Robert Jenssen
Reservoir computing approaches for representation and classification of multivariate time series
Nov 06, 2018
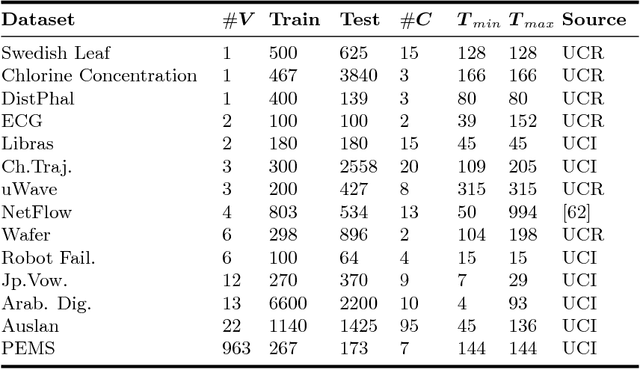
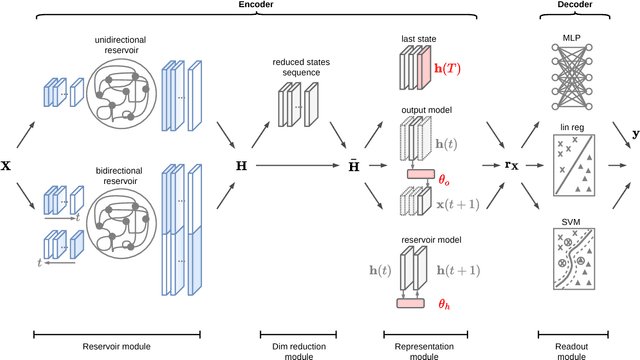
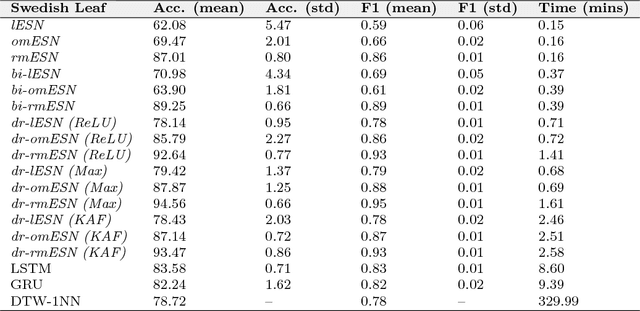
Abstract:Classification of multivariate time series (MTS) has been tackled with a large variety of methodologies and applied to a wide range of scenarios. Among the existing approaches, reservoir computing (RC) techniques, which implement a fixed and high-dimensional recurrent network to process sequential data, are computationally efficient tools to generate a vectorial, fixed-size representation of the MTS that can be further processed by standard classifiers. Despite their unrivaled training speed, MTS classifiers based on a standard RC architecture fail to achieve the same accuracy of other classifiers, such as those exploiting fully trainable recurrent networks. In this paper we introduce the reservoir model space, an RC approach to learn vectorial representations of MTS in an unsupervised fashion. Each MTS is encoded within the parameters of a linear model trained to predict a low-dimensional embedding of the reservoir dynamics. Our model space yields a powerful representation of the MTS and, thanks to an intermediate dimensionality reduction procedure, attains computational performance comparable to other RC methods. As a second contribution we propose a modular RC framework for MTS classification, with an associated open source Python library. By combining the different modules it is possible to seamlessly implement advanced RC architectures, including our proposed unsupervised representation, bidirectional reservoirs, and non-linear readouts, such as deep neural networks with both fixed and flexible activation functions. Results obtained on benchmark and real-world MTS datasets show that RC classifiers are dramatically faster and, when implemented using our proposed representation, also achieve superior classification accuracy.
Understanding Convolutional Neural Network Training with Information Theory
Oct 12, 2018
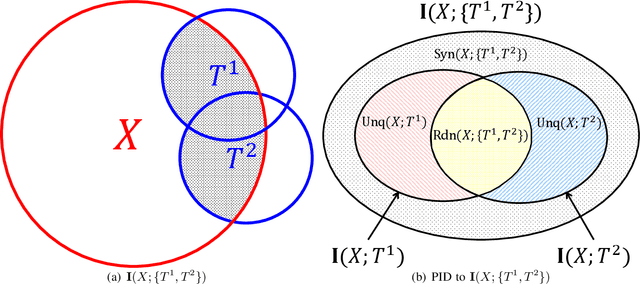

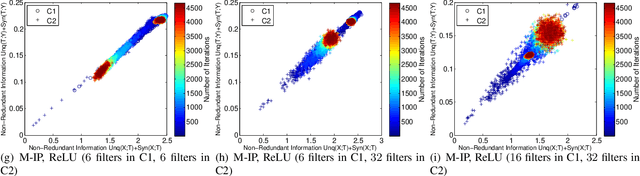
Abstract:Using information theoretic concepts to understand and explore the inner organization of deep neural networks (DNNs) remains a big challenge. Recently, the concept of an information plane (coupled with the famed information bottleneck principle) began to shed light on the analysis of multilayer perceptrons (MLPs). We provided an in-depth insight into stacked autoencoders (SAEs) using a novel matrix-based Renyi's {\alpha}-entropy functional, enabling for the first time the analysis of the dynamics of learning using information flow in the real-world scenario involving complex network architecture and large data. Despite the great potential of these past works, there are several open questions when it comes to applying information theoretic concepts to understand convolutional neural networks (CNNs). These include for instance the accurate estimation of information quantities among multiple variables, and the many different training methodologies. By extending the novel matrix-based Renyi's {\alpha}-entropy functional to a multivariate scenario and introducing the partial information decomposition (PID) framework, this paper presents a systematic method to analyze CNNs training using information theory. Our results validate two fundamental data processing inequalities in CNNs, and also reveals some fundamental issues embedded in the training phase of CNNs.
Multivariate Extension of Matrix-based Renyi's α-order Entropy Functional
Aug 23, 2018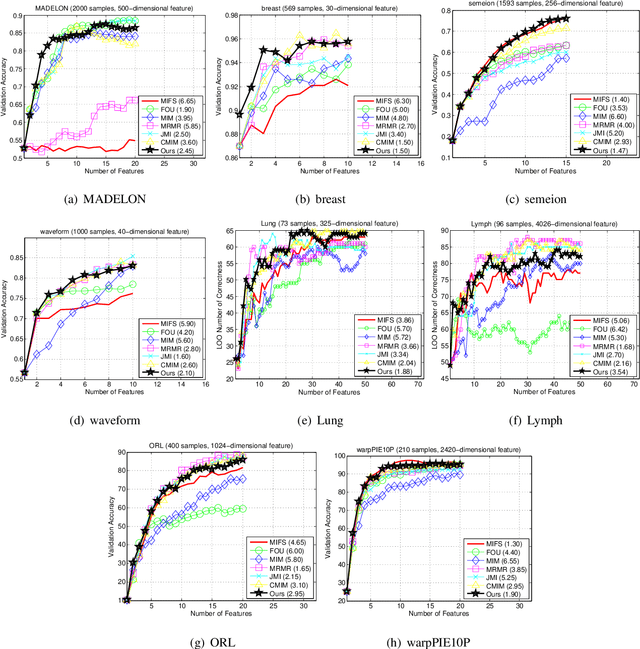
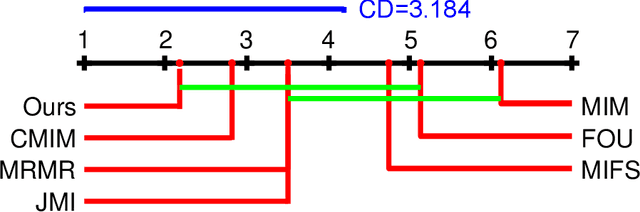
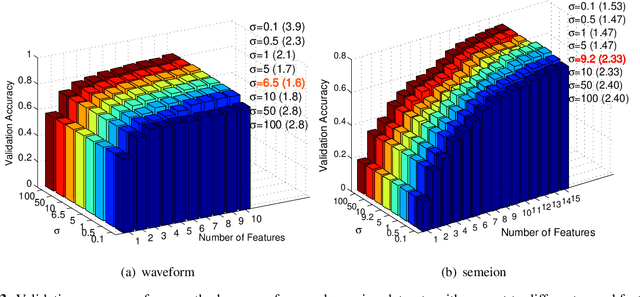
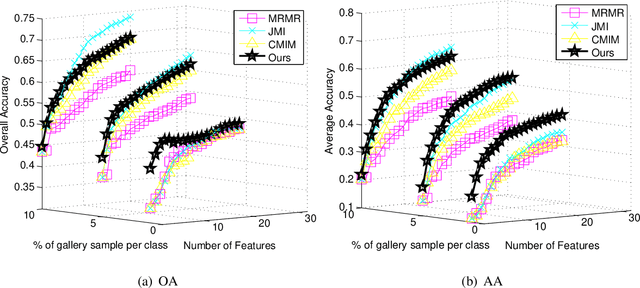
Abstract:The matrix-based Renyi's {\alpha}-order entropy functional was recently introduced using the normalized eigenspectrum of an Hermitian matrix of the projected data in the reproducing kernel Hilbert space (RKHS). However, the current theory in the matrix-based Renyi's {\alpha}-order entropy functional only defines the entropy of a single variable or mutual information between two random variables. In information theory and machine learning communities, one is also frequently interested in multivariate information quantities, such as the multivariate joint entropy and different interactive quantities among multiple variables. In this paper, we first define the matrix-based Renyi's {\alpha}-order joint entropy among multiple variables. We then show how this definition can ease the estimation of various information quantities that measure the interactions among multiple variables, such as interactive information and total correlation. We finally present an application to feature selection to show how our definition provides a simple yet powerful way to estimate a widely-acknowledged intractable quantity from data. A real example on hyperspectral image (HSI) band selection is also provided.
The Deep Kernelized Autoencoder
Jul 23, 2018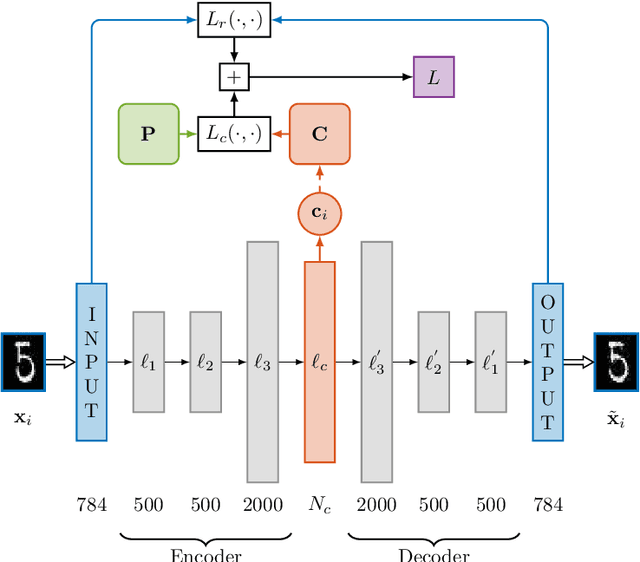
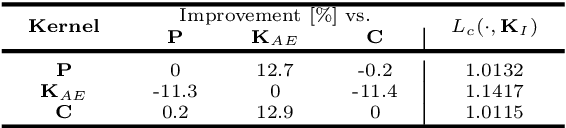
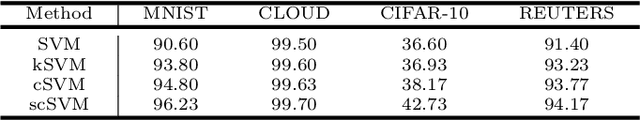

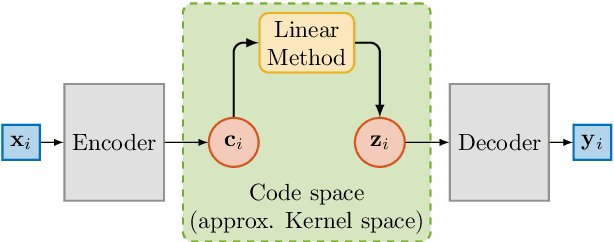

Abstract:Autoencoders learn data representations (codes) in such a way that the input is reproduced at the output of the network. However, it is not always clear what kind of properties of the input data need to be captured by the codes. Kernel machines have experienced great success by operating via inner-products in a theoretically well-defined reproducing kernel Hilbert space, hence capturing topological properties of input data. In this paper, we enhance the autoencoder's ability to learn effective data representations by aligning inner products between codes with respect to a kernel matrix. By doing so, the proposed kernelized autoencoder allows learning similarity-preserving embeddings of input data, where the notion of similarity is explicitly controlled by the user and encoded in a positive semi-definite kernel matrix. Experiments are performed for evaluating both reconstruction and kernel alignment performance in classification tasks and visualization of high-dimensional data. Additionally, we show that our method is capable to emulate kernel principal component analysis on a denoising task, obtaining competitive results at a much lower computational cost.
An overview and comparative analysis of Recurrent Neural Networks for Short Term Load Forecasting
Jul 20, 2018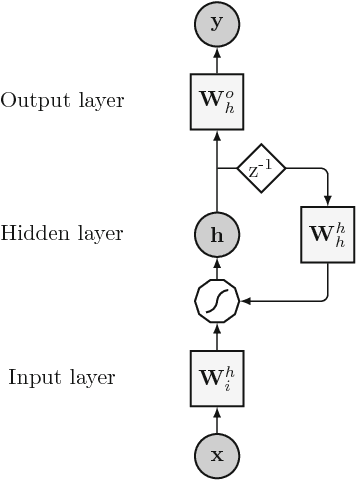
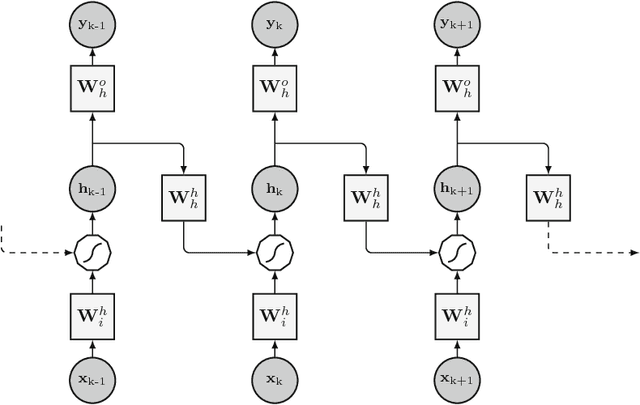
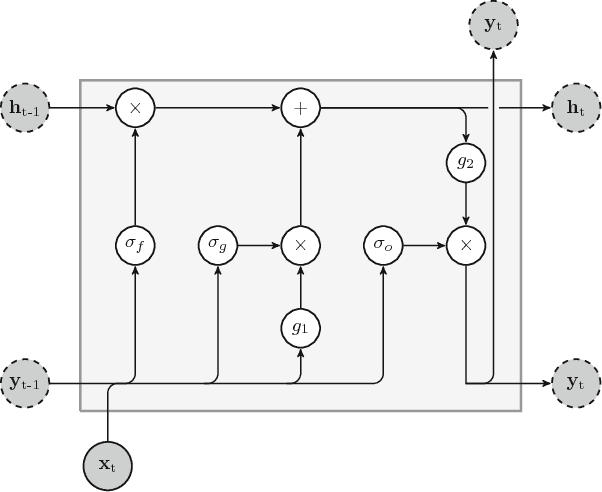
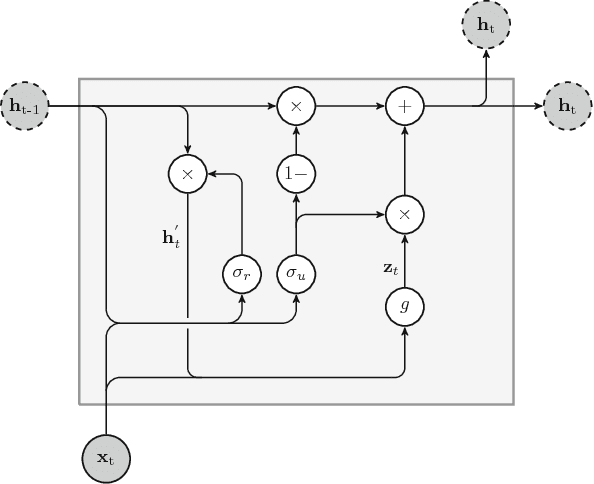
Abstract:The key component in forecasting demand and consumption of resources in a supply network is an accurate prediction of real-valued time series. Indeed, both service interruptions and resource waste can be reduced with the implementation of an effective forecasting system. Significant research has thus been devoted to the design and development of methodologies for short term load forecasting over the past decades. A class of mathematical models, called Recurrent Neural Networks, are nowadays gaining renewed interest among researchers and they are replacing many practical implementation of the forecasting systems, previously based on static methods. Despite the undeniable expressive power of these architectures, their recurrent nature complicates their understanding and poses challenges in the training procedures. Recently, new important families of recurrent architectures have emerged and their applicability in the context of load forecasting has not been investigated completely yet. In this paper we perform a comparative study on the problem of Short-Term Load Forecast, by using different classes of state-of-the-art Recurrent Neural Networks. We test the reviewed models first on controlled synthetic tasks and then on different real datasets, covering important practical cases of study. We provide a general overview of the most important architectures and we define guidelines for configuring the recurrent networks to predict real-valued time series.
Uncertainty and Interpretability in Convolutional Neural Networks for Semantic Segmentation of Colorectal Polyps
Jul 16, 2018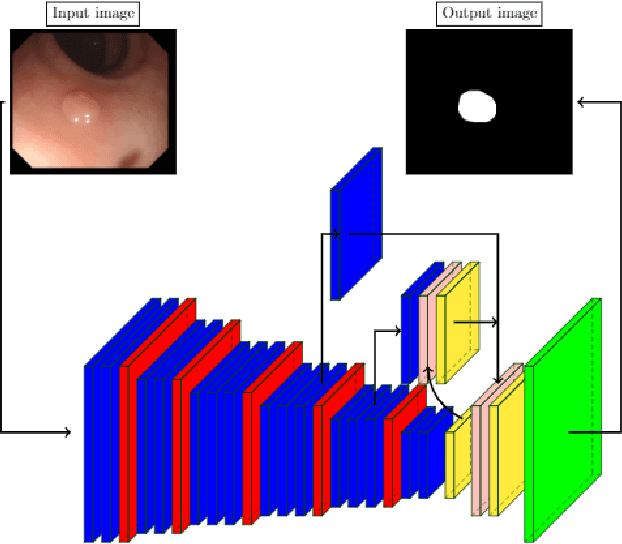
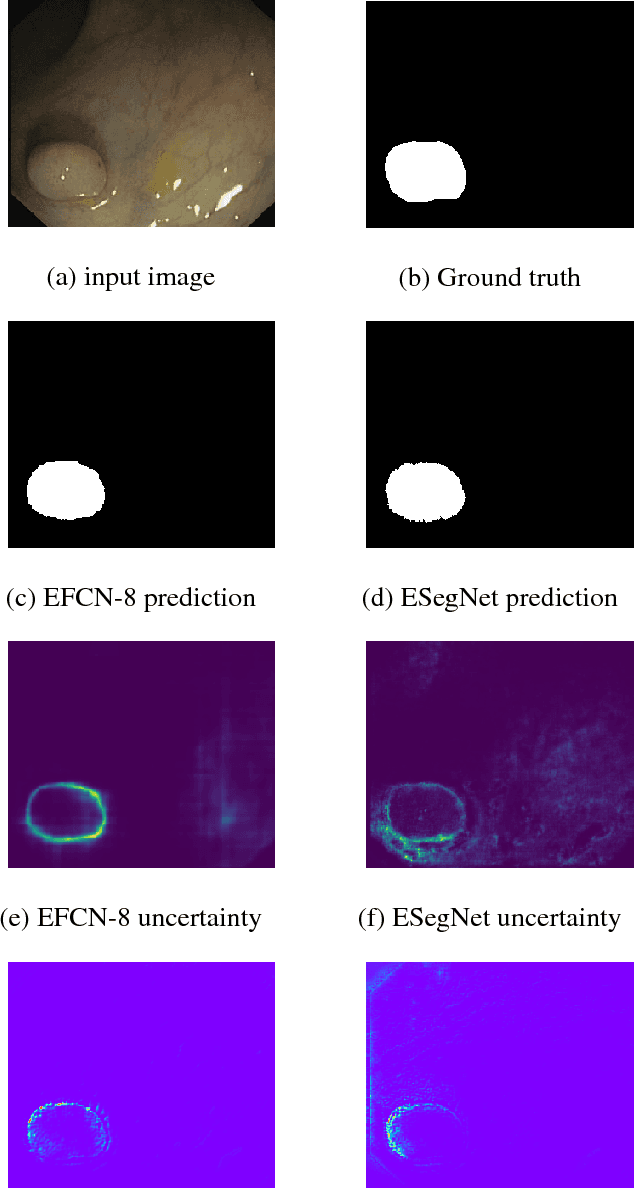
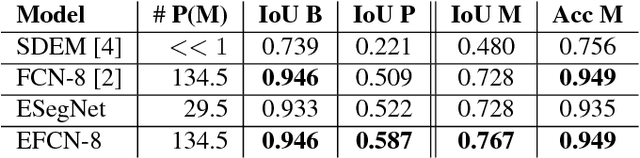
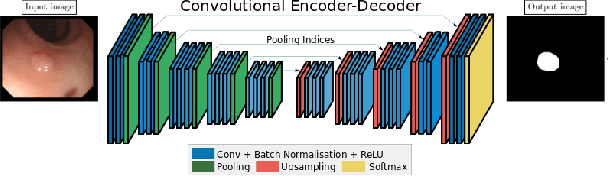

Abstract:Convolutional Neural Networks (CNNs) are propelling advances in a range of different computer vision tasks such as object detection and object segmentation. Their success has motivated research in applications of such models for medical image analysis. If CNN-based models are to be helpful in a medical context, they need to be precise, interpretable, and uncertainty in predictions must be well understood. In this paper, we develop and evaluate recent advances in uncertainty estimation and model interpretability in the context of semantic segmentation of polyps from colonoscopy images. We evaluate and enhance several architectures of Fully Convolutional Networks (FCNs) for semantic segmentation of colorectal polyps and provide a comparison between these models. Our highest performing model achieves a 76.06\% mean IOU accuracy on the EndoScene dataset, a considerable improvement over the previous state-of-the-art.
Segment-Based Credit Scoring Using Latent Clusters in the Variational Autoencoder
Jun 07, 2018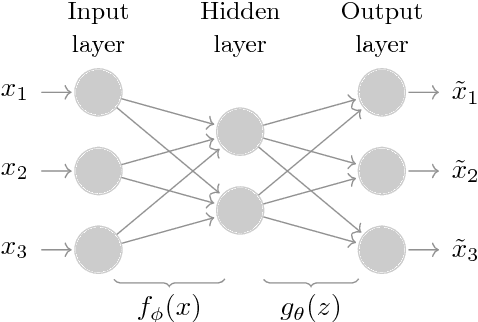
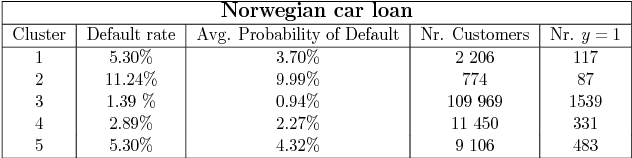

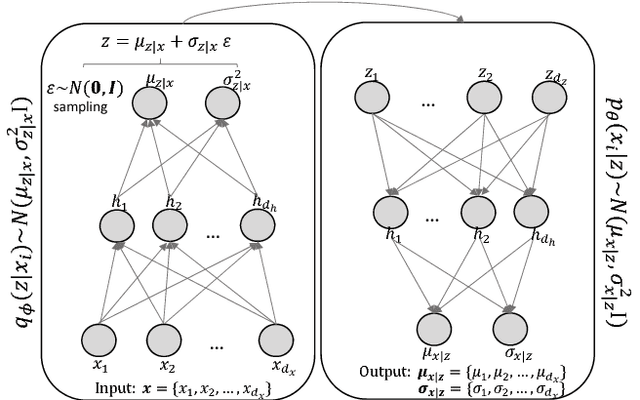
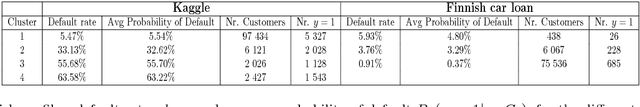
Abstract:Identifying customer segments in retail banking portfolios with different risk profiles can improve the accuracy of credit scoring. The Variational Autoencoder (VAE) has shown promising results in different research domains, and it has been documented the powerful information embedded in the latent space of the VAE. We use the VAE and show that transforming the input data into a meaningful representation, it is possible to steer configurations in the latent space of the VAE. Specifically, the Weight of Evidence (WoE) transformation encapsulates the propensity to fall into financial distress and the latent space in the VAE preserves this characteristic in a well-defined clustering structure. These clusters have considerably different risk profiles and therefore are suitable not only for credit scoring but also for marketing and customer purposes. This new clustering methodology offers solutions to some of the challenges in the existing clustering algorithms, e.g., suggests the number of clusters, assigns cluster labels to new customers, enables cluster visualization, scales to large datasets, captures non-linear relationships among others. Finally, for portfolios with a large number of customers in each cluster, developing one classifier model per cluster can improve the credit scoring assessment.
Learning representations for multivariate time series with missing data using Temporal Kernelized Autoencoders
May 09, 2018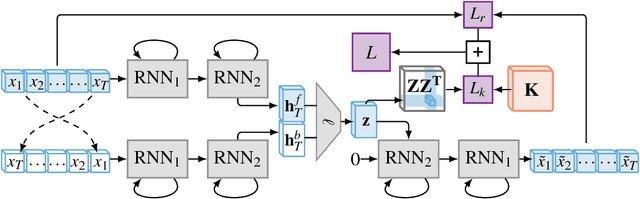
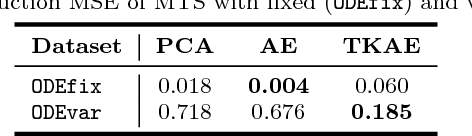
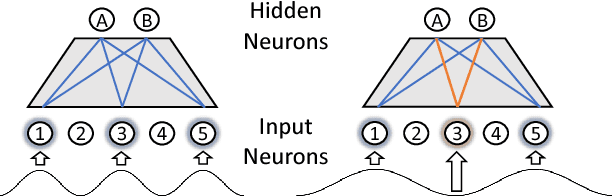
Abstract:Learning compressed representations of multivariate time series (MTS) facilitate the analysis and process of the data in presence of noise, redundant information, and large amount of variables and time steps. However, classic dimensionality reduction approaches are not designed to process sequential data, especially in the presence of missing values. In this work, we propose a novel autoencoder architecture based on recurrent neural networks to generate compressed representations of MTS, which may contain missing values and have variable lengths. Our autoencoder learns fixed-length vectorial representations, whose pairwise similarities are aligned with a kernel function that operates in input space and handles missing values. This, allows to preserve relationships in the low-dimensional vector space even in presence of missing values. To highlight the main features of the proposed autoencoder, we first investigate its performance in controlled experiments. Successively, we show how the learned representations can be exploited both in several benchmark and real-world classification tasks on medical data. Finally, based on the proposed architecture, we conceive a framework for one-class classification and imputation of missing data in time series extracted from ECG signals.
Urban Land Cover Classification with Missing Data Modalities Using Deep Convolutional Neural Networks
May 08, 2018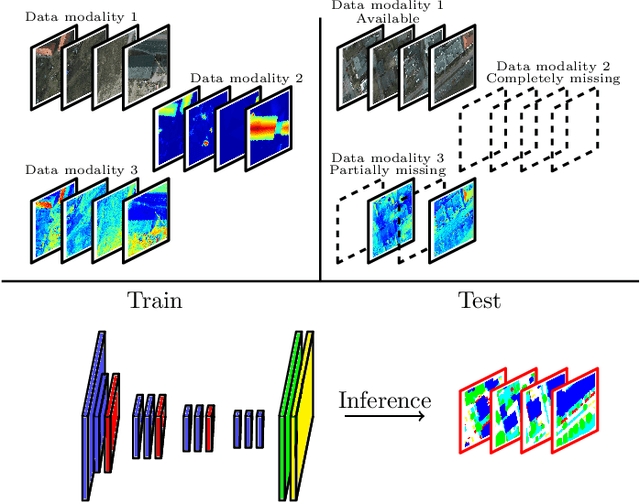
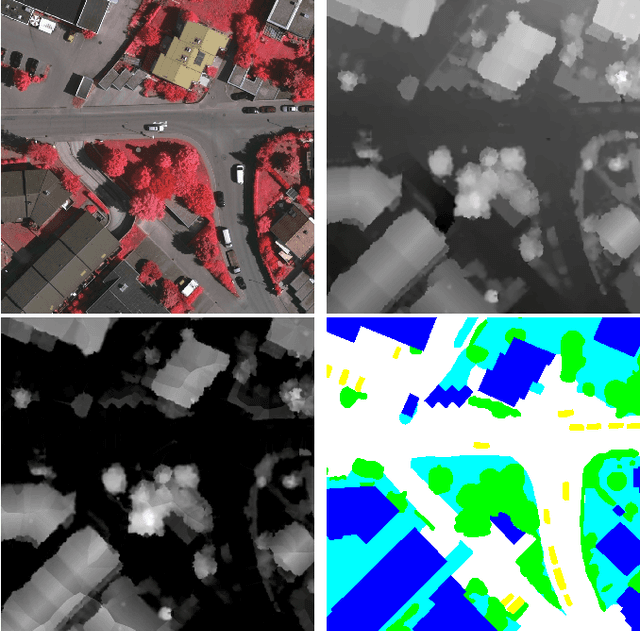
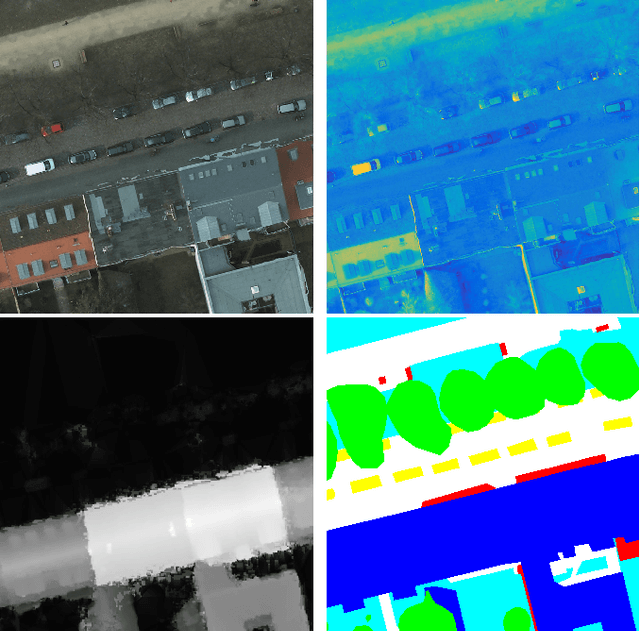


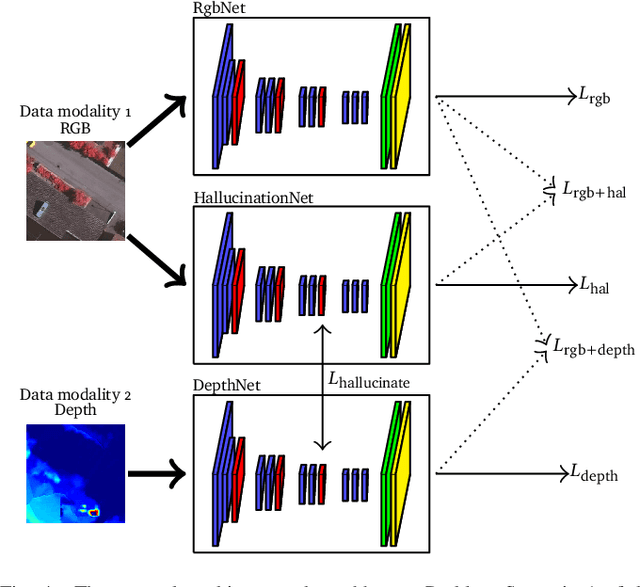
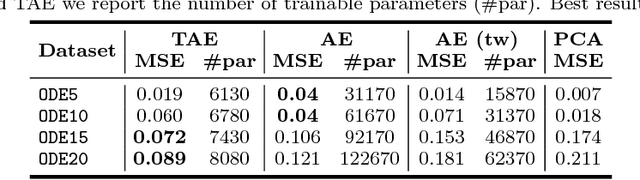
Abstract:Automatic urban land cover classification is a fundamental problem in remote sensing, e.g. for environmental monitoring. The problem is highly challenging, as classes generally have high inter-class and low intra-class variance. Techniques to improve urban land cover classification performance in remote sensing include fusion of data from different sensors with different data modalities. However, such techniques require all modalities to be available to the classifier in the decision-making process, i.e. at test time, as well as in training. If a data modality is missing at test time, current state-of-the-art approaches have in general no procedure available for exploiting information from these modalities. This represents a waste of potentially useful information. We propose as a remedy a convolutional neural network (CNN) architecture for urban land cover classification which is able to embed all available training modalities in a so-called hallucination network. The network will in effect replace missing data modalities in the test phase, enabling fusion capabilities even when data modalities are missing in testing. We demonstrate the method using two datasets consisting of optical and digital surface model (DSM) images. We simulate missing modalities by assuming that DSM images are missing during testing. Our method outperforms both standard CNNs trained only on optical images as well as an ensemble of two standard CNNs. We further evaluate the potential of our method to handle situations where only some DSM images are missing during testing. Overall, we show that we can clearly exploit training time information of the missing modality during testing.
An Unsupervised Multivariate Time Series Kernel Approach for Identifying Patients with Surgical Site Infection from Blood Samples
Mar 21, 2018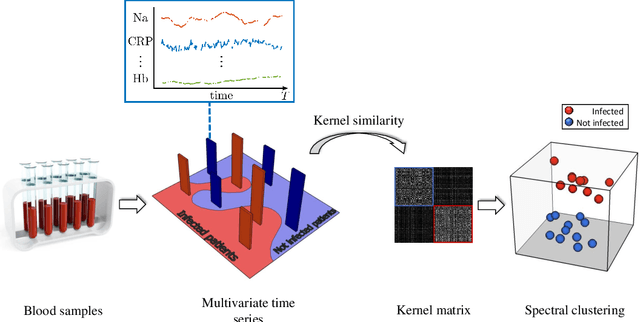
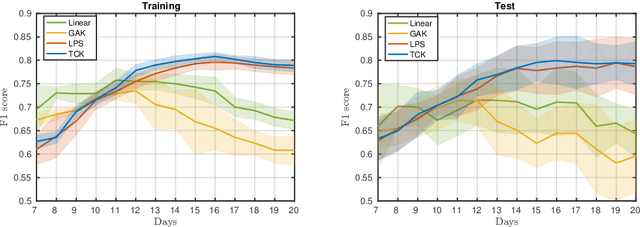
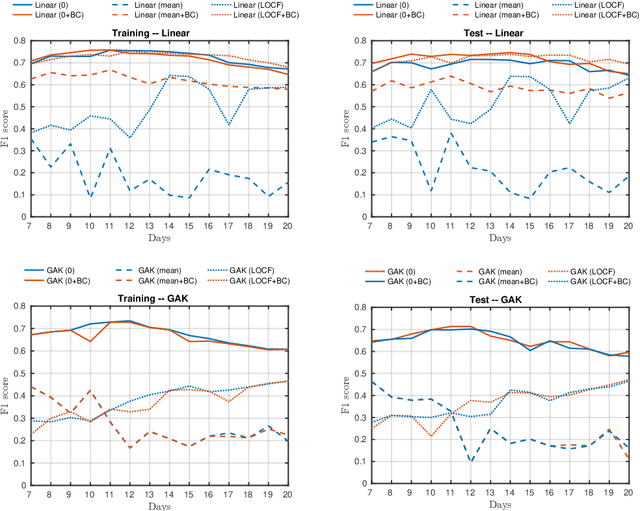
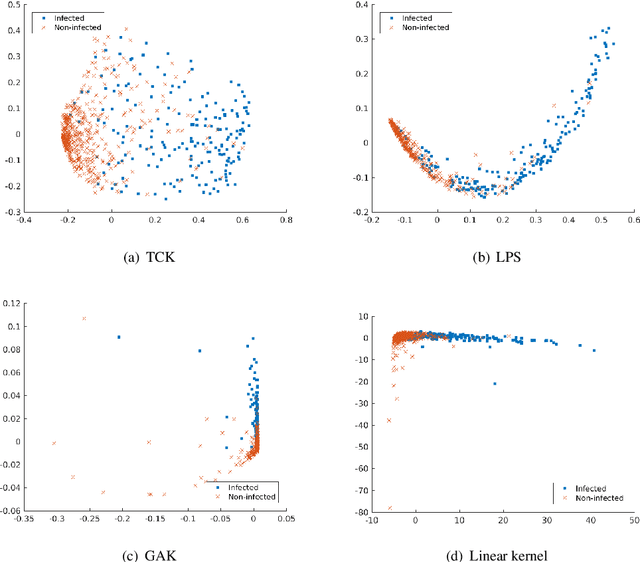
Abstract:A large fraction of the electronic health records consists of clinical measurements collected over time, such as blood tests, which provide important information about the health status of a patient. These sequences of clinical measurements are naturally represented as time series, characterized by multiple variables and the presence of missing data, which complicate analysis. In this work, we propose a surgical site infection detection framework for patients undergoing colorectal cancer surgery that is completely unsupervised, hence alleviating the problem of getting access to labelled training data. The framework is based on powerful kernels for multivariate time series that account for missing data when computing similarities. Our approach show superior performance compared to baselines that have to resort to imputation techniques and performs comparable to a supervised classification baseline.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge