncertainty
Jul 08, 2021Lin Wang, Lie Ju, Donghao Zhang, Xin Wang, Wanji He, Yelin Huang, Zhiwen Yang, Xuan Yao, Xin Zhao, Xiufen Ye, Zongyuan Ge
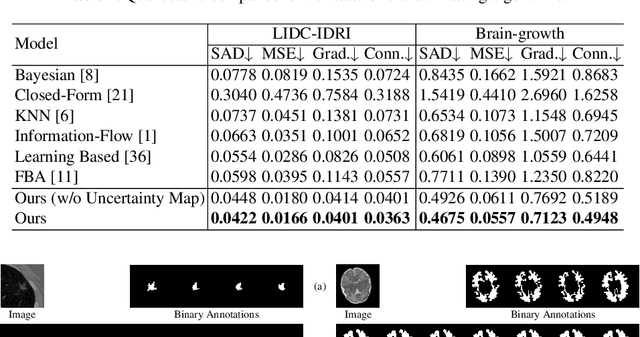
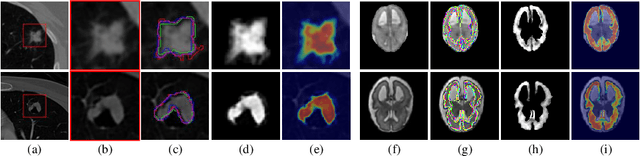


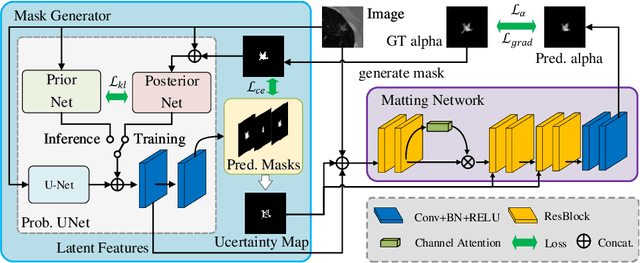
In medical image segmentation, it is difficult to mark ambiguous areas accurately with binary masks, especially when dealing with small lesions. Therefore, it is a challenge for radiologists to reach a consensus by using binary masks under the condition of multiple annotations. However, these areas may contain anatomical structures that are conducive to diagnosis. Uncertainty is introduced to study these situations. Nevertheless, the uncertainty is usually measured by the variances between predictions in a multiple trial way. It is not intuitive, and there is no exact correspondence in the image. Inspired by image matting, we introduce matting as a soft segmentation method and a new perspective to deal with and represent uncertain regions into medical scenes, namely medical matting. More specifically, because there is no available medical matting dataset, we first labeled two medical datasets with alpha matte. Secondly, the matting method applied to the natural image is not suitable for the medical scene, so we propose a new architecture to generate binary masks and alpha matte in a row. Thirdly, the uncertainty map is introduced to highlight the ambiguous regions from the binary results and improve the matting performance. Evaluated on these datasets, the proposed model outperformed state-of-the-art matting algorithms by a large margin, and alpha matte is proved to be a more efficient labeling form than a binary mask.
A Comparison for Patch-level Classification of Deep Learning Methods on Transparent Images: from Convolutional Neural Networks to Visual Transformers
Jun 22, 2021Hechen Yang, Chen Li, Peng Zhao, Ao Chen, Xin Zhao, Marcin Grzegorzek




Nowadays, analysis of transparent images in the field of computer vision has gradually become a hot spot. In this paper, we compare the classification performance of different deep learning for the problem that transparent images are difficult to analyze. We crop the transparent images into 8 * 8 and 224 * 224 pixels patches in the same proportion, and then divide the two different pixels patches into foreground and background according to groundtruch. We also use 4 types of convolutional neural networks and a novel ViT network model to compare the foreground and background classification experiments. We conclude that ViT performs the worst in classifying 8 * 8 pixels patches, but it outperforms most convolutional neural networks in classifying 224 * 224.
A State-of-the-art Survey of Object Detection Techniques in Microorganism Image Analysis: from Traditional Image Processing and Classical Machine Learning to Current Deep Convolutional Neural Networks and Potential Visual Transformers
May 07, 2021Chen Li, Pingli Ma, Md Mamunur Rahaman, Yudong Yao, Jiawei Zhang, Shuojia Zou, Xin Zhao, Marcin Grzegorzek
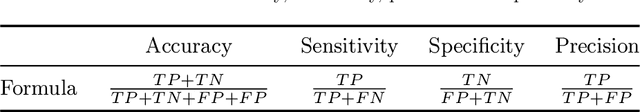
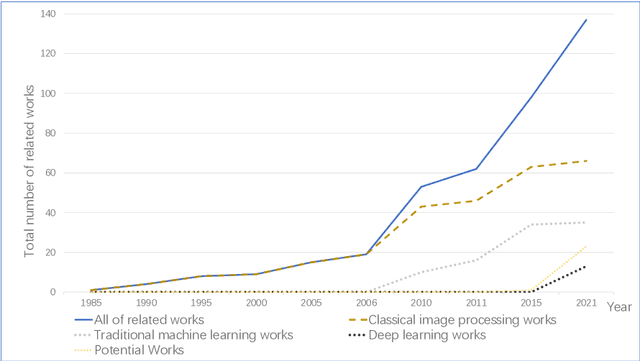
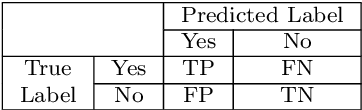
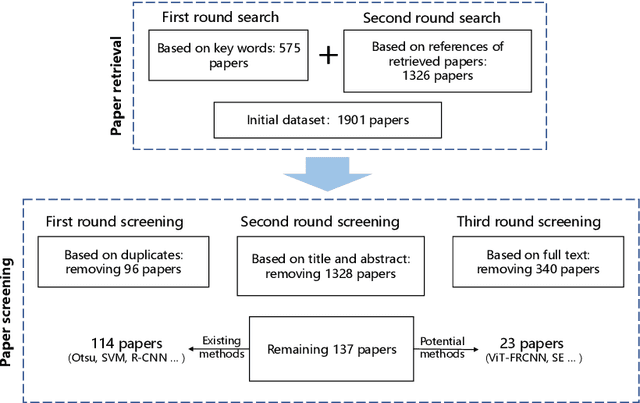

Microorganisms play a vital role in human life. Therefore, microorganism detection is of great significance to human beings. However, the traditional manual microscopic detection methods have the disadvantages of long detection cycle, low detection accuracy in large orders, and great difficulty in detecting uncommon microorganisms. Therefore, it is meaningful to apply computer image analysis technology to the field of microorganism detection. Computer image analysis can realize high-precision and high-efficiency detection of microorganisms. In this review, first,we analyse the existing microorganism detection methods in chronological order, from traditional image processing and traditional machine learning to deep learning methods. Then, we analyze and summarize these existing methods and introduce some potential methods, including visual transformers. In the end, the future development direction and challenges of microorganism detection are discussed. In general, we have summarized 137 related technical papers from 1985 to the present. This review will help researchers have a more comprehensive understanding of the development process, research status, and future trends in the field of microorganism detection and provide a reference for researchers in other fields.
Relational Subsets Knowledge Distillation for Long-tailed Retinal Diseases Recognition
Apr 22, 2021Lie Ju, Xin Wang, Lin Wang, Tongliang Liu, Xin Zhao, Tom Drummond, Dwarikanath Mahapatra, Zongyuan Ge
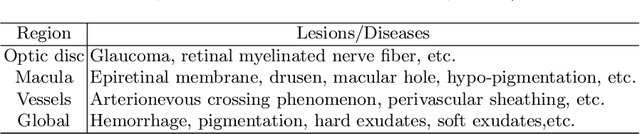
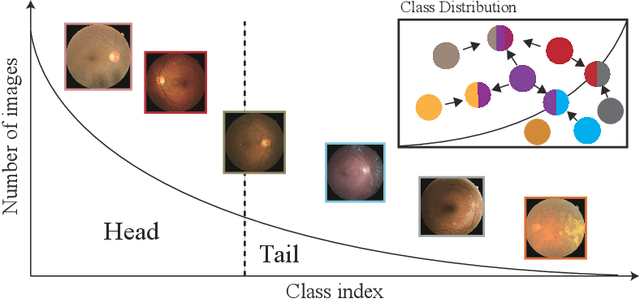

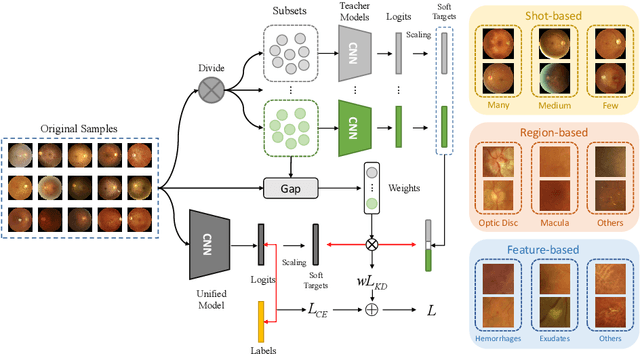
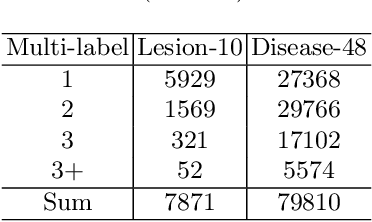
In the real world, medical datasets often exhibit a long-tailed data distribution (i.e., a few classes occupy most of the data, while most classes have rarely few samples), which results in a challenging imbalance learning scenario. For example, there are estimated more than 40 different kinds of retinal diseases with variable morbidity, however with more than 30+ conditions are very rare from the global patient cohorts, which results in a typical long-tailed learning problem for deep learning-based screening models. In this study, we propose class subset learning by dividing the long-tailed data into multiple class subsets according to prior knowledge, such as regions and phenotype information. It enforces the model to focus on learning the subset-specific knowledge. More specifically, there are some relational classes that reside in the fixed retinal regions, or some common pathological features are observed in both the majority and minority conditions. With those subsets learnt teacher models, then we are able to distill the multiple teacher models into a unified model with weighted knowledge distillation loss. The proposed framework proved to be effective for the long-tailed retinal diseases recognition task. The experimental results on two different datasets demonstrate that our method is flexible and can be easily plugged into many other state-of-the-art techniques with significant improvements.
A Comprehensive Review of Image Analysis Methods for Microorganism Counting: From Classical Image Processing to Deep Learning Approaches
Mar 25, 2021Chen Li, Jiawei Zhang, Md Mamunur Rahaman, Yudong Yao, Pingli Ma, Jinghua Zhang, Xin Zhao, Tao Jiang, Marcin Grzegorzek

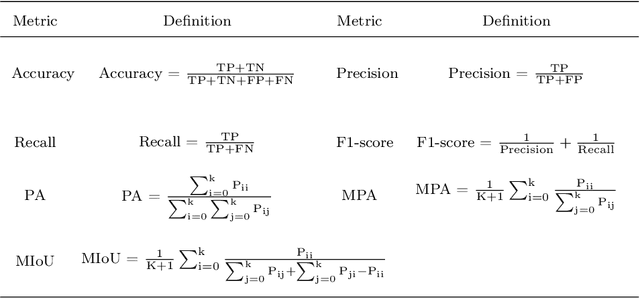
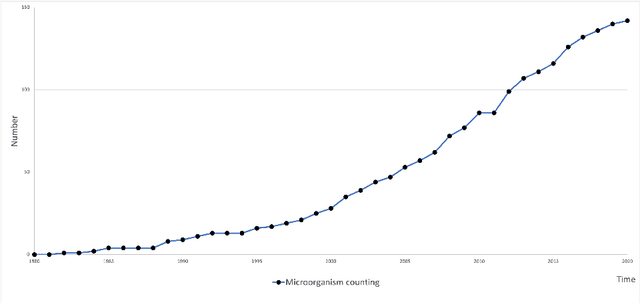
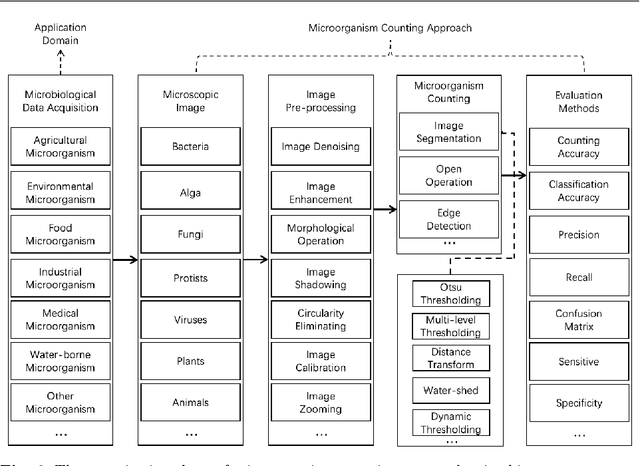
Microorganisms such as bacteria and fungi play essential roles in many application fields, like biotechnique, medical technique and industrial domain. Microorganism counting techniques are crucial in microorganism analysis, helping biologists and related researchers quantitatively analyze the microorganisms and calculate their characteristics, such as biomass concentration and biological activity. However, traditional microorganism manual counting methods are time consuming and subjective, which cannot be applied in large-scale applications. In order to improve this situation, image analysis-based microorganism counting systems are developed since 1980s, which consists of digital image processing, image segmentation, image classification and so on. Moreover, image analysis-based microorganism counting methods are efficient comparing with traditional plate counting methods. In this article, we have studied the development of microorganism counting methods using digital image analysis. Firstly, the microorganisms are grouped as bacteria and other microorganisms. Then, the related articles are summarized based on image segmentation methods. Each part of articles are reviewed by time periods. Moreover, commonly used image processing methods for microorganism counting are summarized and analyzed to find technological common points. More than 142 papers are summarized in this article. In conclusion, this article can be referred to researchers to determine the development trend in microorganism counting field and further analyze the potential applications
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge