Thanuja D. Ambegoda
Air Signing and Privacy-Preserving Signature Verification for Digital Documents
May 17, 2024



Abstract:This paper presents a novel approach to the digital signing of electronic documents through the use of a camera-based interaction system, single-finger tracking for sign recognition, and multi commands executing hand gestures. The proposed solution, referred to as "Air Signature," involves writing the signature in front of the camera, rather than relying on traditional methods such as mouse drawing or physically signing on paper and showing it to a web camera. The goal is to develop a state-of-the-art method for detecting and tracking gestures and objects in real-time. The proposed methods include applying existing gesture recognition and object tracking systems, improving accuracy through smoothing and line drawing, and maintaining continuity during fast finger movements. An evaluation of the fingertip detection, sketching, and overall signing process is performed to assess the effectiveness of the proposed solution. The secondary objective of this research is to develop a model that can effectively recognize the unique signature of a user. This type of signature can be verified by neural cores that analyze the movement, speed, and stroke pixels of the signing in real time. The neural cores use machine learning algorithms to match air signatures to the individual's stored signatures, providing a secure and efficient method of verification. Our proposed System does not require sensors or any hardware other than the camera.
Estimation of Z-Thickness and XY-Anisotropy of Electron Microscopy Images using Gaussian Processes
Feb 04, 2020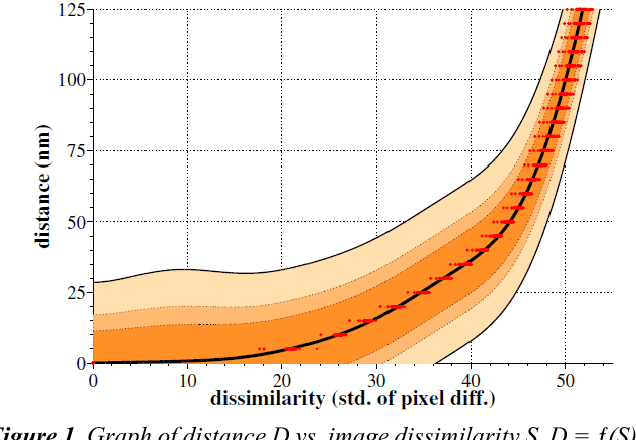
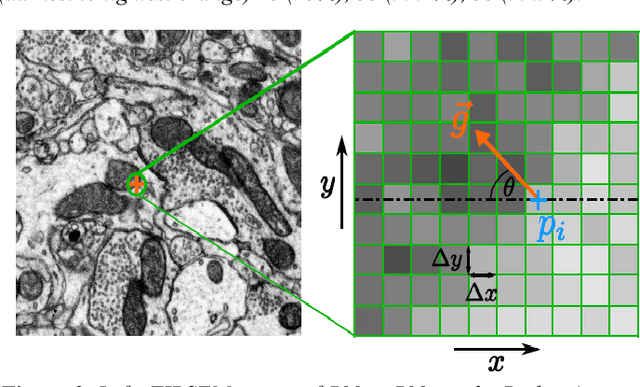

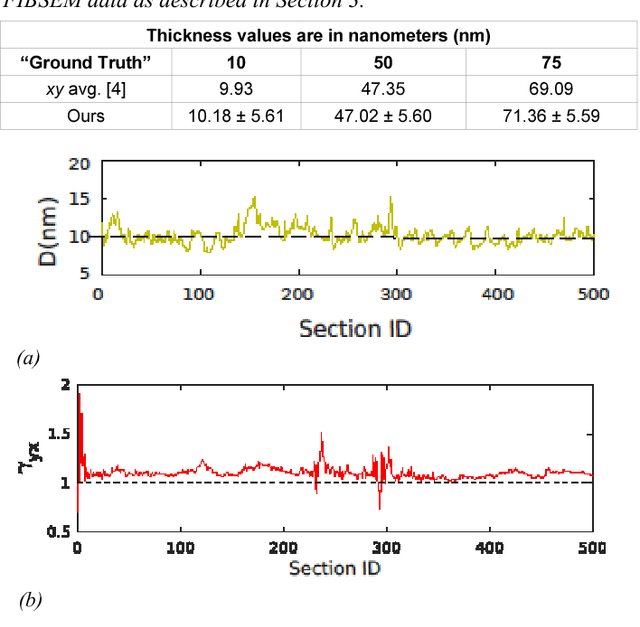
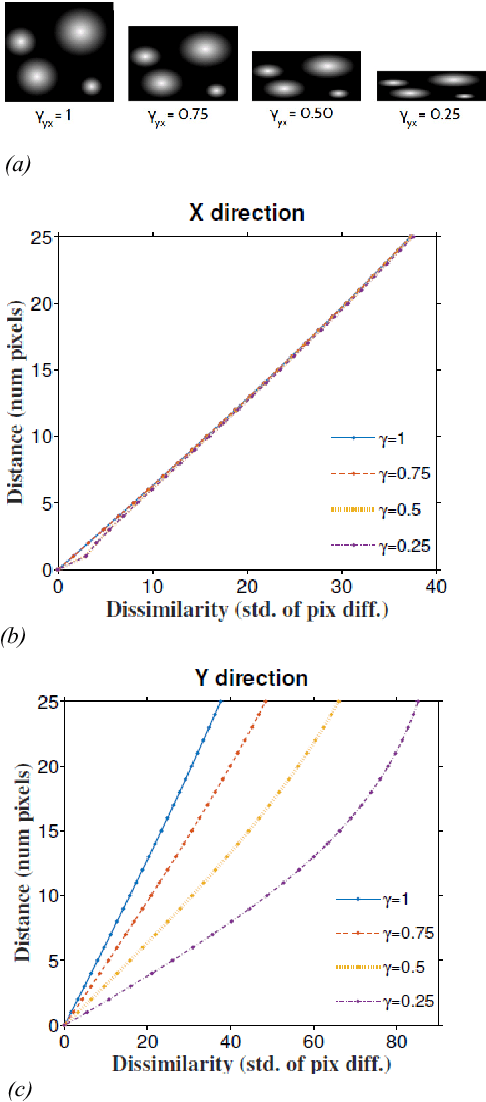
Abstract:Serial section electron microscopy (ssEM) is a widely used technique for obtaining volumetric information of biological tissues at nanometer scale. However, accurate 3D reconstructions of identified cellular structures and volumetric quantifications require precise estimates of section thickness and anisotropy (or stretching) along the XY imaging plane. In fact, many image processing algorithms simply assume isotropy within the imaging plane. To ameliorate this problem, we present a method for estimating thickness and stretching of electron microscopy sections using non-parametric Bayesian regression of image statistics. We verify our thickness and stretching estimates using direct measurements obtained by atomic force microscopy (AFM) and show that our method has a lower estimation error compared to a recent indirect thickness estimation method as well as a relative Z coordinate estimation method. Furthermore, we have made the first dataset of ssSEM images with directly measured section thickness values publicly available for the evaluation of indirect thickness estimation methods.
Efficient 2D neuron boundary segmentation with local topological constraints
Feb 03, 2020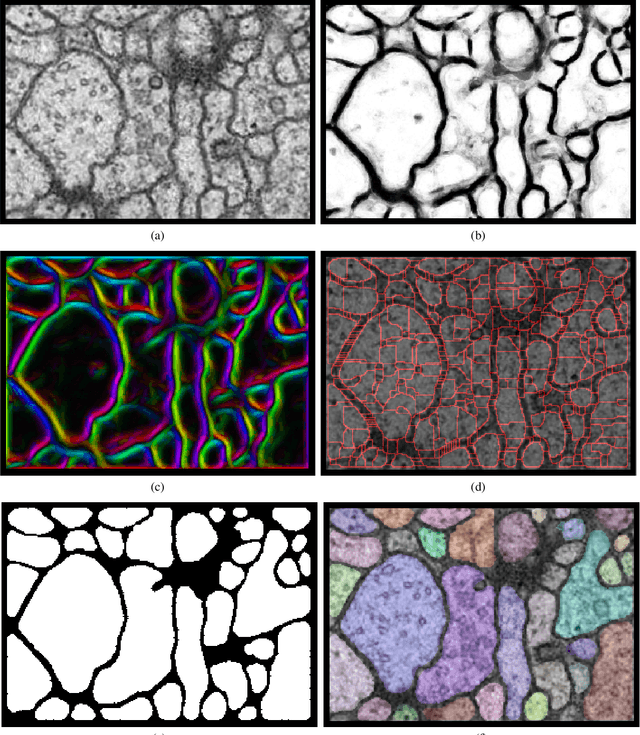
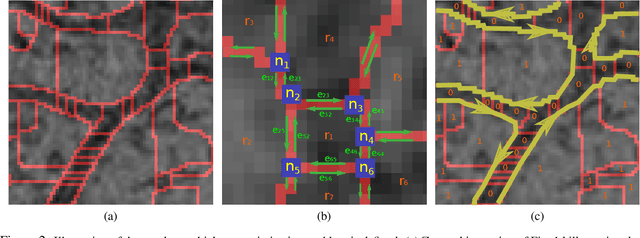
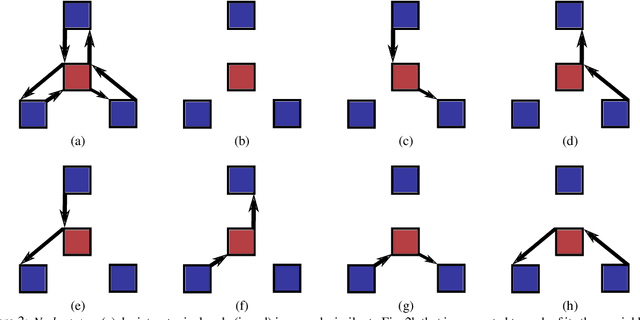

Abstract:We present a method for segmenting neuron membranes in 2D electron microscopy imagery. This segmentation task has been a bottleneck to reconstruction efforts of the brain's synaptic circuits. One common problem is the misclassification of blurry membrane fragments as cell interior, which leads to merging of two adjacent neuron sections into one via the blurry membrane region. Human annotators can easily avoid such errors by implicitly performing gap completion, taking into account the continuity of membranes. Drawing inspiration from these human strategies, we formulate the segmentation task as an edge labeling problem on a graph with local topological constraints. We derive an integer linear program (ILP) that enforces membrane continuity, i.e. the absence of gaps. The cost function of the ILP is the pixel-wise deviation of the segmentation from a priori membrane probabilities derived from the data. Based on membrane probability maps obtained using random forest classifiers and convolutional neural networks, our method improves the neuron boundary segmentation accuracy compared to a variety of standard segmentation approaches. Our method successfully performs gap completion and leads to fewer topological errors. The method could potentially also be incorporated into other image segmentation pipelines with known topological constraints.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge