Interpretable Graph Neural Networks for Connectome-Based Brain Disorder Analysis
Jun 30, 2022Hejie Cui, Wei Dai, Yanqiao Zhu, Xiaoxiao Li, Lifang He, Carl Yang
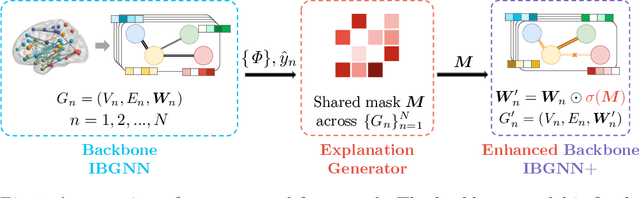
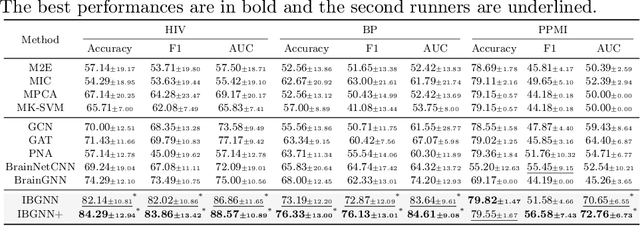
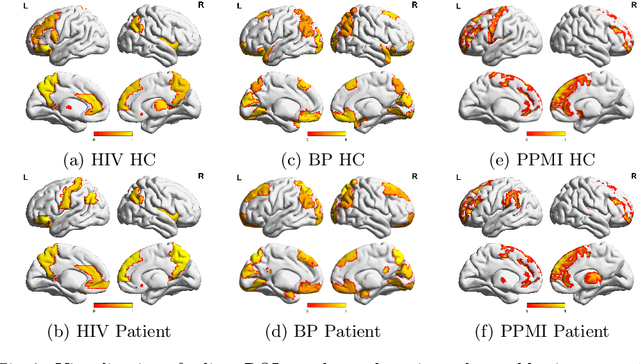
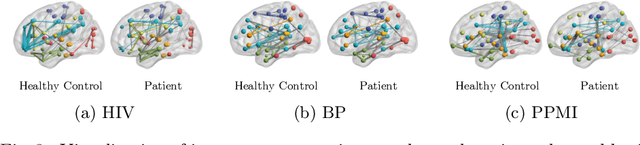
Human brains lie at the core of complex neurobiological systems, where the neurons, circuits, and subsystems interact in enigmatic ways. Understanding the structural and functional mechanisms of the brain has long been an intriguing pursuit for neuroscience research and clinical disorder therapy. Mapping the connections of the human brain as a network is one of the most pervasive paradigms in neuroscience. Graph Neural Networks (GNNs) have recently emerged as a potential method for modeling complex network data. Deep models, on the other hand, have low interpretability, which prevents their usage in decision-critical contexts like healthcare. To bridge this gap, we propose an interpretable framework to analyze disorder-specific Regions of Interest (ROIs) and prominent connections. The proposed framework consists of two modules: a brain-network-oriented backbone model for disease prediction and a globally shared explanation generator that highlights disorder-specific biomarkers including salient ROIs and important connections. We conduct experiments on three real-world datasets of brain disorders. The results verify that our framework can obtain outstanding performance and also identify meaningful biomarkers. All code for this work is available at https://github.com/HennyJie/IBGNN.git.
Dynamic Bank Learning for Semi-supervised Federated Image Diagnosis with Class Imbalance
Jun 27, 2022Meirui Jiang, Hongzheng Yang, Xiaoxiao Li, Quande Liu, Pheng-Ann Heng, Qi Dou
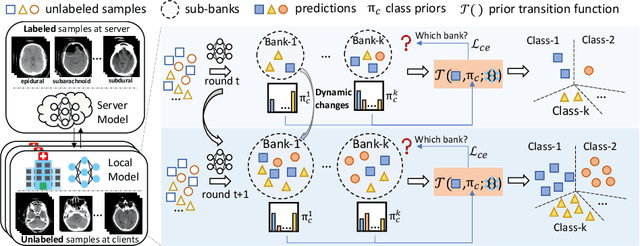
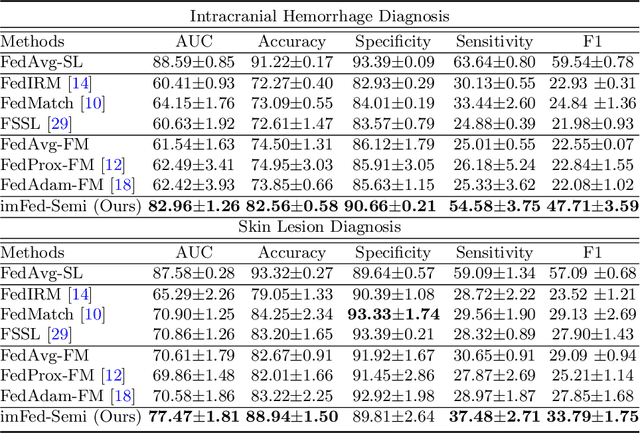
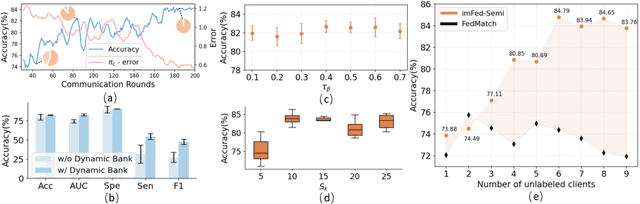
Despite recent progress on semi-supervised federated learning (FL) for medical image diagnosis, the problem of imbalanced class distributions among unlabeled clients is still unsolved for real-world use. In this paper, we study a practical yet challenging problem of class imbalanced semi-supervised FL (imFed-Semi), which allows all clients to have only unlabeled data while the server just has a small amount of labeled data. This imFed-Semi problem is addressed by a novel dynamic bank learning scheme, which improves client training by exploiting class proportion information. This scheme consists of two parts, i.e., the dynamic bank construction to distill various class proportions for each local client, and the sub-bank classification to impose the local model to learn different class proportions. We evaluate our approach on two public real-world medical datasets, including the intracranial hemorrhage diagnosis with 25,000 CT slices and skin lesion diagnosis with 10,015 dermoscopy images. The effectiveness of our method has been validated with significant performance improvements (7.61% and 4.69%) compared with the second-best on the accuracy, as well as comprehensive analytical studies. Code is available at https://github.com/med-air/imFedSemi.
Hypernetwork-based Personalized Federated Learning for Multi-Institutional CT Imaging
Jun 09, 2022Ziyuan Yang, Wenjun Xia, Zexin Lu, Yingyu Chen, Xiaoxiao Li, Yi Zhang
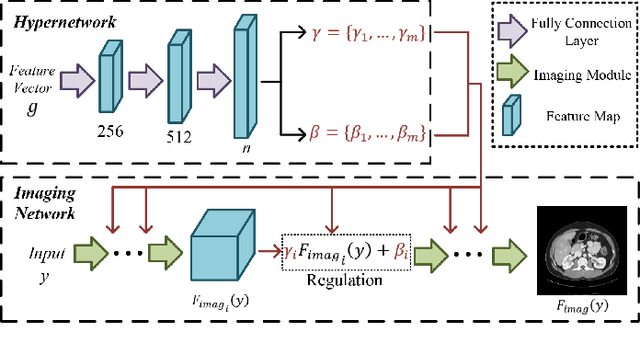
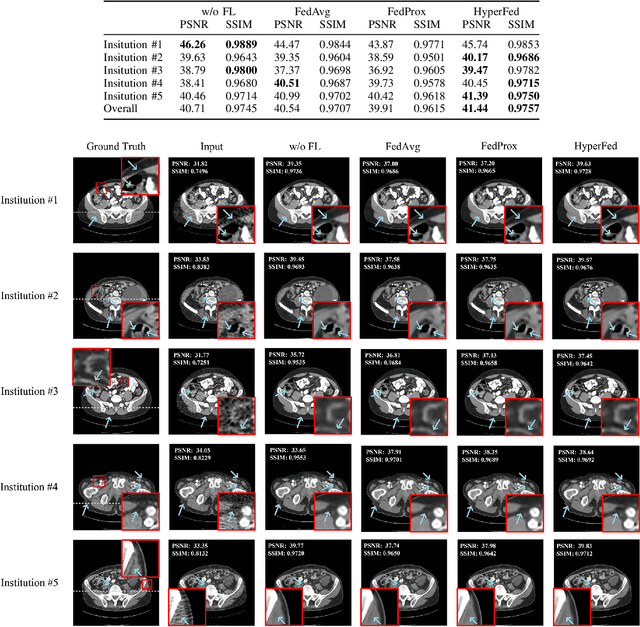
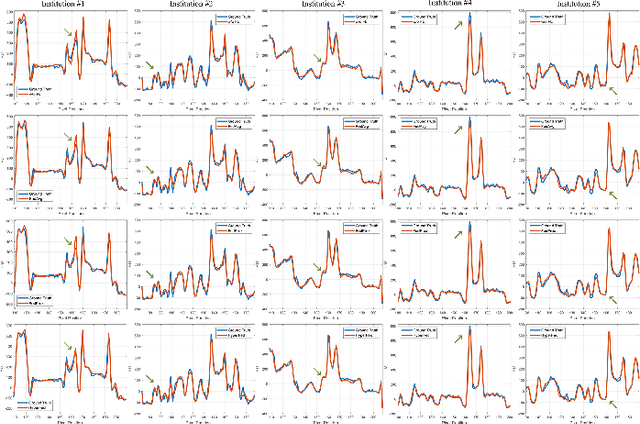
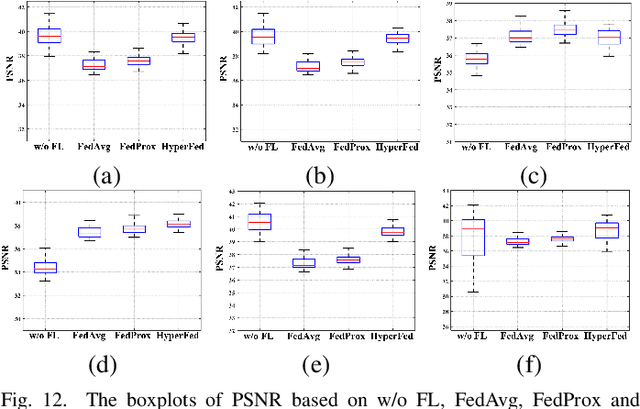
Computed tomography (CT) is of great importance in clinical practice due to its powerful ability to provide patients' anatomical information without any invasive inspection, but its potential radiation risk is raising people's concerns. Deep learning-based methods are considered promising in CT reconstruction, but these network models are usually trained with the measured data obtained from specific scanning protocol and need to centralizedly collect large amounts of data, which will lead to serious data domain shift, and privacy concerns. To relieve these problems, in this paper, we propose a hypernetwork-based federated learning method for personalized CT imaging, dubbed as HyperFed. The basic assumption of HyperFed is that the optimization problem for each institution can be divided into two parts: the local data adaption problem and the global CT imaging problem, which are implemented by an institution-specific hypernetwork and a global-sharing imaging network, respectively. The purpose of global-sharing imaging network is to learn stable and effective common features from different institutions. The institution-specific hypernetwork is carefully designed to obtain hyperparameters to condition the global-sharing imaging network for personalized local CT reconstruction. Experiments show that HyperFed achieves competitive performance in CT reconstruction compared with several other state-of-the-art methods. It is believed as a promising direction to improve CT imaging quality and achieve personalized demands of different institutions or scanners without privacy data sharing. The codes will be released at https://github.com/Zi-YuanYang/HyperFed.
MISSU: 3D Medical Image Segmentation via Self-distilling TransUNet
Jun 02, 2022Nan Wang, Shaohui Lin, Xiaoxiao Li, Ke Li, Yunhang Shen, Yue Gao, Lizhuang Ma
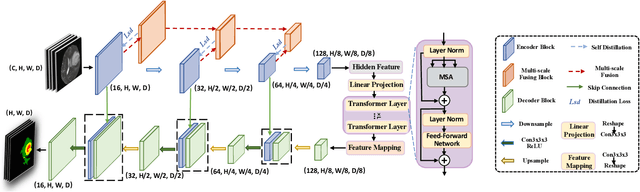
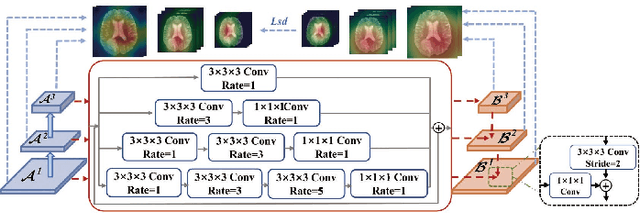
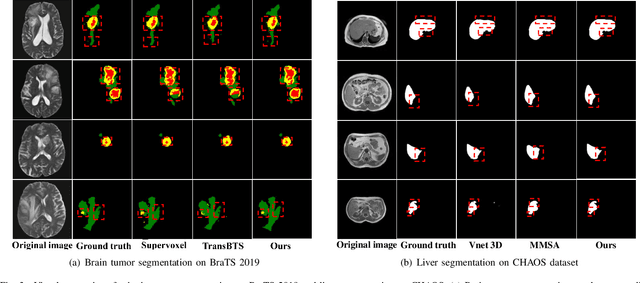
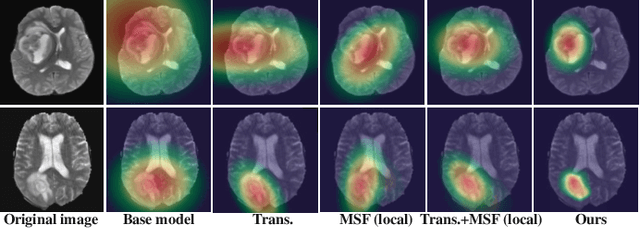
U-Nets have achieved tremendous success in medical image segmentation. Nevertheless, it may suffer limitations in global (long-range) contextual interactions and edge-detail preservation. In contrast, Transformer has an excellent ability to capture long-range dependencies by leveraging the self-attention mechanism into the encoder. Although Transformer was born to model the long-range dependency on the extracted feature maps, it still suffers from extreme computational and spatial complexities in processing high-resolution 3D feature maps. This motivates us to design the efficiently Transformer-based UNet model and study the feasibility of Transformer-based network architectures for medical image segmentation tasks. To this end, we propose to self-distill a Transformer-based UNet for medical image segmentation, which simultaneously learns global semantic information and local spatial-detailed features. Meanwhile, a local multi-scale fusion block is first proposed to refine fine-grained details from the skipped connections in the encoder by the main CNN stem through self-distillation, only computed during training and removed at inference with minimal overhead. Extensive experiments on BraTS 2019 and CHAOS datasets show that our MISSU achieves the best performance over previous state-of-the-art methods. Code and models are available at \url{https://github.com/wangn123/MISSU.git}
Federated Learning from Only Unlabeled Data with Class-Conditional-Sharing Clients
Apr 07, 2022Nan Lu, Zhao Wang, Xiaoxiao Li, Gang Niu, Qi Dou, Masashi Sugiyama
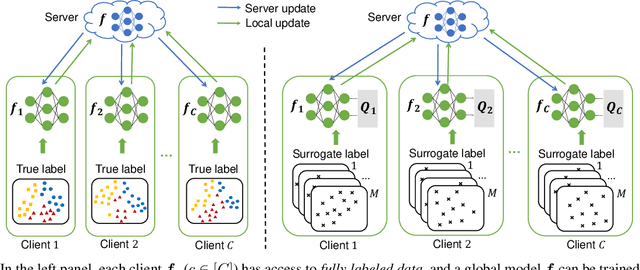
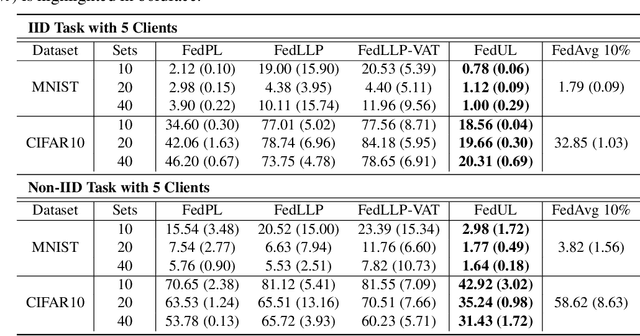
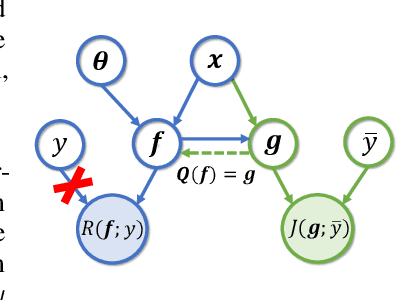
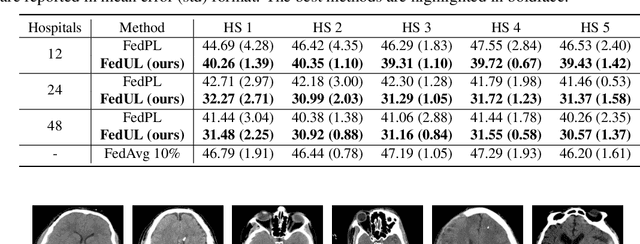
Supervised federated learning (FL) enables multiple clients to share the trained model without sharing their labeled data. However, potential clients might even be reluctant to label their own data, which could limit the applicability of FL in practice. In this paper, we show the possibility of unsupervised FL whose model is still a classifier for predicting class labels, if the class-prior probabilities are shifted while the class-conditional distributions are shared among the unlabeled data owned by the clients. We propose federation of unsupervised learning (FedUL), where the unlabeled data are transformed into surrogate labeled data for each of the clients, a modified model is trained by supervised FL, and the wanted model is recovered from the modified model. FedUL is a very general solution to unsupervised FL: it is compatible with many supervised FL methods, and the recovery of the wanted model can be theoretically guaranteed as if the data have been labeled. Experiments on benchmark and real-world datasets demonstrate the effectiveness of FedUL. Code is available at https://github.com/lunanbit/FedUL.
GATE: Graph CCA for Temporal SElf-supervised Learning for Label-efficient fMRI Analysis
Mar 17, 2022Liang Peng, Nan Wang, Jie Xu, Xiaofeng Zhu, Xiaoxiao Li
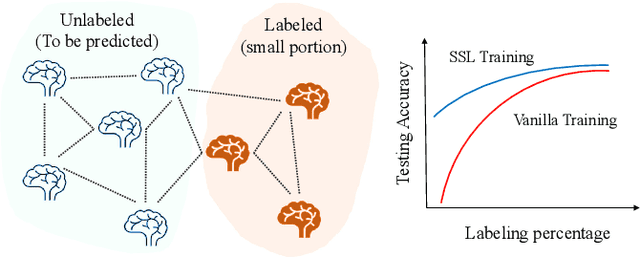
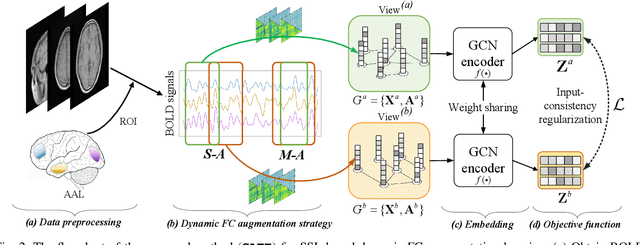
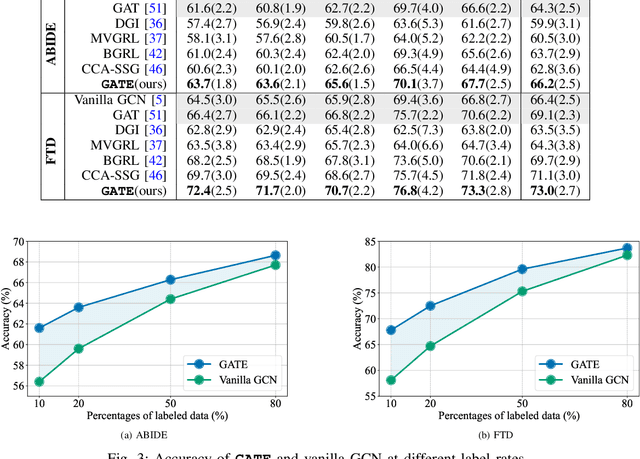
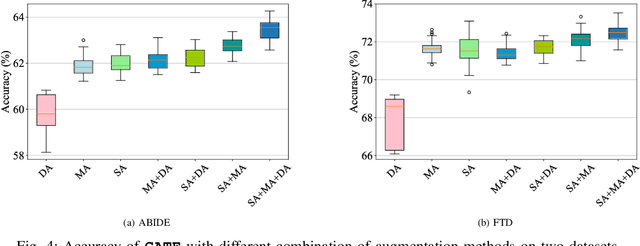
In this work, we focus on the challenging task, neuro-disease classification, using functional magnetic resonance imaging (fMRI). In population graph-based disease analysis, graph convolutional neural networks (GCNs) have achieved remarkable success. However, these achievements are inseparable from abundant labeled data and sensitive to spurious signals. To improve fMRI representation learning and classification under a label-efficient setting, we propose a novel and theory-driven self-supervised learning (SSL) framework on GCNs, namely Graph CCA for Temporal self-supervised learning on fMRI analysis GATE. Concretely, it is demanding to design a suitable and effective SSL strategy to extract formation and robust features for fMRI. To this end, we investigate several new graph augmentation strategies from fMRI dynamic functional connectives (FC) for SSL training. Further, we leverage canonical-correlation analysis (CCA) on different temporal embeddings and present the theoretical implications. Consequently, this yields a novel two-step GCN learning procedure comprised of (i) SSL on an unlabeled fMRI population graph and (ii) fine-tuning on a small labeled fMRI dataset for a classification task. Our method is tested on two independent fMRI datasets, demonstrating superior performance on autism and dementia diagnosis.
Evaluating Explainable AI on a Multi-Modal Medical Imaging Task: Can Existing Algorithms Fulfill Clinical Requirements?
Mar 12, 2022Weina Jin, Xiaoxiao Li, Ghassan Hamarneh
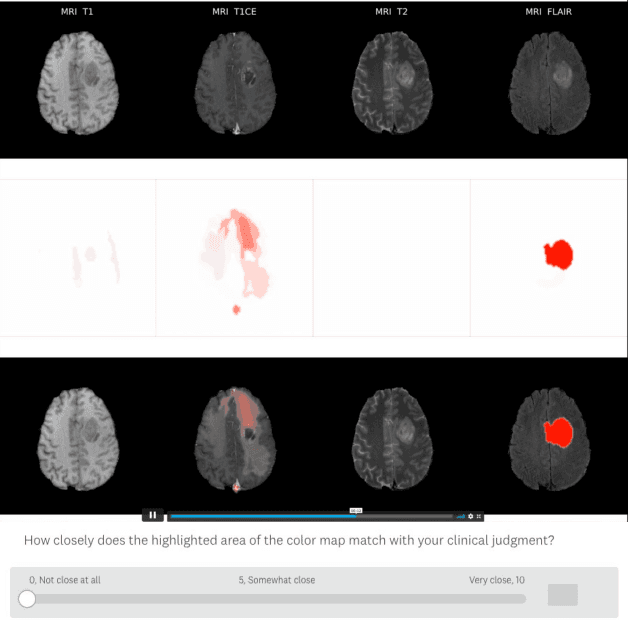
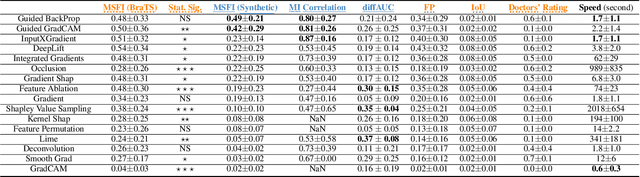
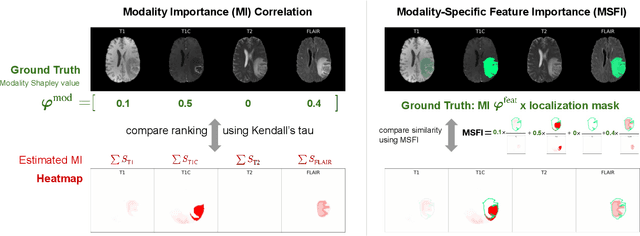
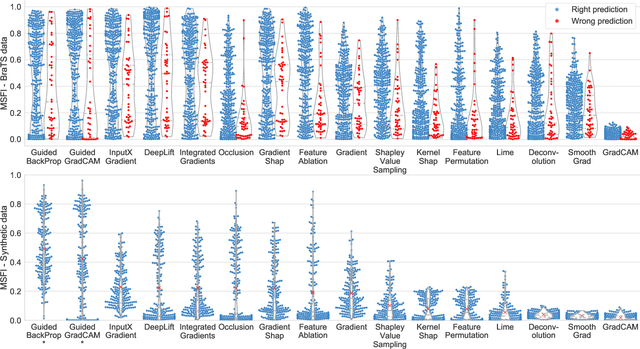
Being able to explain the prediction to clinical end-users is a necessity to leverage the power of artificial intelligence (AI) models for clinical decision support. For medical images, a feature attribution map, or heatmap, is the most common form of explanation that highlights important features for AI models' prediction. However, it is unknown how well heatmaps perform on explaining decisions on multi-modal medical images, where each image modality or channel visualizes distinct clinical information of the same underlying biomedical phenomenon. Understanding such modality-dependent features is essential for clinical users' interpretation of AI decisions. To tackle this clinically important but technically ignored problem, we propose the modality-specific feature importance (MSFI) metric. It encodes clinical image and explanation interpretation patterns of modality prioritization and modality-specific feature localization. We conduct a clinical requirement-grounded, systematic evaluation using computational methods and a clinician user study. Results show that the examined 16 heatmap algorithms failed to fulfill clinical requirements to correctly indicate AI model decision process or decision quality. The evaluation and MSFI metric can guide the design and selection of XAI algorithms to meet clinical requirements on multi-modal explanation.
Guidelines and evaluation for clinical explainable AI on medical image analysis
Feb 16, 2022Weina Jin, Xiaoxiao Li, Mostafa Fatehi, Ghassan Hamarneh
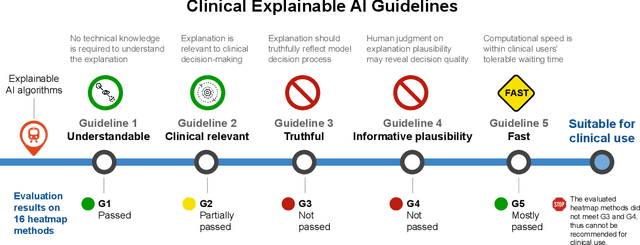
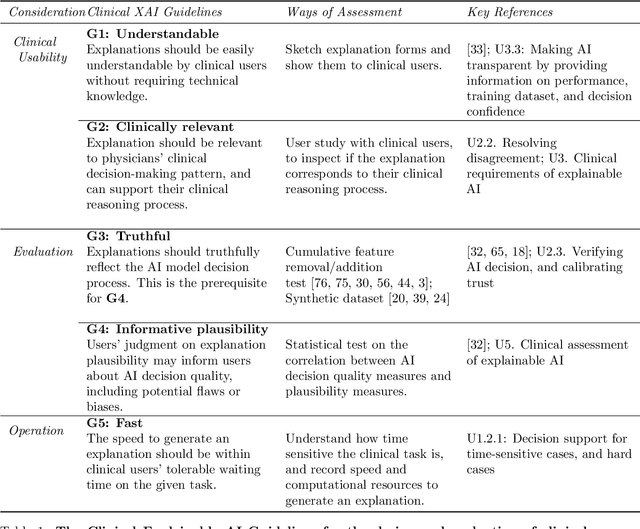
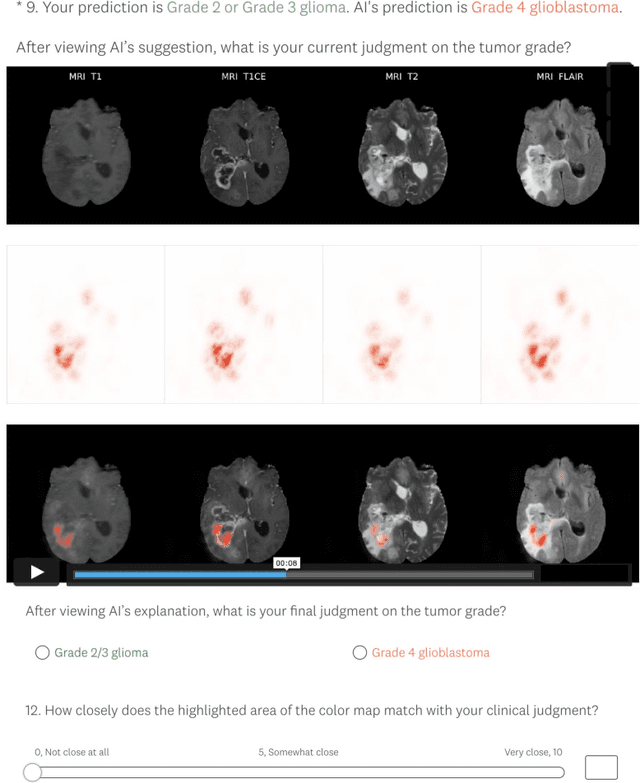
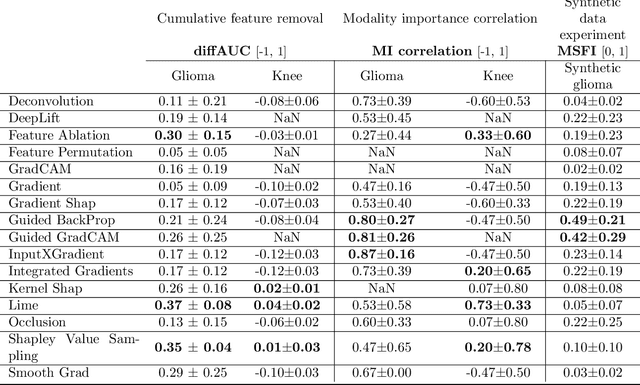
Explainable artificial intelligence (XAI) is essential for enabling clinical users to get informed decision support from AI and comply with evidence-based medical practice. Applying XAI in clinical settings requires proper evaluation criteria to ensure the explanation technique is both technically sound and clinically useful, but specific support is lacking to achieve this goal. To bridge the research gap, we propose the Clinical XAI Guidelines that consist of five criteria a clinical XAI needs to be optimized for. The guidelines recommend choosing an explanation form based on Guideline 1 (G1) Understandability and G2 Clinical relevance. For the chosen explanation form, its specific XAI technique should be optimized for G3 Truthfulness, G4 Informative plausibility, and G5 Computational efficiency. Following the guidelines, we conducted a systematic evaluation on a novel problem of multi-modal medical image explanation with two clinical tasks, and proposed new evaluation metrics accordingly. The evaluated 16 commonly-used heatmap XAI techniques were not suitable for clinical use due to their failure in \textbf{G3} and \textbf{G4}. Our evaluation demonstrated the use of Clinical XAI Guidelines to support the design and evaluation for clinically viable XAI.
RestoreDet: Degradation Equivariant Representation for Object Detection in Low Resolution Images
Jan 07, 2022Ziteng Cui, Yingying Zhu, Lin Gu, Guo-Jun Qi, Xiaoxiao Li, Peng Gao, Zenghui Zhang, Tatsuya Harada
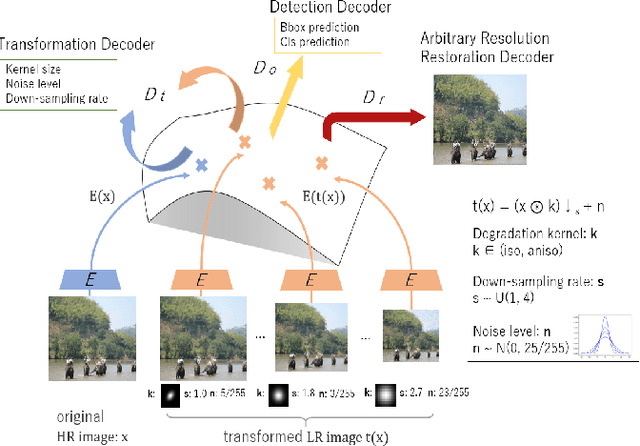
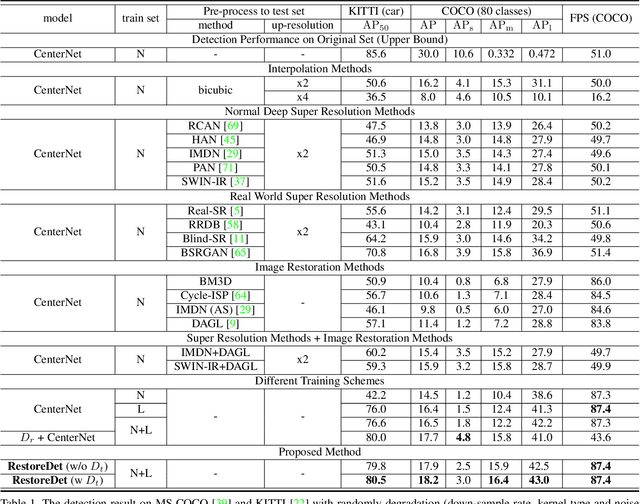
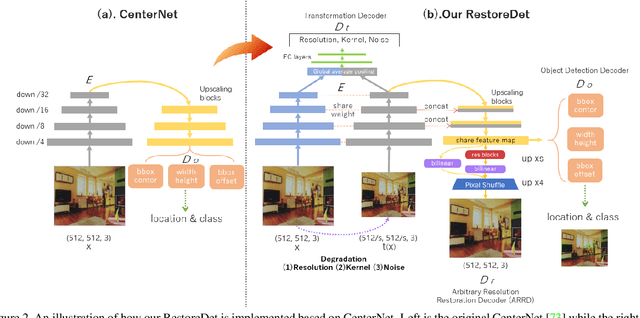
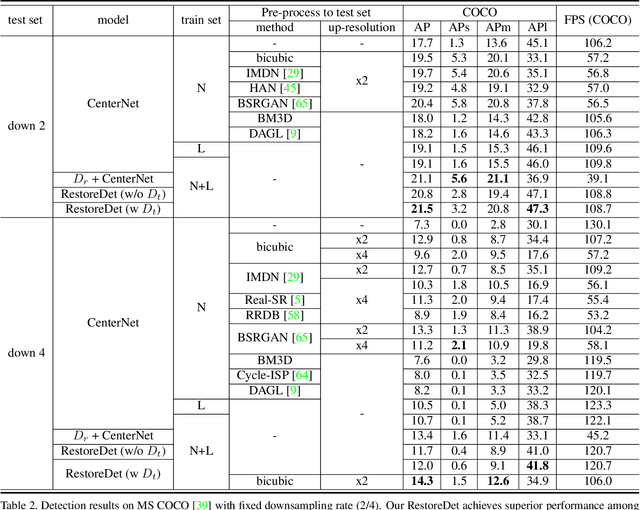
Image restoration algorithms such as super resolution (SR) are indispensable pre-processing modules for object detection in degraded images. However, most of these algorithms assume the degradation is fixed and known a priori. When the real degradation is unknown or differs from assumption, both the pre-processing module and the consequent high-level task such as object detection would fail. Here, we propose a novel framework, RestoreDet, to detect objects in degraded low resolution images. RestoreDet utilizes the downsampling degradation as a kind of transformation for self-supervised signals to explore the equivariant representation against various resolutions and other degradation conditions. Specifically, we learn this intrinsic visual structure by encoding and decoding the degradation transformation from a pair of original and randomly degraded images. The framework could further take the advantage of advanced SR architectures with an arbitrary resolution restoring decoder to reconstruct the original correspondence from the degraded input image. Both the representation learning and object detection are optimized jointly in an end-to-end training fashion. RestoreDet is a generic framework that could be implemented on any mainstream object detection architectures. The extensive experiment shows that our framework based on CenterNet has achieved superior performance compared with existing methods when facing variant degradation situations. Our code would be released soon.
FedNI: Federated Graph Learning with Network Inpainting for Population-Based Disease Prediction
Dec 19, 2021Liang Peng, Nan Wang, Nicha Dvornek, Xiaofeng Zhu, Xiaoxiao Li
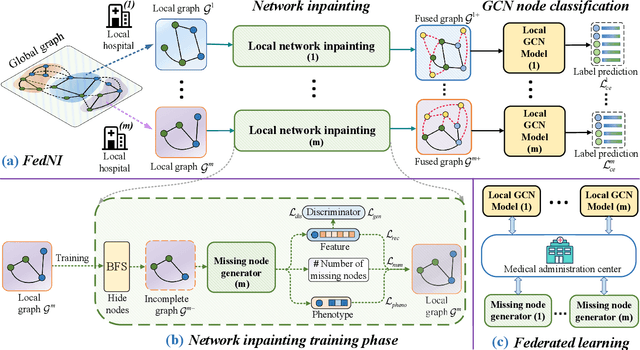
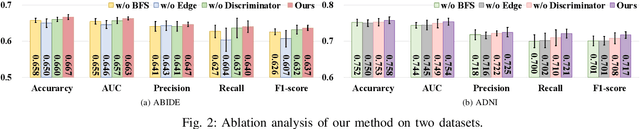
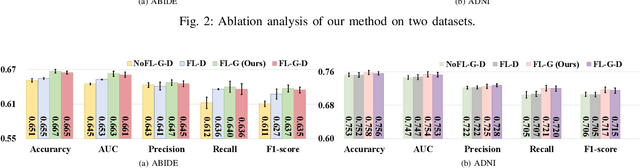
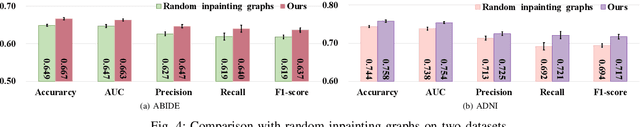
Graph Convolutional Neural Networks (GCNs) are widely used for graph analysis. Specifically, in medical applications, GCNs can be used for disease prediction on a population graph, where graph nodes represent individuals and edges represent individual similarities. However, GCNs rely on a vast amount of data, which is challenging to collect for a single medical institution. In addition, a critical challenge that most medical institutions continue to face is addressing disease prediction in isolation with incomplete data information. To address these issues, Federated Learning (FL) allows isolated local institutions to collaboratively train a global model without data sharing. In this work, we propose a framework, FedNI, to leverage network inpainting and inter-institutional data via FL. Specifically, we first federatively train missing node and edge predictor using a graph generative adversarial network (GAN) to complete the missing information of local networks. Then we train a global GCN node classifier across institutions using a federated graph learning platform. The novel design enables us to build more accurate machine learning models by leveraging federated learning and also graph learning approaches. We demonstrate that our federated model outperforms local and baseline FL methods with significant margins on two public neuroimaging datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge