Mohammad Javad Shafiee
StochasticNet: Forming Deep Neural Networks via Stochastic Connectivity
Nov 10, 2015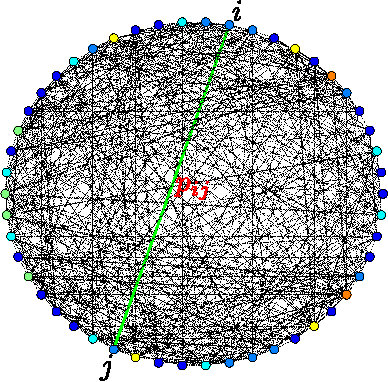
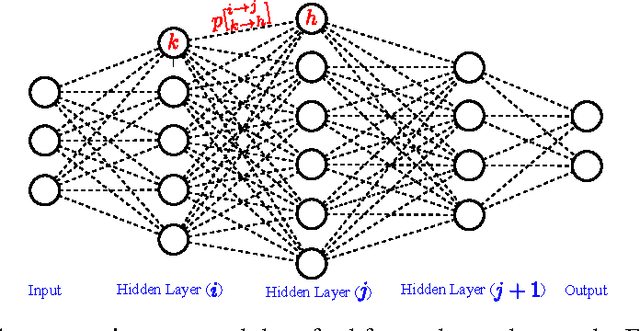
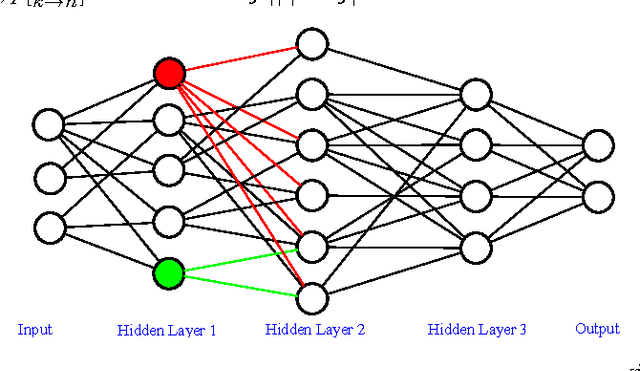
Abstract:Deep neural networks is a branch in machine learning that has seen a meteoric rise in popularity due to its powerful abilities to represent and model high-level abstractions in highly complex data. One area in deep neural networks that is ripe for exploration is neural connectivity formation. A pivotal study on the brain tissue of rats found that synaptic formation for specific functional connectivity in neocortical neural microcircuits can be surprisingly well modeled and predicted as a random formation. Motivated by this intriguing finding, we introduce the concept of StochasticNet, where deep neural networks are formed via stochastic connectivity between neurons. As a result, any type of deep neural networks can be formed as a StochasticNet by allowing the neuron connectivity to be stochastic. Stochastic synaptic formations, in a deep neural network architecture, can allow for efficient utilization of neurons for performing specific tasks. To evaluate the feasibility of such a deep neural network architecture, we train a StochasticNet using four different image datasets (CIFAR-10, MNIST, SVHN, and STL-10). Experimental results show that a StochasticNet, using less than half the number of neural connections as a conventional deep neural network, achieves comparable accuracy and reduces overfitting on the CIFAR-10, MNIST and SVHN dataset. Interestingly, StochasticNet with less than half the number of neural connections, achieved a higher accuracy (relative improvement in test error rate of ~6% compared to ConvNet) on the STL-10 dataset than a conventional deep neural network. Finally, StochasticNets have faster operational speeds while achieving better or similar accuracy performances.
Discovery Radiomics for Multi-Parametric MRI Prostate Cancer Detection
Oct 20, 2015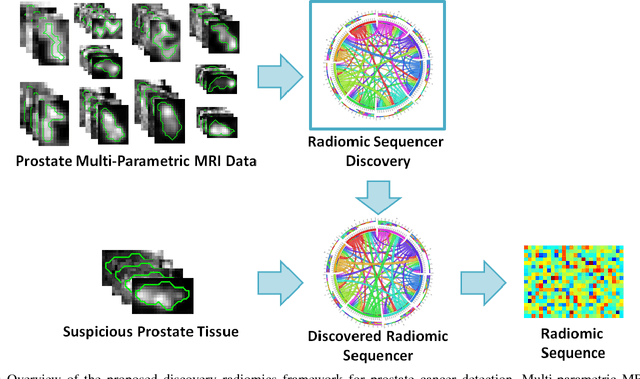
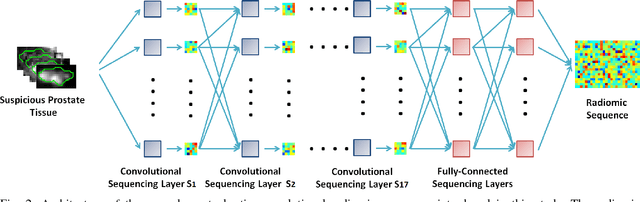
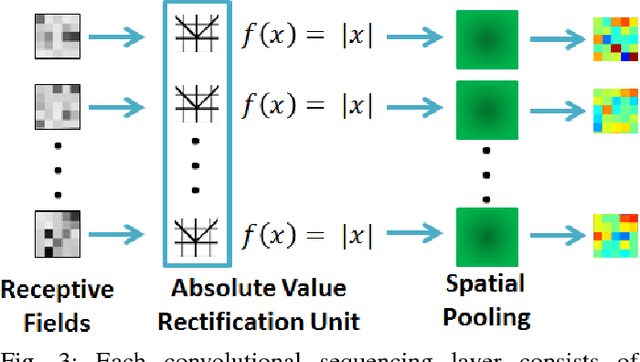

Abstract:Prostate cancer is the most diagnosed form of cancer in Canadian men, and is the third leading cause of cancer death. Despite these statistics, prognosis is relatively good with a sufficiently early diagnosis, making fast and reliable prostate cancer detection crucial. As imaging-based prostate cancer screening, such as magnetic resonance imaging (MRI), requires an experienced medical professional to extensively review the data and perform a diagnosis, radiomics-driven methods help streamline the process and has the potential to significantly improve diagnostic accuracy and efficiency, and thus improving patient survival rates. These radiomics-driven methods currently rely on hand-crafted sets of quantitative imaging-based features, which are selected manually and can limit their ability to fully characterize unique prostate cancer tumour phenotype. In this study, we propose a novel \textit{discovery radiomics} framework for generating custom radiomic sequences tailored for prostate cancer detection. Discovery radiomics aims to uncover abstract imaging-based features that capture highly unique tumour traits and characteristics beyond what can be captured using predefined feature models. In this paper, we discover new custom radiomic sequencers for generating new prostate radiomic sequences using multi-parametric MRI data. We evaluated the performance of the discovered radiomic sequencer against a state-of-the-art hand-crafted radiomic sequencer for computer-aided prostate cancer detection with a feedforward neural network using real clinical prostate multi-parametric MRI data. Results for the discovered radiomic sequencer demonstrate good performance in prostate cancer detection and clinical decision support relative to the hand-crafted radiomic sequencer. The use of discovery radiomics shows potential for more efficient and reliable automatic prostate cancer detection.
Forming A Random Field via Stochastic Cliques: From Random Graphs to Fully Connected Random Fields
Jun 30, 2015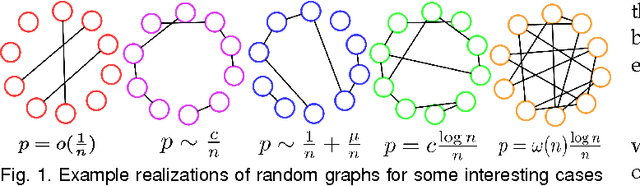
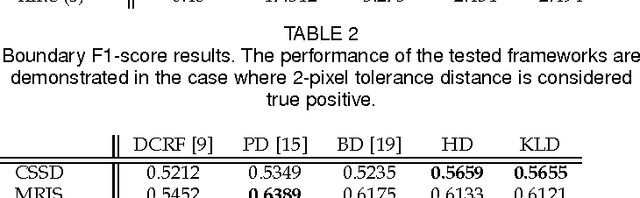
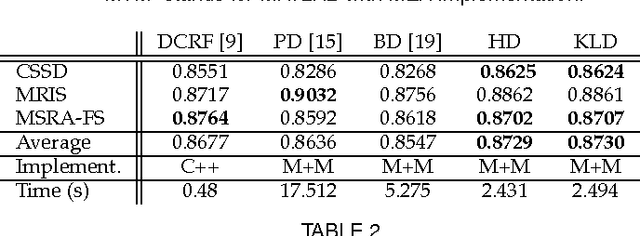
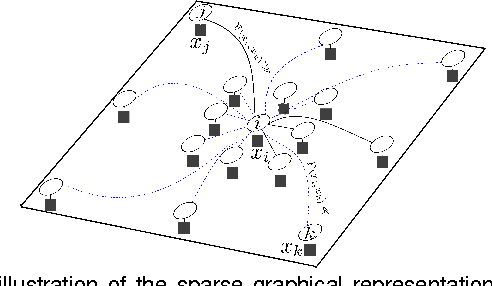

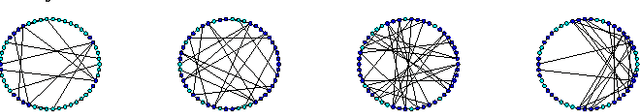
Abstract:Random fields have remained a topic of great interest over past decades for the purpose of structured inference, especially for problems such as image segmentation. The local nodal interactions commonly used in such models often suffer the short-boundary bias problem, which are tackled primarily through the incorporation of long-range nodal interactions. However, the issue of computational tractability becomes a significant issue when incorporating such long-range nodal interactions, particularly when a large number of long-range nodal interactions (e.g., fully-connected random fields) are modeled. In this work, we introduce a generalized random field framework based around the concept of stochastic cliques, which addresses the issue of computational tractability when using fully-connected random fields by stochastically forming a sparse representation of the random field. The proposed framework allows for efficient structured inference using fully-connected random fields without any restrictions on the potential functions that can be utilized. Several realizations of the proposed framework using graph cuts are presented and evaluated, and experimental results demonstrate that the proposed framework can provide competitive performance for the purpose of image segmentation when compared to existing fully-connected and principled deep random field frameworks.
A deep-structured fully-connected random field model for structured inference
May 27, 2015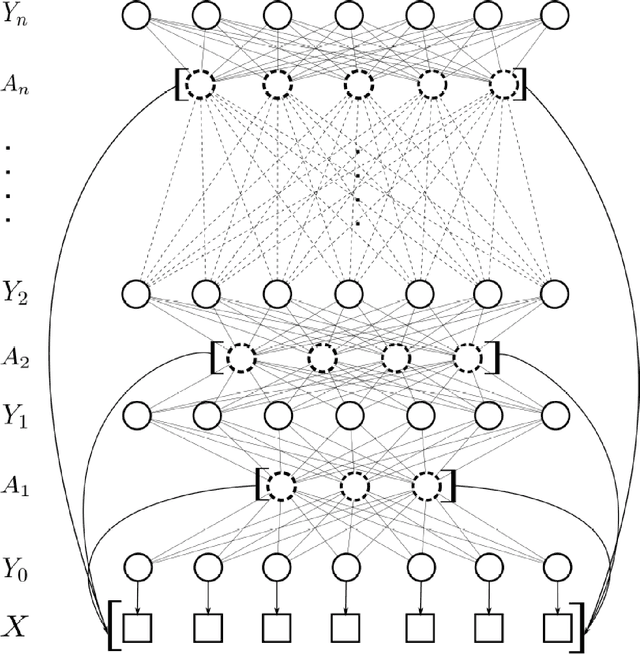
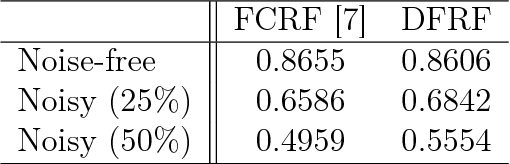

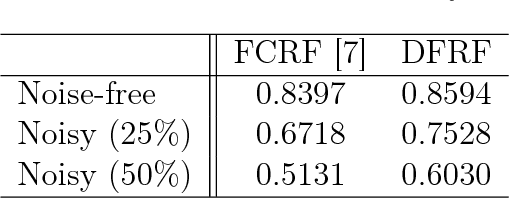
Abstract:There has been significant interest in the use of fully-connected graphical models and deep-structured graphical models for the purpose of structured inference. However, fully-connected and deep-structured graphical models have been largely explored independently, leaving the unification of these two concepts ripe for exploration. A fundamental challenge with unifying these two types of models is in dealing with computational complexity. In this study, we investigate the feasibility of unifying fully-connected and deep-structured models in a computationally tractable manner for the purpose of structured inference. To accomplish this, we introduce a deep-structured fully-connected random field (DFRF) model that integrates a series of intermediate sparse auto-encoding layers placed between state layers to significantly reduce computational complexity. The problem of image segmentation was used to illustrate the feasibility of using the DFRF for structured inference in a computationally tractable manner. Results in this study show that it is feasible to unify fully-connected and deep-structured models in a computationally tractable manner for solving structured inference problems such as image segmentation.
* Accepted, 13 pages
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge