Paul Fieguth
Multi-Channel Feature Extraction for Virtual Histological Staining of Photon Absorption Remote Sensing Images
Jul 04, 2023Abstract:Accurate and fast histological staining is crucial in histopathology, impacting diagnostic precision and reliability. Traditional staining methods are time-consuming and subjective, causing delays in diagnosis. Digital pathology plays a vital role in advancing and optimizing histology processes to improve efficiency and reduce turnaround times. This study introduces a novel deep learning-based framework for virtual histological staining using photon absorption remote sensing (PARS) images. By extracting features from PARS time-resolved signals using a variant of the K-means method, valuable multi-modal information is captured. The proposed multi-channel cycleGAN (MC-GAN) model expands on the traditional cycleGAN framework, allowing the inclusion of additional features. Experimental results reveal that specific combinations of features outperform the conventional channels by improving the labeling of tissue structures prior to model training. Applied to human skin and mouse brain tissue, the results underscore the significance of choosing the optimal combination of features, as it reveals a substantial visual and quantitative concurrence between the virtually stained and the gold standard chemically stained hematoxylin and eosin (H&E) images, surpassing the performance of other feature combinations. Accurate virtual staining is valuable for reliable diagnostic information, aiding pathologists in disease classification, grading, and treatment planning. This study aims to advance label-free histological imaging and opens doors for intraoperative microscopy applications.
Virtual Histology with Photon Absorption Remote Sensing using a Cycle-Consistent Generative Adversarial Network with Weakly Registered Pairs
Jun 26, 2023



Abstract:Modern histopathology relies on the microscopic examination of thin tissue sections stained with histochemical techniques, typically using brightfield or fluorescence microscopy. However, the staining of samples can permanently alter their chemistry and structure, meaning an individual tissue section must be prepared for each desired staining contrast. This not only consumes valuable tissue samples but also introduces delays in essential diagnostic timelines. In this work, virtual histochemical staining is developed using label-free photon absorption remote sensing (PARS) microscopy. We present a method that generates virtually stained histology images that are indistinguishable from the gold standard hematoxylin and eosin (H&E) staining. First, PARS label-free ultraviolet absorption images are captured directly within unstained tissue specimens. The radiative and non-radiative absorption images are then preprocessed, and virtually stained through the presented pathway. The preprocessing pipeline features a self-supervised Noise2Void denoising convolutional neural network (CNN) as well as a novel algorithm for pixel-level mechanical scanning error correction. These developments significantly enhance the recovery of sub-micron tissue structures, such as nucleoli location and chromatin distribution. Finally, we used a cycle-consistent generative adversarial network CycleGAN architecture to virtually stain the preprocessed PARS data. Virtual staining is applied to thin unstained sections of malignant human skin and breast tissue samples. Clinically relevant details are revealed, with comparable contrast and quality to gold standard H&E-stained images. This work represents a crucial step to deploying label-free microscopy as an alternative to standard histopathology techniques.
Is Generative Modeling-based Stylization Necessary for Domain Adaptation in Regression Tasks?
Jun 02, 2023Abstract:Unsupervised domain adaptation (UDA) aims to bridge the gap between source and target domains in the absence of target domain labels using two main techniques: input-level alignment (such as generative modeling and stylization) and feature-level alignment (which matches the distribution of the feature maps, e.g. gradient reversal layers). Motivated from the success of generative modeling for image classification, stylization-based methods were recently proposed for regression tasks, such as pose estimation. However, use of input-level alignment via generative modeling and stylization incur additional overhead and computational complexity which limit their use in real-world DA tasks. To investigate the role of input-level alignment for DA, we ask the following question: Is generative modeling-based stylization necessary for visual domain adaptation in regression? Surprisingly, we find that input-alignment has little effect on regression tasks as compared to classification. Based on these insights, we develop a non-parametric feature-level domain alignment method -- Implicit Stylization (ImSty) -- which results in consistent improvements over SOTA regression task, without the need for computationally intensive stylization and generative modeling. Our work conducts a critical evaluation of the role of generative modeling and stylization, at a time when these are also gaining popularity for domain generalization.
CLVOS23: A Long Video Object Segmentation Dataset for Continual Learning
Apr 09, 2023



Abstract:Continual learning in real-world scenarios is a major challenge. A general continual learning model should have a constant memory size and no predefined task boundaries, as is the case in semi-supervised Video Object Segmentation (VOS), where continual learning challenges particularly present themselves in working on long video sequences. In this article, we first formulate the problem of semi-supervised VOS, specifically online VOS, as a continual learning problem, and then secondly provide a public VOS dataset, CLVOS23, focusing on continual learning. Finally, we propose and implement a regularization-based continual learning approach on LWL, an existing online VOS baseline, to demonstrate the efficacy of continual learning when applied to online VOS and to establish a CLVOS23 baseline. We apply the proposed baseline to the Long Videos dataset as well as to two short video VOS datasets, DAVIS16 and DAVIS17. To the best of our knowledge, this is the first time that VOS has been defined and addressed as a continual learning problem.
Machine Learning Challenges of Biological Factors in Insect Image Data
Nov 04, 2022


Abstract:The BIOSCAN project, led by the International Barcode of Life Consortium, seeks to study changes in biodiversity on a global scale. One component of the project is focused on studying the species interaction and dynamics of all insects. In addition to genetically barcoding insects, over 1.5 million images per year will be collected, each needing taxonomic classification. With the immense volume of incoming images, relying solely on expert taxonomists to label the images would be impossible; however, artificial intelligence and computer vision technology may offer a viable high-throughput solution. Additional tasks including manually weighing individual insects to determine biomass, remain tedious and costly. Here again, computer vision may offer an efficient and compelling alternative. While the use of computer vision methods is appealing for addressing these problems, significant challenges resulting from biological factors present themselves. These challenges are formulated in the context of machine learning in this paper.
Building Spatio-temporal Transformers for Egocentric 3D Pose Estimation
Jun 09, 2022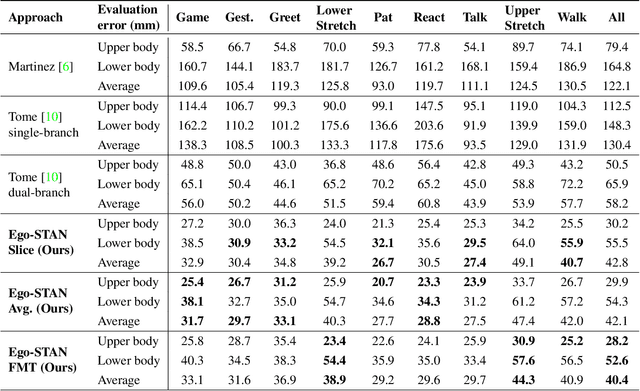
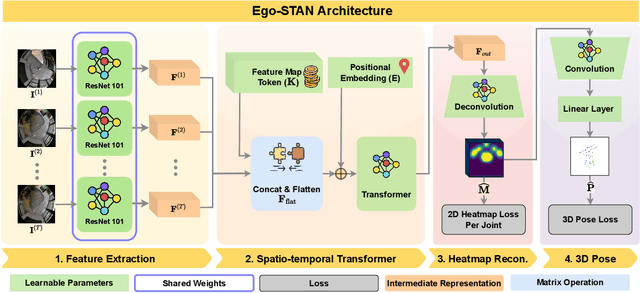
Abstract:Egocentric 3D human pose estimation (HPE) from images is challenging due to severe self-occlusions and strong distortion introduced by the fish-eye view from the head mounted camera. Although existing works use intermediate heatmap-based representations to counter distortion with some success, addressing self-occlusion remains an open problem. In this work, we leverage information from past frames to guide our self-attention-based 3D HPE estimation procedure -- Ego-STAN. Specifically, we build a spatio-temporal Transformer model that attends to semantically rich convolutional neural network-based feature maps. We also propose feature map tokens: a new set of learnable parameters to attend to these feature maps. Finally, we demonstrate Ego-STAN's superior performance on the xR-EgoPose dataset where it achieves a 30.6% improvement on the overall mean per-joint position error, while leading to a 22% drop in parameters compared to the state-of-the-art.
Virtual Histological Staining of Label-Free Total Absorption Photoacoustic Remote Sensing (TA-PARS)
Apr 01, 2022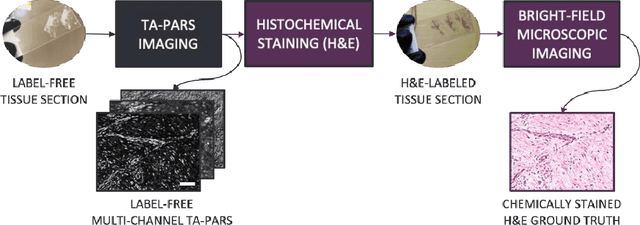
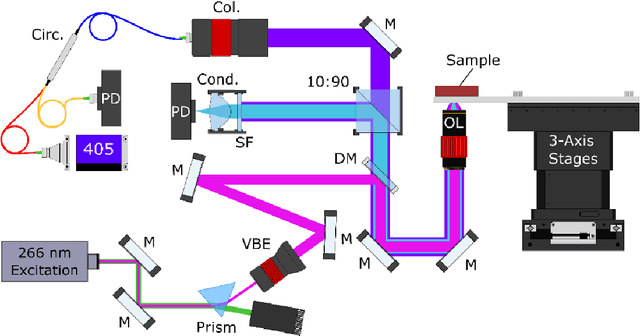
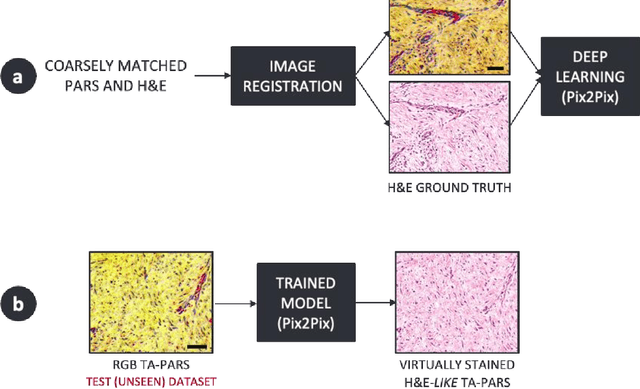
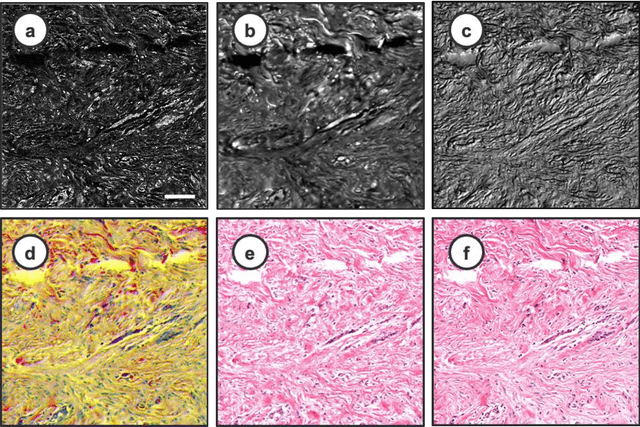
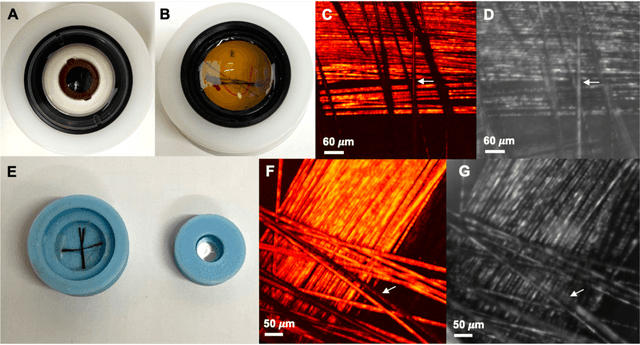
Abstract:Histopathological visualizations are a pillar of modern medicine and biological research. Surgical oncology relies exclusively on post-operative histology to determine definitive surgical success and guide adjuvant treatments. The current histology workflow is based on bright-field microscopic assessment of histochemical stained tissues and has some major limitations. For example, the preparation of stained specimens for brightfield assessment requires lengthy sample processing, delaying interventions for days or even weeks. Hence, there is a pressing need for improved histopathology methods. In this paper, we present a deep-learning-based approach for virtual label-free histochemical staining of total-absorption photoacoustic remote sensing (TA-PARS) images of unstained tissue. TA-PARS provides an array of directly measured label-free contrasts such as scattering and total absorption (radiative and non-radiative), ideal for developing H&E colorizations without the need to infer arbitrary tissue structures. We use a Pix2Pix generative adversarial network (GAN) to develop visualizations analogous to H&E staining from label-free TA-PARS images. Thin sections of human skin tissue were first virtually stained with the TA-PARS, then were chemically stained with H&E producing a one-to-one comparison between the virtual and chemical staining. The one-to-one matched virtually- and chemically- stained images exhibit high concordance validating the digital colorization of the TA-PARS images against the gold standard H&E. TA-PARS images were reviewed by four dermatologic pathologists who confirmed they are of diagnostic quality, and that resolution, contrast, and color permitted interpretation as if they were H&E. The presented approach paves the way for the development of TA-PARS slide-free histology, which promises to dramatically reduce the time from specimen resection to histological imaging.
Normalized K-Means for Noise-Insensitive Multi-Dimensional Feature Learning
Feb 15, 2022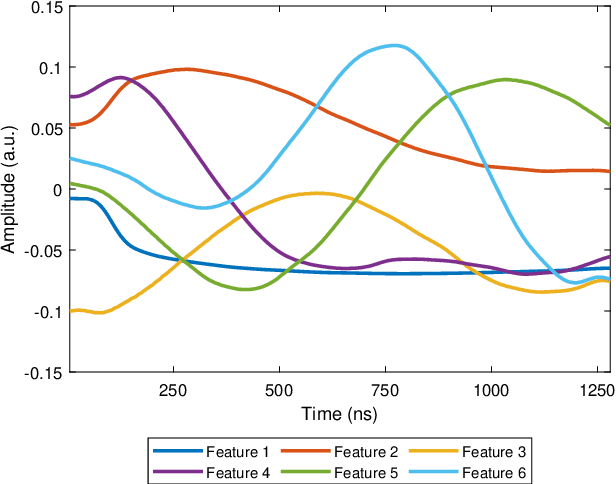
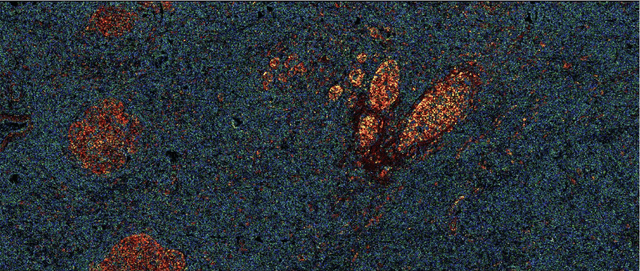
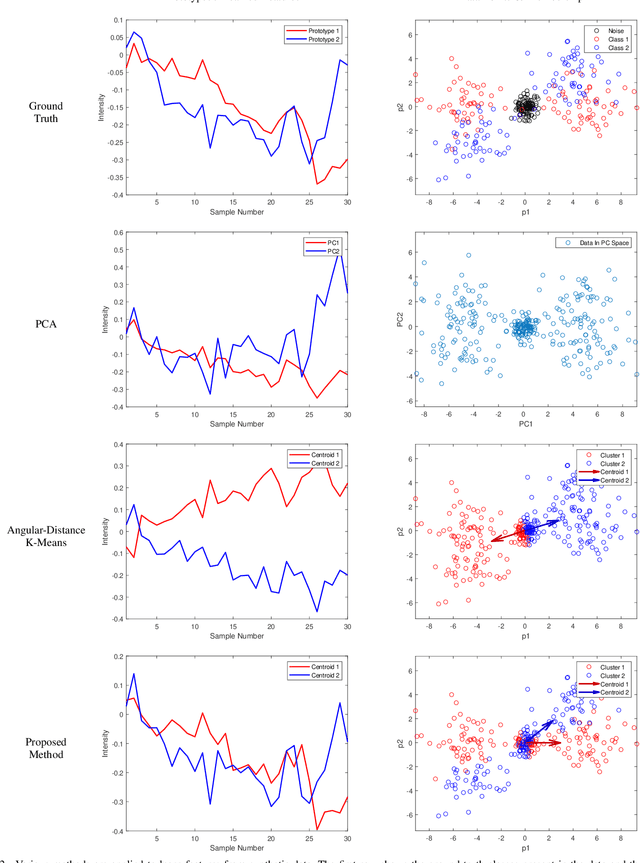
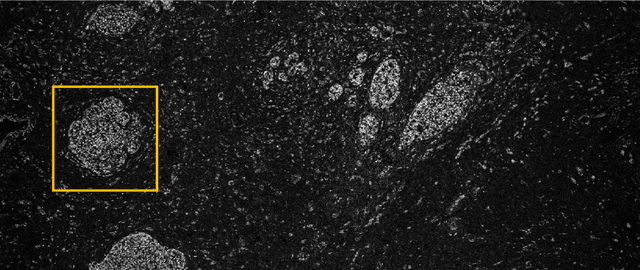

Abstract:Many measurement modalities which perform imaging by probing an object pixel-by-pixel, such as via Photoacoustic Microscopy, produce a multi-dimensional feature (typically a time-domain signal) at each pixel. In principle, the many degrees of freedom in the time-domain signal would admit the possibility of significant multi-modal information being implicitly present, much more than a single scalar "brightness", regarding the underlying targets being observed. However, the measured signal is neither a weighted-sum of basis functions (such as principal components) nor one of a set of prototypes (K-means), which has motivated the novel clustering method proposed here, capable of learning centroids (signal shapes) that are related to the underlying, albeit unknown, target characteristics in a scalable and noise-robust manner.
Survey of Deep Learning Methods for Inverse Problems
Nov 13, 2021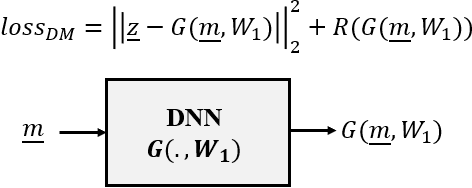
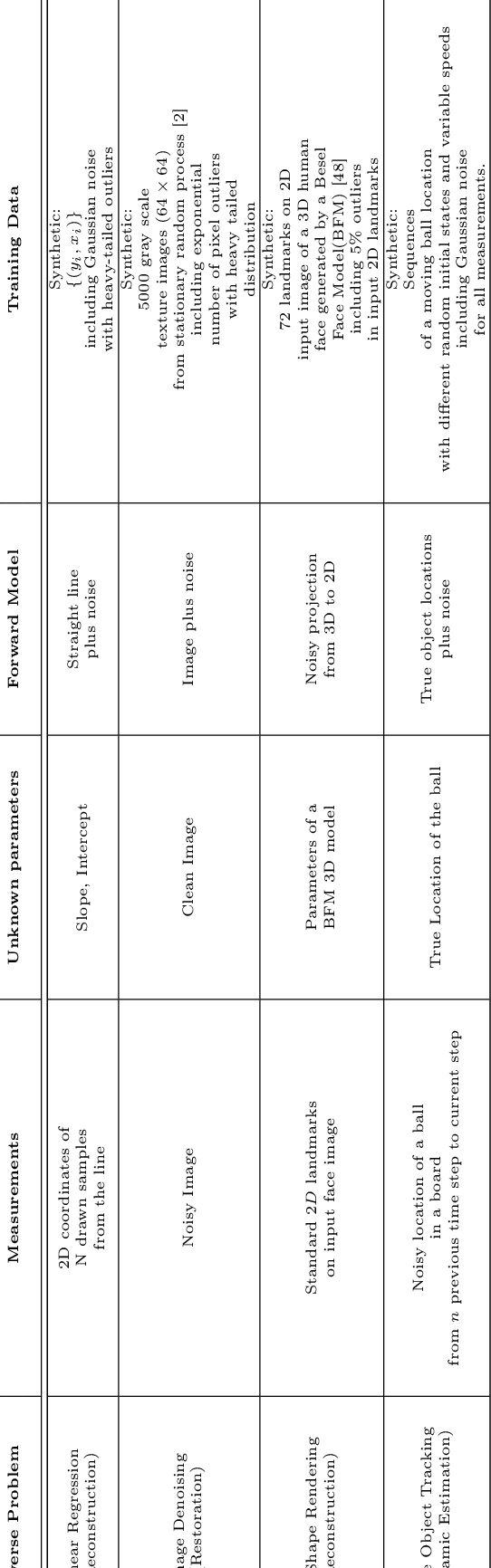
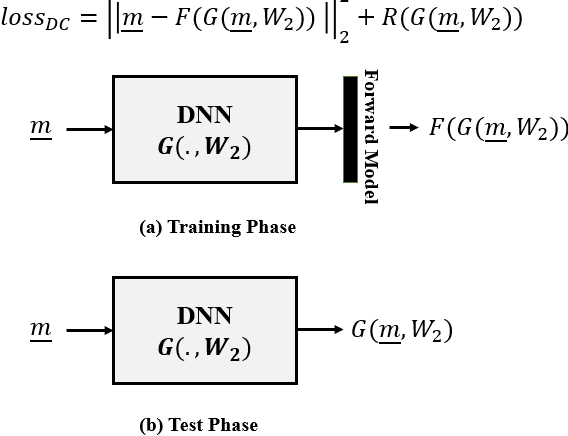
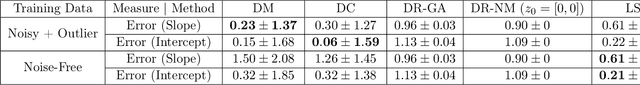
Abstract:In this paper we investigate a variety of deep learning strategies for solving inverse problems. We classify existing deep learning solutions for inverse problems into three categories of Direct Mapping, Data Consistency Optimizer, and Deep Regularizer. We choose a sample of each inverse problem type, so as to compare the robustness of the three categories, and report a statistical analysis of their differences. We perform extensive experiments on the classic problem of linear regression and three well-known inverse problems in computer vision, namely image denoising, 3D human face inverse rendering, and object tracking, selected as representative prototypes for each class of inverse problems. The overall results and the statistical analyses show that the solution categories have a robustness behaviour dependent on the type of inverse problem domain, and specifically dependent on whether or not the problem includes measurement outliers. Based on our experimental results, we conclude by proposing the most robust solution category for each inverse problem class.
In vivo functional and structural retina imaging using multimodal photoacoustic remote sensing microscopy and optical coherence tomography
Aug 26, 2021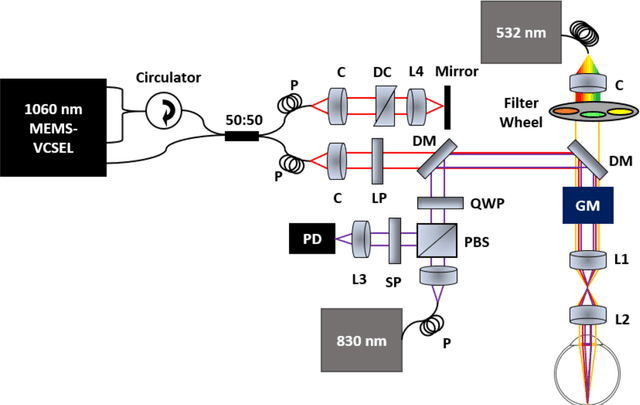

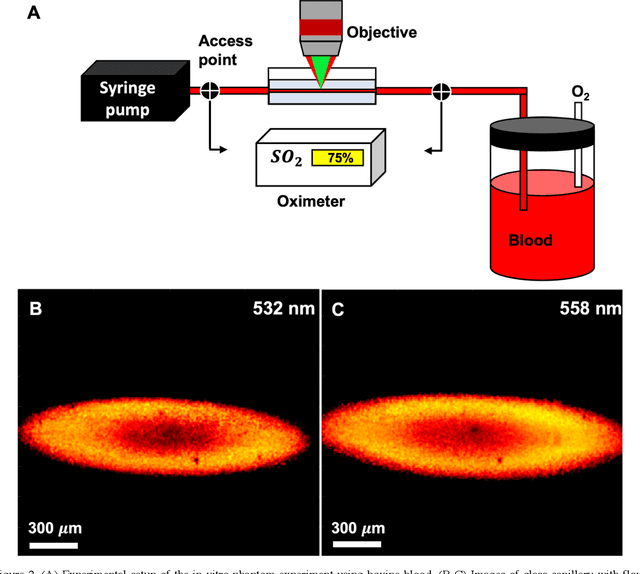
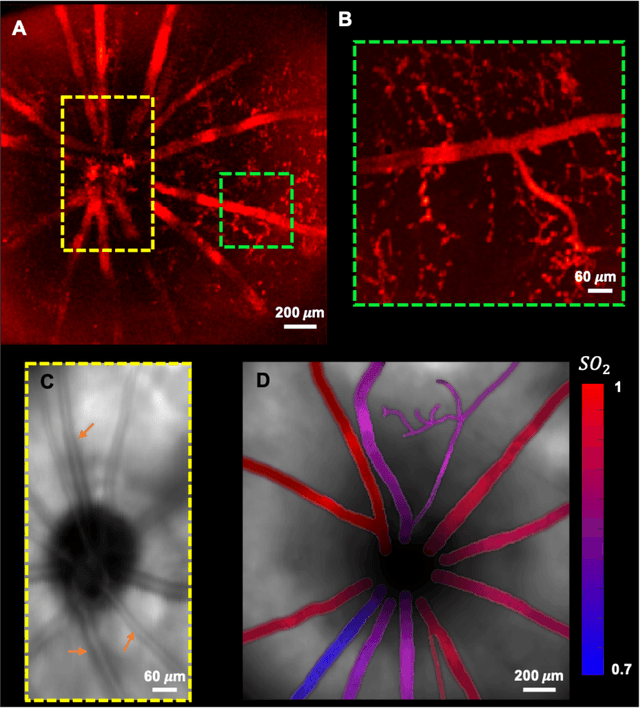
Abstract:We have developed a multimodal photoacoustic remote sensing (PARS) microscope combined with swept source optical coherence tomography for in vivo, non-contact retinal imaging. Building on the proven strength of multiwavelength PARS imaging, the system is applied for estimating retinal oxygen saturation in the rat retina. The capability of the technology is demonstrated by imaging both microanatomy and the microvasculature of the retina in vivo. To our knowledge this is the first time a non-contact photoacoustic imaging technique is employed for in vivo oxygen saturation measurement in the retina.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge