Shohei Shimizu
Bayesian estimation of possible causal direction in the presence of latent confounders using a linear non-Gaussian acyclic structural equation model with individual-specific effects
May 20, 2014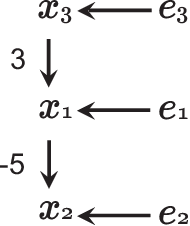
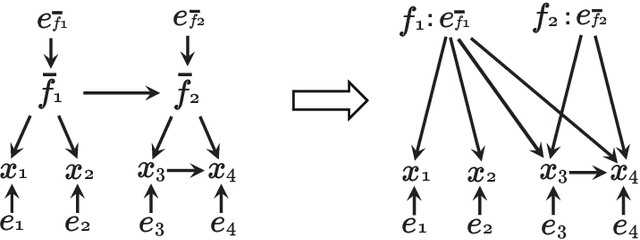

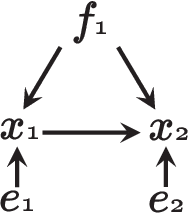
Abstract:We consider learning the possible causal direction of two observed variables in the presence of latent confounding variables. Several existing methods have been shown to consistently estimate causal direction assuming linear or some type of nonlinear relationship and no latent confounders. However, the estimation results could be distorted if either assumption is actually violated. In this paper, we first propose a new linear non-Gaussian acyclic structural equation model with individual-specific effects that allows latent confounders to be considered. We then propose an empirical Bayesian approach for estimating possible causal direction using the new model. We demonstrate the effectiveness of our method using artificial and real-world data.
Causal Discovery in a Binary Exclusive-or Skew Acyclic Model: BExSAM
Jan 22, 2014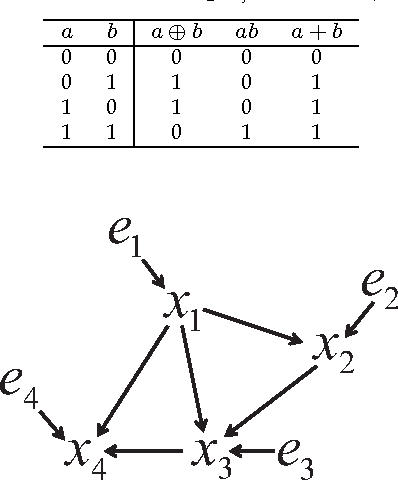
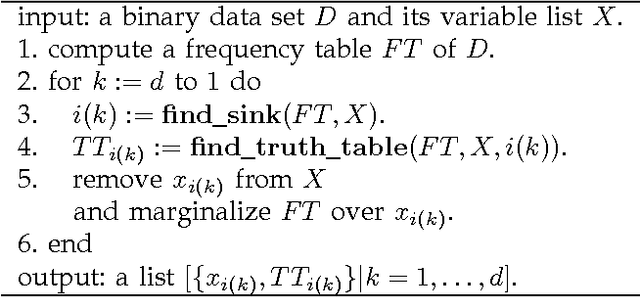
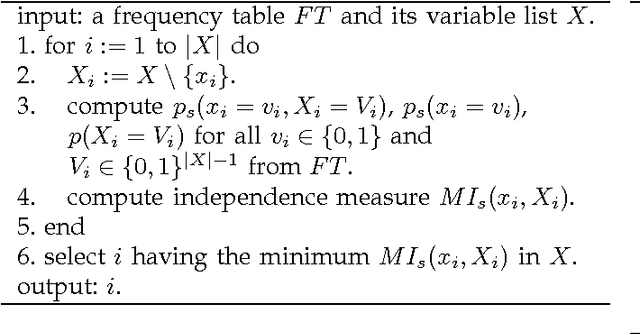

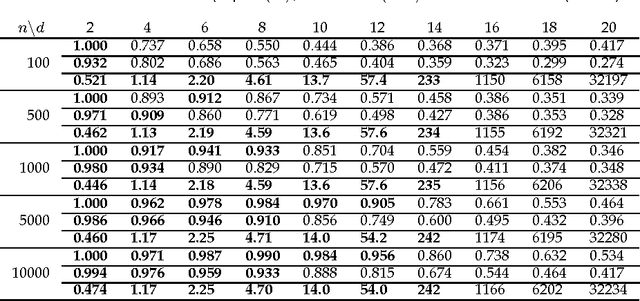

Abstract:Discovering causal relations among observed variables in a given data set is a major objective in studies of statistics and artificial intelligence. Recently, some techniques to discover a unique causal model have been explored based on non-Gaussianity of the observed data distribution. However, most of these are limited to continuous data. In this paper, we present a novel causal model for binary data and propose an efficient new approach to deriving the unique causal model governing a given binary data set under skew distributions of external binary noises. Experimental evaluation shows excellent performance for both artificial and real world data sets.
Identifiability of an Integer Modular Acyclic Additive Noise Model and its Causal Structure Discovery
Jan 22, 2014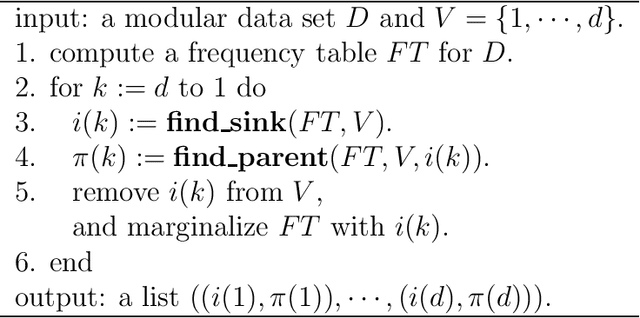
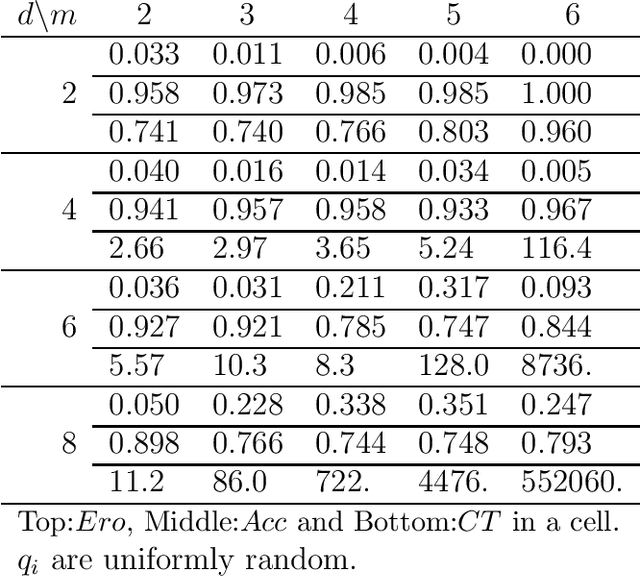
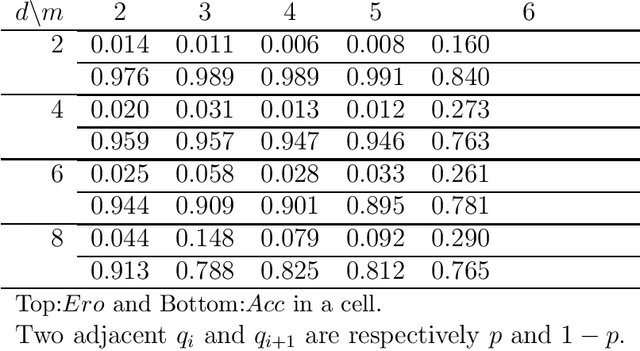
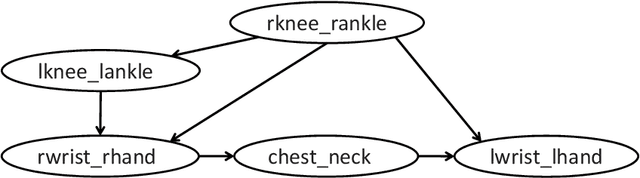
Abstract:The notion of causality is used in many situations dealing with uncertainty. We consider the problem whether causality can be identified given data set generated by discrete random variables rather than continuous ones. In particular, for non-binary data, thus far it was only known that causality can be identified except rare cases. In this paper, we present necessary and sufficient condition for an integer modular acyclic additive noise (IMAN) of two variables. In addition, we relate bivariate and multivariate causal identifiability in a more explicit manner, and develop a practical algorithm to find the order of variables and their parent sets. We demonstrate its performance in applications to artificial data and real world body motion data with comparisons to conventional methods.
ParceLiNGAM: A causal ordering method robust against latent confounders
Jul 29, 2013Abstract:We consider learning a causal ordering of variables in a linear non-Gaussian acyclic model called LiNGAM. Several existing methods have been shown to consistently estimate a causal ordering assuming that all the model assumptions are correct. But, the estimation results could be distorted if some assumptions actually are violated. In this paper, we propose a new algorithm for learning causal orders that is robust against one typical violation of the model assumptions: latent confounders. The key idea is to detect latent confounders by testing independence between estimated external influences and find subsets (parcels) that include variables that are not affected by latent confounders. We demonstrate the effectiveness of our method using artificial data and simulated brain imaging data.
Learning LiNGAM based on data with more variables than observations
Aug 21, 2012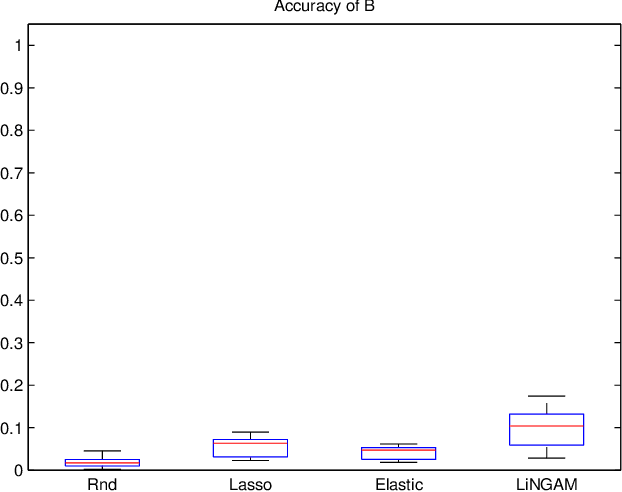
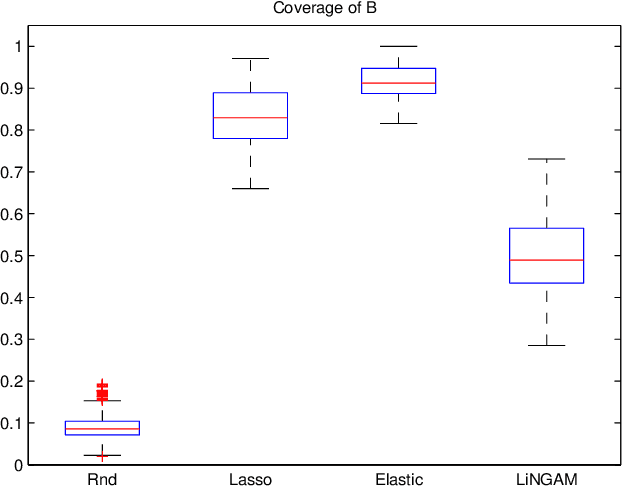
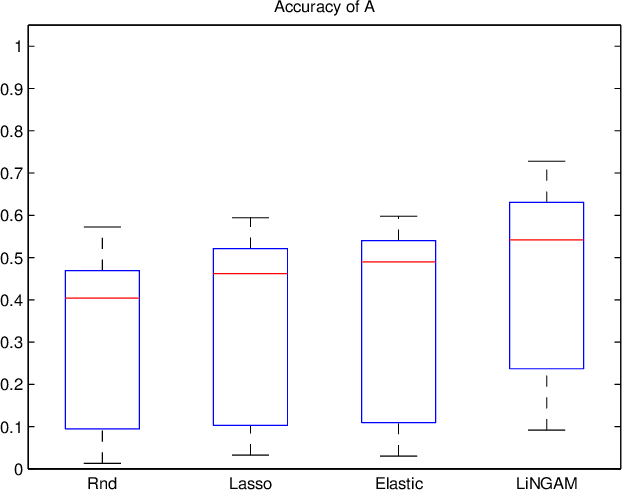
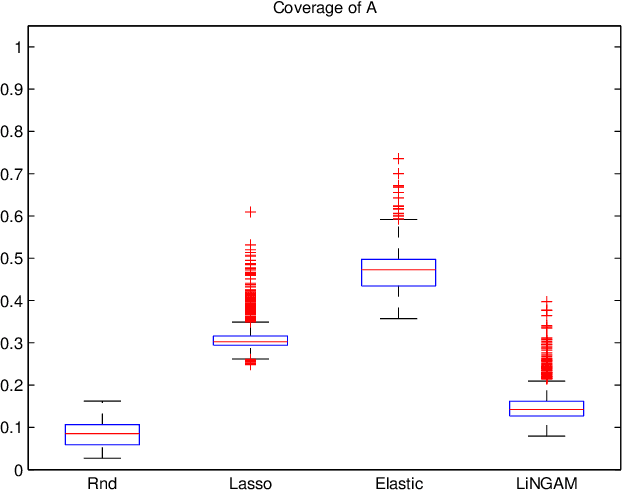
Abstract:A very important topic in systems biology is developing statistical methods that automatically find causal relations in gene regulatory networks with no prior knowledge of causal connectivity. Many methods have been developed for time series data. However, discovery methods based on steady-state data are often necessary and preferable since obtaining time series data can be more expensive and/or infeasible for many biological systems. A conventional approach is causal Bayesian networks. However, estimation of Bayesian networks is ill-posed. In many cases it cannot uniquely identify the underlying causal network and only gives a large class of equivalent causal networks that cannot be distinguished between based on the data distribution. We propose a new discovery algorithm for uniquely identifying the underlying causal network of genes. To the best of our knowledge, the proposed method is the first algorithm for learning gene networks based on a fully identifiable causal model called LiNGAM. We here compare our algorithm with competing algorithms using artificially-generated data, although it is definitely better to test it based on real microarray gene expression data.
Discovery of non-gaussian linear causal models using ICA
Jul 04, 2012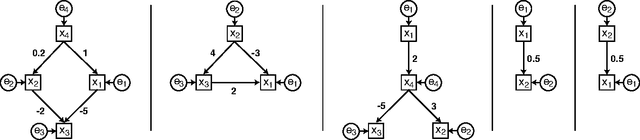
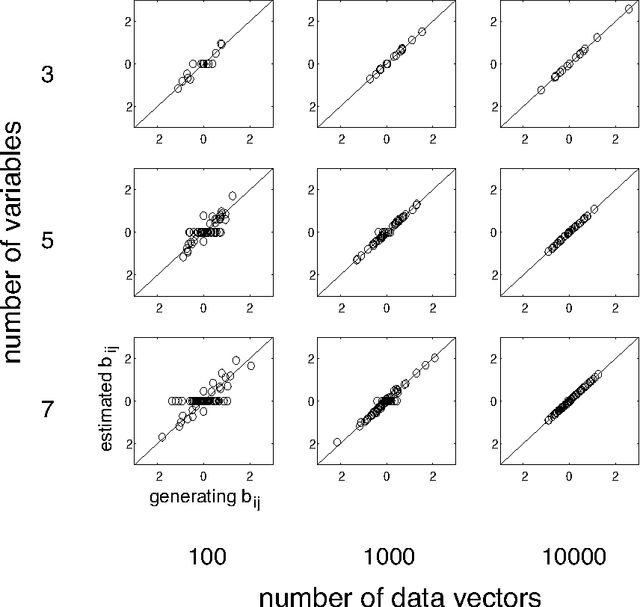
Abstract:In recent years, several methods have been proposed for the discovery of causal structure from non-experimental data (Spirtes et al. 2000; Pearl 2000). Such methods make various assumptions on the data generating process to facilitate its identification from purely observational data. Continuing this line of research, we show how to discover the complete causal structure of continuous-valued data, under the assumptions that (a) the data generating process is linear, (b) there are no unobserved confounders, and (c) disturbance variables have non-gaussian distributions of non-zero variances. The solution relies on the use of the statistical method known as independent component analysis (ICA), and does not require any pre-specified time-ordering of the variables. We provide a complete Matlab package for performing this LiNGAM analysis (short for Linear Non-Gaussian Acyclic Model), and demonstrate the effectiveness of the method using artificially generated data.
Causal discovery of linear acyclic models with arbitrary distributions
Jun 13, 2012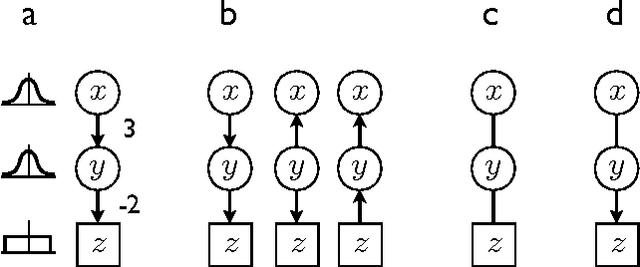
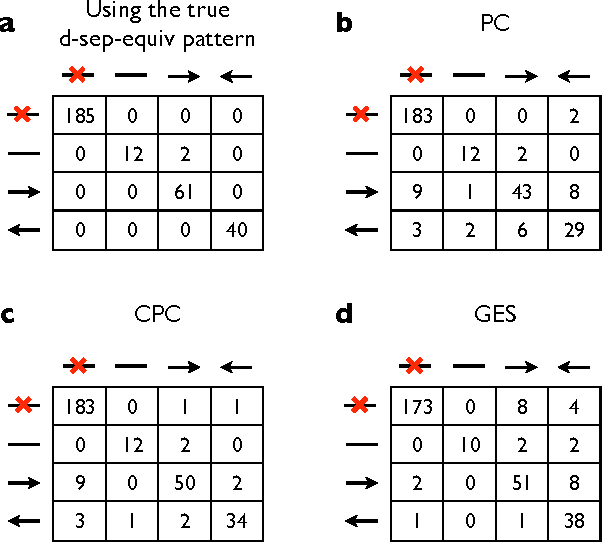
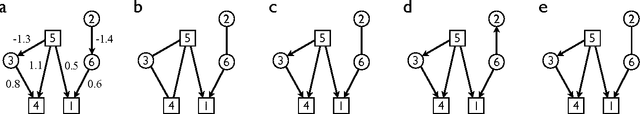
Abstract:An important task in data analysis is the discovery of causal relationships between observed variables. For continuous-valued data, linear acyclic causal models are commonly used to model the data-generating process, and the inference of such models is a well-studied problem. However, existing methods have significant limitations. Methods based on conditional independencies (Spirtes et al. 1993; Pearl 2000) cannot distinguish between independence-equivalent models, whereas approaches purely based on Independent Component Analysis (Shimizu et al. 2006) are inapplicable to data which is partially Gaussian. In this paper, we generalize and combine the two approaches, to yield a method able to learn the model structure in many cases for which the previous methods provide answers that are either incorrect or are not as informative as possible. We give exact graphical conditions for when two distinct models represent the same family of distributions, and empirically demonstrate the power of our method through thorough simulations.
Estimation of causal orders in a linear non-Gaussian acyclic model: a method robust against latent confounders
Apr 09, 2012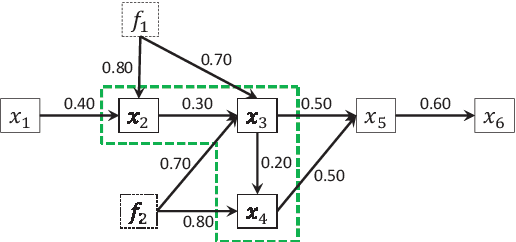
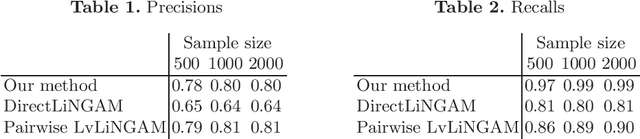
Abstract:We consider to learn a causal ordering of variables in a linear non-Gaussian acyclic model called LiNGAM. Several existing methods have been shown to consistently estimate a causal ordering assuming that all the model assumptions are correct. But, the estimation results could be distorted if some assumptions actually are violated. In this paper, we propose a new algorithm for learning causal orders that is robust against one typical violation of the model assumptions: latent confounders. We demonstrate the effectiveness of our method using artificial data.
Discovering causal structures in binary exclusive-or skew acyclic models
Feb 14, 2012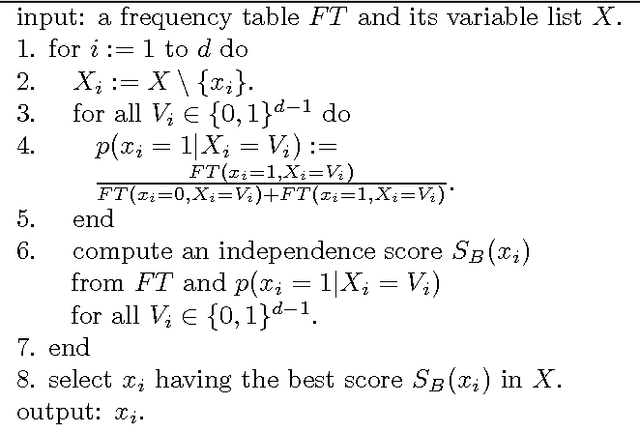
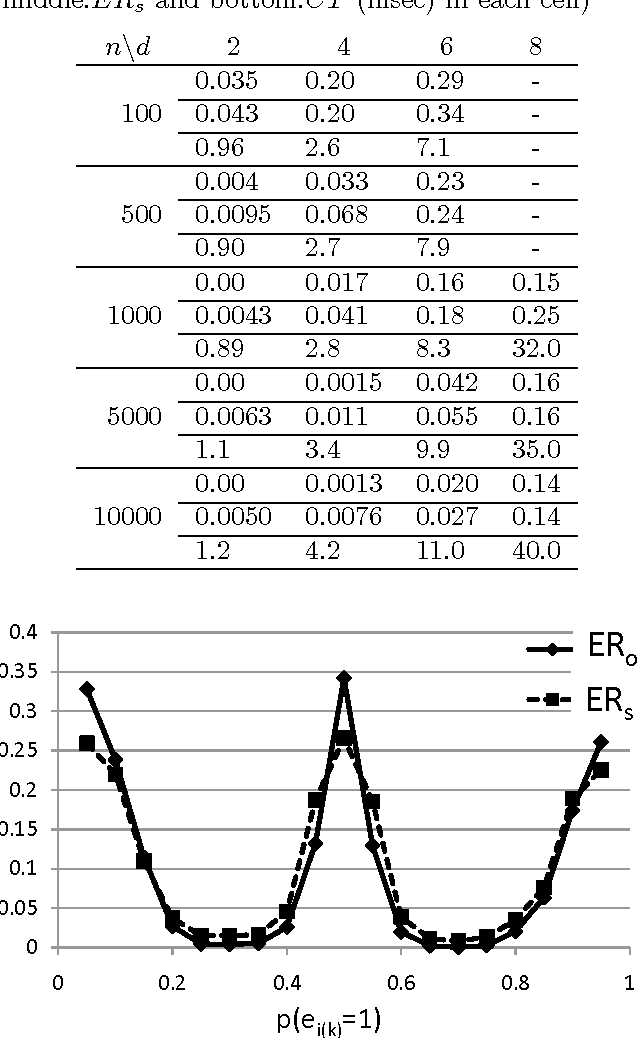
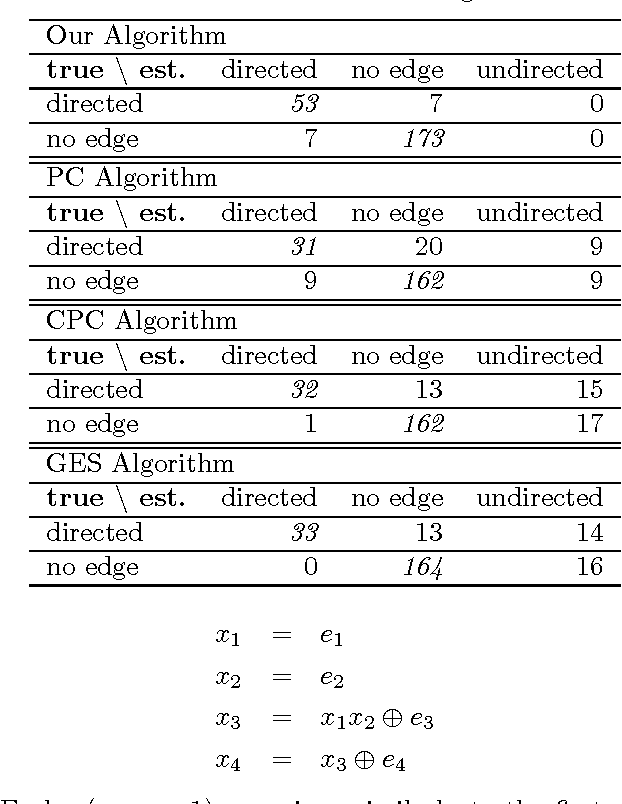
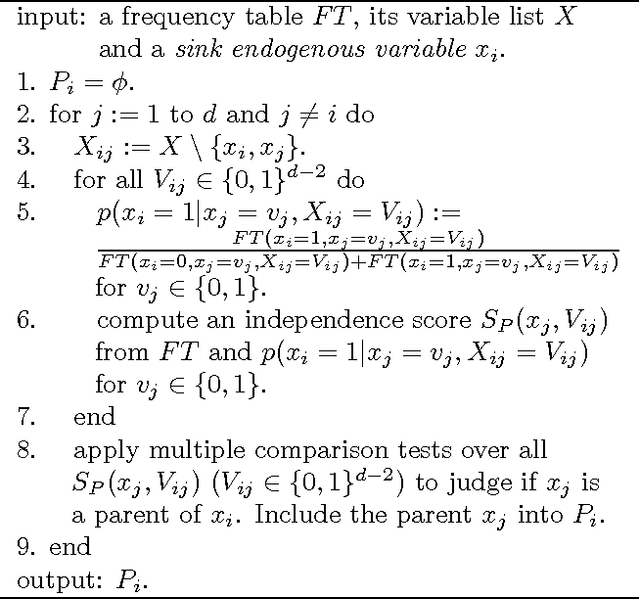
Abstract:Discovering causal relations among observed variables in a given data set is a main topic in studies of statistics and artificial intelligence. Recently, some techniques to discover an identifiable causal structure have been explored based on non-Gaussianity of the observed data distribution. However, most of these are limited to continuous data. In this paper, we present a novel causal model for binary data and propose a new approach to derive an identifiable causal structure governing the data based on skew Bernoulli distributions of external noise. Experimental evaluation shows excellent performance for both artificial and real world data sets.
Joint estimation of linear non-Gaussian acyclic models
Nov 30, 2011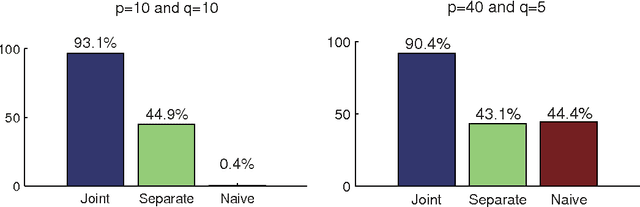
Abstract:A linear non-Gaussian structural equation model called LiNGAM is an identifiable model for exploratory causal analysis. Previous methods estimate a causal ordering of variables and their connection strengths based on a single dataset. However, in many application domains, data are obtained under different conditions, that is, multiple datasets are obtained rather than a single dataset. In this paper, we present a new method to jointly estimate multiple LiNGAMs under the assumption that the models share a causal ordering but may have different connection strengths and differently distributed variables. In simulations, the new method estimates the models more accurately than estimating them separately.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge