A Framework for Individualizing Predictions of Disease Trajectories by Exploiting Multi-Resolution Structure
Jan 21, 2016Peter Schulam, Suchi Saria
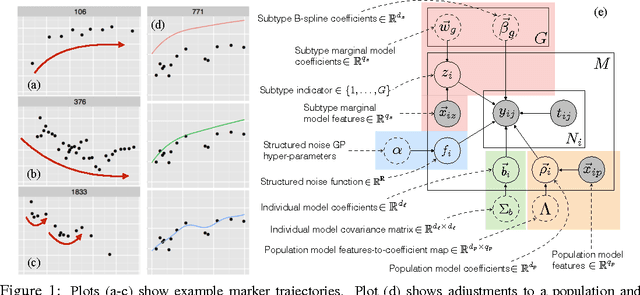
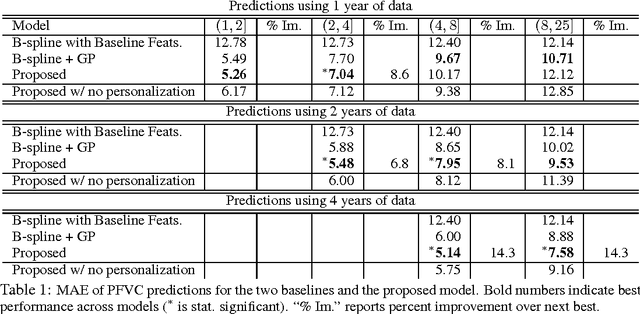
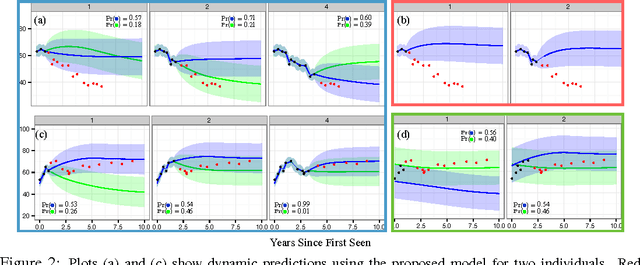
For many complex diseases, there is a wide variety of ways in which an individual can manifest the disease. The challenge of personalized medicine is to develop tools that can accurately predict the trajectory of an individual's disease, which can in turn enable clinicians to optimize treatments. We represent an individual's disease trajectory as a continuous-valued continuous-time function describing the severity of the disease over time. We propose a hierarchical latent variable model that individualizes predictions of disease trajectories. This model shares statistical strength across observations at different resolutions--the population, subpopulation and the individual level. We describe an algorithm for learning population and subpopulation parameters offline, and an online procedure for dynamically learning individual-specific parameters. Finally, we validate our model on the task of predicting the course of interstitial lung disease, a leading cause of death among patients with the autoimmune disease scleroderma. We compare our approach against state-of-the-art and demonstrate significant improvements in predictive accuracy.
Deformable Distributed Multiple Detector Fusion for Multi-Person Tracking
Dec 18, 2015Andy J Ma, Pong C Yuen, Suchi Saria
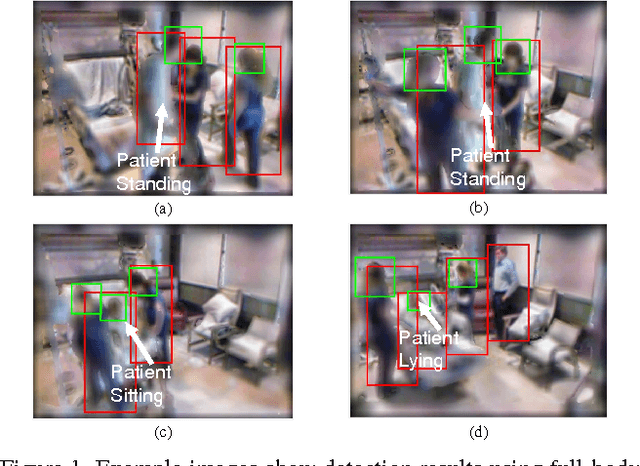
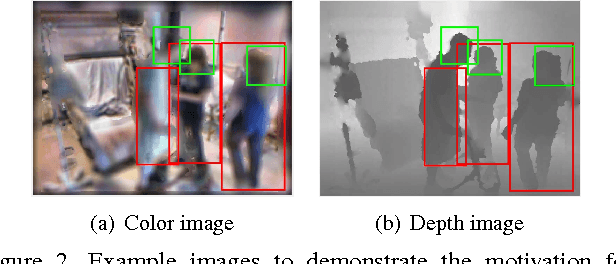

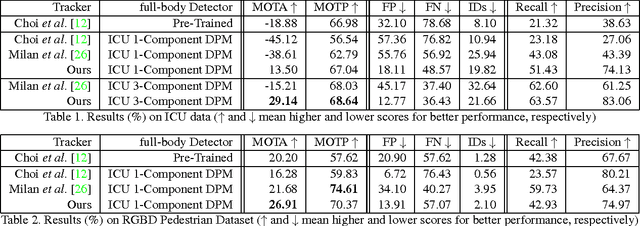


This paper addresses fully automated multi-person tracking in complex environments with challenging occlusion and extensive pose variations. Our solution combines multiple detectors for a set of different regions of interest (e.g., full-body and head) for multi-person tracking. The use of multiple detectors leads to fewer miss detections as it is able to exploit the complementary strengths of the individual detectors. While the number of false positives may increase with the increased number of bounding boxes detected from multiple detectors, we propose to group the detection outputs by bounding box location and depth information. For robustness to significant pose variations, deformable spatial relationship between detectors are learnt in our multi-person tracking system. On RGBD data from a live Intensive Care Unit (ICU), we show that the proposed method significantly improves multi-person tracking performance over state-of-the-art methods.
Learning (Predictive) Risk Scores in the Presence of Censoring due to Interventions
Jul 27, 2015Kirill Dyagilev, Suchi Saria
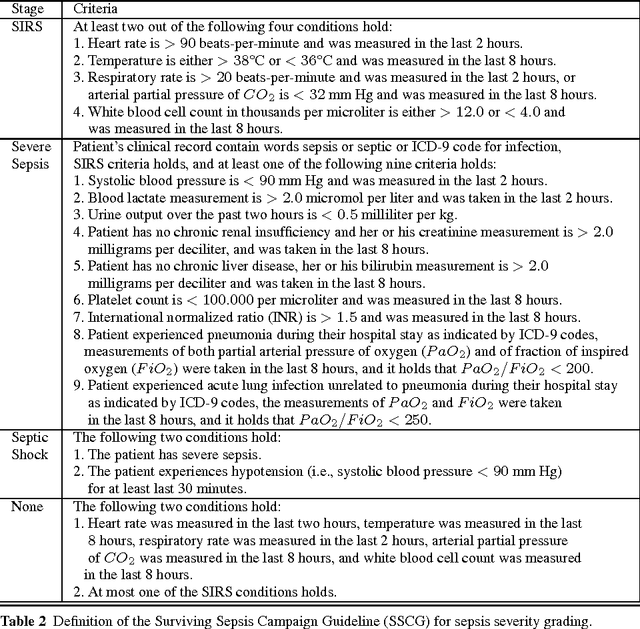
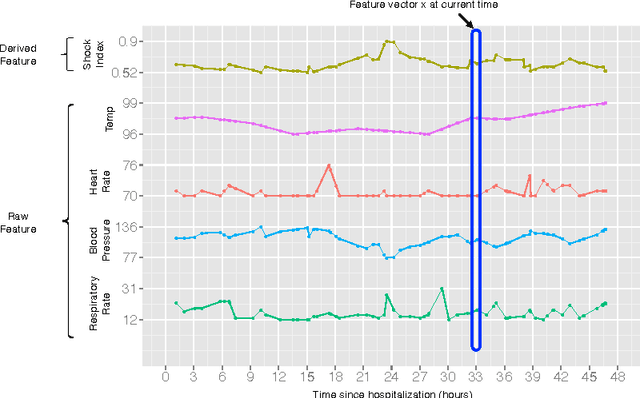
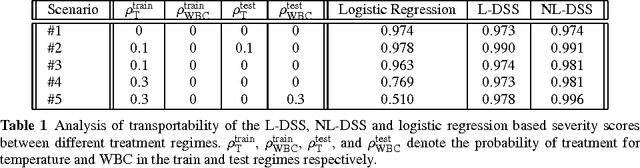
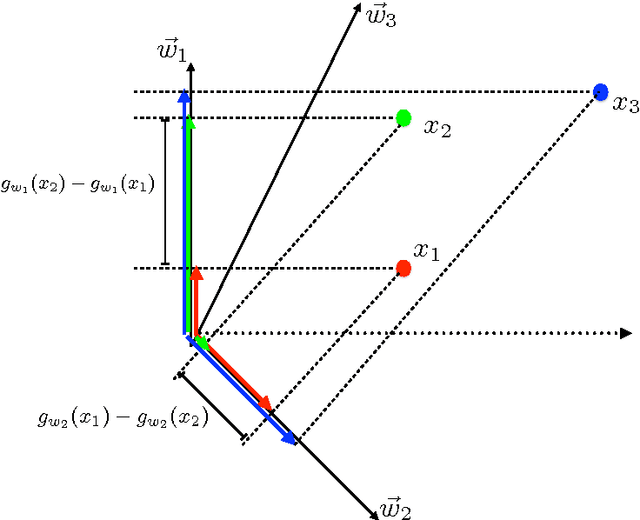

A large and diverse set of measurements are regularly collected during a patient's hospital stay to monitor their health status. Tools for integrating these measurements into severity scores, that accurately track changes in illness severity, can improve clinicians ability to provide timely interventions. Existing approaches for creating such scores either 1) rely on experts to fully specify the severity score, or 2) train a predictive score, using supervised learning, by regressing against a surrogate marker of severity such as the presence of downstream adverse events. The first approach does not extend to diseases where an accurate score cannot be elicited from experts. The second approach often produces scores that suffer from bias due to treatment-related censoring (Paxton, 2013). We propose a novel ranking based framework for disease severity score learning (DSSL). DSSL exploits the following key observation: while it is challenging for experts to quantify the disease severity at any given time, it is often easy to compare the disease severity at two different times. Extending existing ranking algorithms, DSSL learns a function that maps a vector of patient's measurements to a scalar severity score such that the resulting score is temporally smooth and consistent with the expert's ranking of pairs of disease states. We apply DSSL to the problem of learning a sepsis severity score using a large, real-world dataset. The learned scores significantly outperform state-of-the-art clinical scores in ranking patient states by severity and in early detection of future adverse events. We also show that the learned disease severity trajectories are consistent with clinical expectations of disease evolution. Further, using simulated datasets, we show that DSSL exhibits better generalization performance to changes in treatment patterns compared to the above approaches.
Reasoning at the Right Time Granularity
Jun 20, 2012Suchi Saria, Uri Nodelman, Daphne Koller
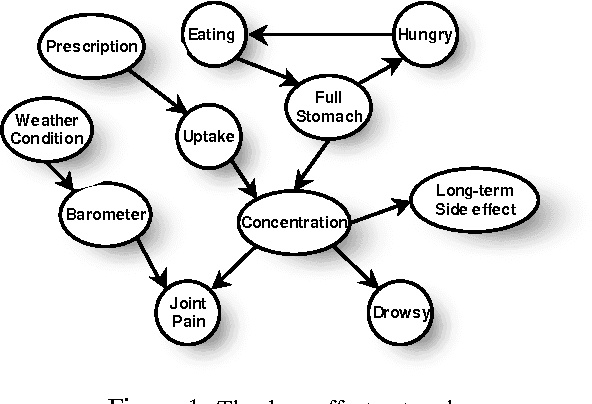
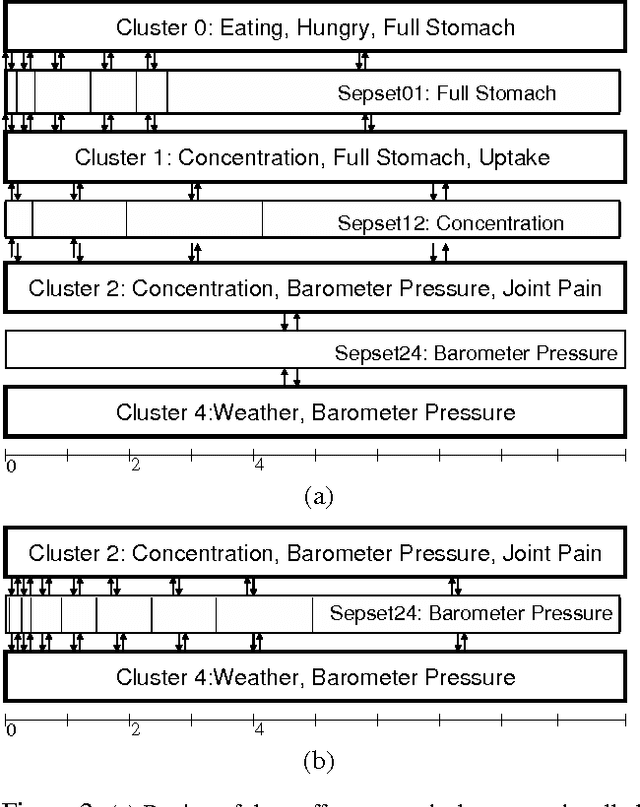
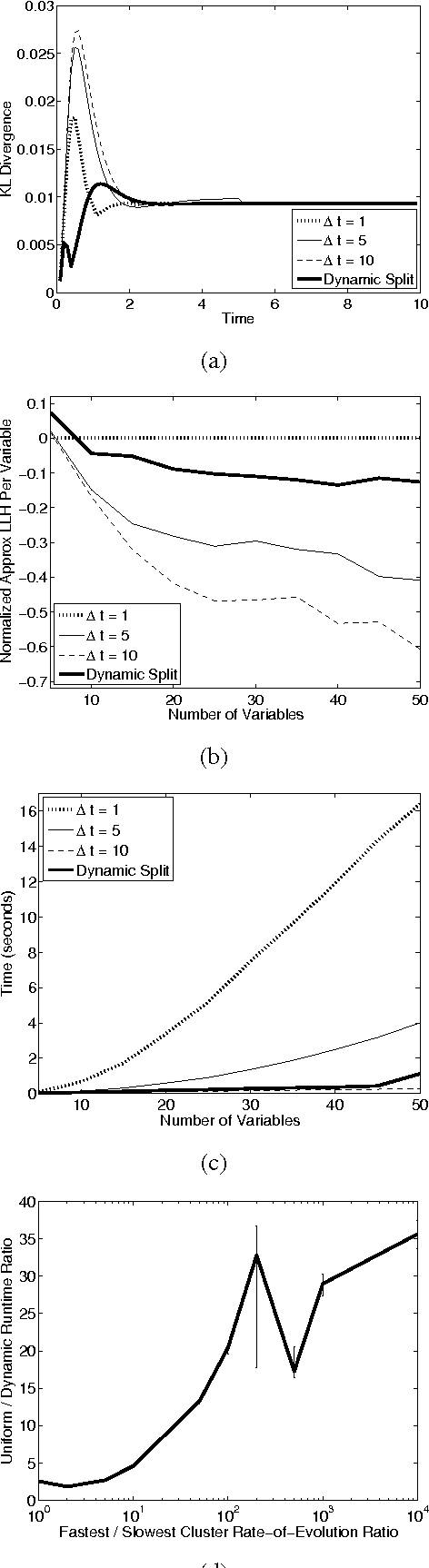
Most real-world dynamic systems are composed of different components that often evolve at very different rates. In traditional temporal graphical models, such as dynamic Bayesian networks, time is modeled at a fixed granularity, generally selected based on the rate at which the fastest component evolves. Inference must then be performed at this fastest granularity, potentially at significant computational cost. Continuous Time Bayesian Networks (CTBNs) avoid time-slicing in the representation by modeling the system as evolving continuously over time. The expectation-propagation (EP) inference algorithm of Nodelman et al. (2005) can then vary the inference granularity over time, but the granularity is uniform across all parts of the system, and must be selected in advance. In this paper, we provide a new EP algorithm that utilizes a general cluster graph architecture where clusters contain distributions that can overlap in both space (set of variables) and time. This architecture allows different parts of the system to be modeled at very different time granularities, according to their current rate of evolution. We also provide an information-theoretic criterion for dynamically re-partitioning the clusters during inference to tune the level of approximation to the current rate of evolution. This avoids the need to hand-select the appropriate granularity, and allows the granularity to adapt as information is transmitted across the network. We present experiments demonstrating that this approach can result in significant computational savings.
Discovering shared and individual latent structure in multiple time series
Aug 12, 2010Suchi Saria, Daphne Koller, Anna Penn
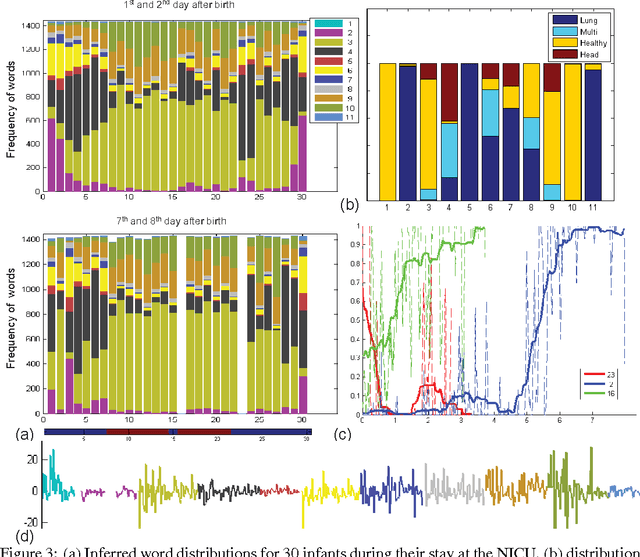
This paper proposes a nonparametric Bayesian method for exploratory data analysis and feature construction in continuous time series. Our method focuses on understanding shared features in a set of time series that exhibit significant individual variability. Our method builds on the framework of latent Diricihlet allocation (LDA) and its extension to hierarchical Dirichlet processes, which allows us to characterize each series as switching between latent ``topics'', where each topic is characterized as a distribution over ``words'' that specify the series dynamics. However, unlike standard applications of LDA, we discover the words as we learn the model. We apply this model to the task of tracking the physiological signals of premature infants; our model obtains clinically significant insights as well as useful features for supervised learning tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge