Rishi Mukhopadhyay
Using Residual Dipolar Couplings from Two Alignment Media to Detect Structural Homology
Nov 06, 2019

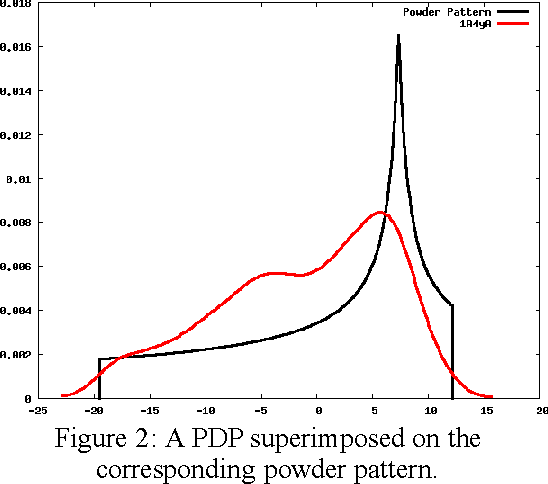
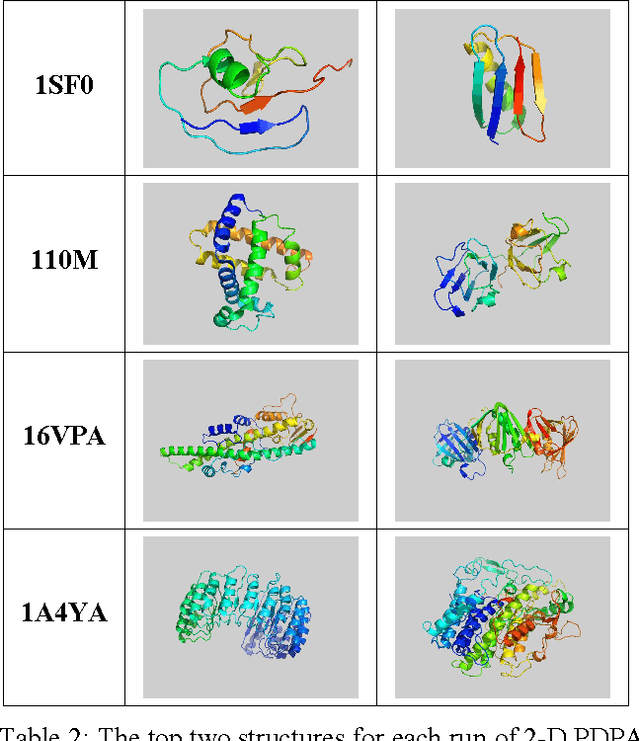
Abstract:The method of Probability Density Profile Analysis has been introduced previously as a tool to find the best match between a set of experimentally generated Residual Dipolar Couplings and a set of known protein structures. While it proved effective on small databases in identifying protein fold families, and for picking the best result from computational protein folding tool ROBETTA, for larger data sets, more data is required. Here, the method of 2-D Probability Density Profile Analysis is presented which incorporates paired RDC data from 2 alignment media for N-H vectors. The method was tested using synthetic RDC data generated with +/-1 Hz error. The results show that the addition of information from a second alignment medium makes 2-D PDPA a much more effective tool that is able to identify a structure from a database of 600 protein fold family representatives.
Protein Fold Family Recognition From Unassigned Residual Dipolar Coupling Data
Nov 01, 2019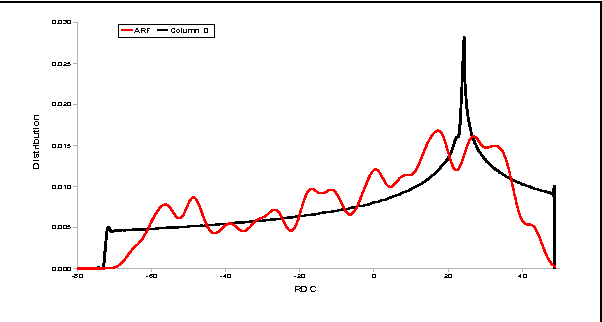
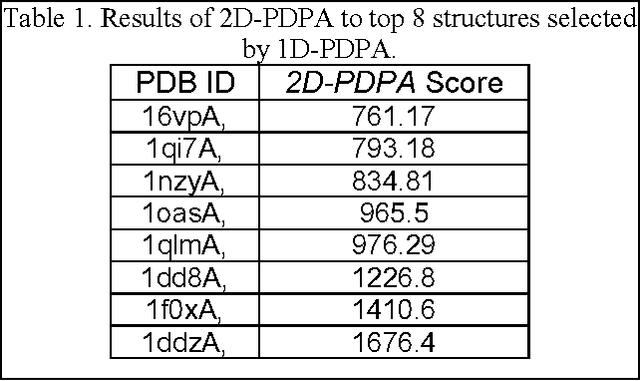
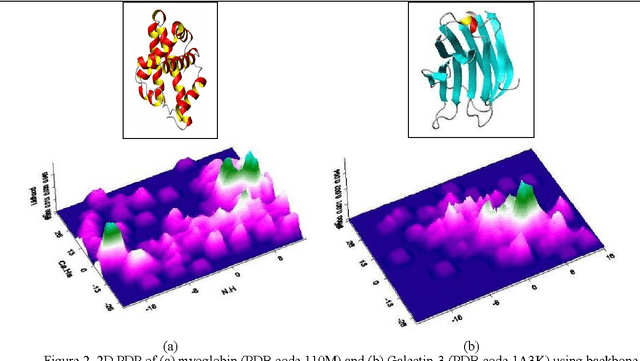
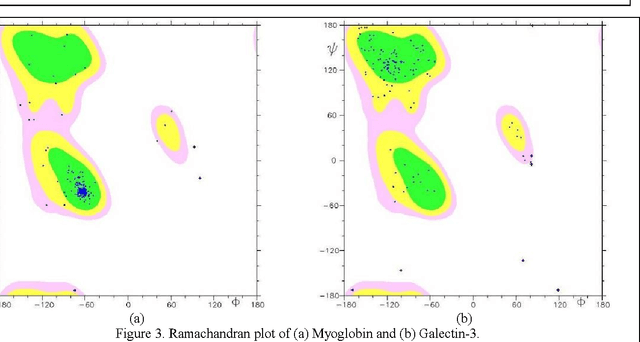
Abstract:Despite many advances in computational modeling of protein structures, these methods have not been widely utilized by experimental structural biologists. Two major obstacles are preventing the transition from a purely-experimental to a purely-computational mode of protein structure determination. The first problem is that most computational methods need a large library of computed structures that span a large variety of protein fold families, while structural genomics initiatives have slowed in their ability to provide novel protein folds in recent years. The second problem is an unwillingness to trust computational models that have no experimental backing. In this paper we test a potential solution to these problems that we have called Probability Density Profile Analysis (PDPA) that utilizes unassigned residual dipolar coupling data that are relatively cheap to acquire from NMR experiments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge