Mohammad Javad Shafiee
Evolution in Groups: A deeper look at synaptic cluster driven evolution of deep neural networks
Apr 07, 2017

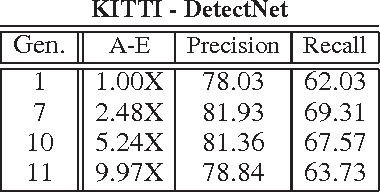

Abstract:A promising paradigm for achieving highly efficient deep neural networks is the idea of evolutionary deep intelligence, which mimics biological evolution processes to progressively synthesize more efficient networks. A crucial design factor in evolutionary deep intelligence is the genetic encoding scheme used to simulate heredity and determine the architectures of offspring networks. In this study, we take a deeper look at the notion of synaptic cluster-driven evolution of deep neural networks which guides the evolution process towards the formation of a highly sparse set of synaptic clusters in offspring networks. Utilizing a synaptic cluster-driven genetic encoding, the probabilistic encoding of synaptic traits considers not only individual synaptic properties but also inter-synaptic relationships within a deep neural network. This process results in highly sparse offspring networks which are particularly tailored for parallel computational devices such as GPUs and deep neural network accelerator chips. Comprehensive experimental results using four well-known deep neural network architectures (LeNet-5, AlexNet, ResNet-56, and DetectNet) on two different tasks (object categorization and object detection) demonstrate the efficiency of the proposed method. Cluster-driven genetic encoding scheme synthesizes networks that can achieve state-of-the-art performance with significantly smaller number of synapses than that of the original ancestor network. ($\sim$125-fold decrease in synapses for MNIST). Furthermore, the improved cluster efficiency in the generated offspring networks ($\sim$9.71-fold decrease in clusters for MNIST and a $\sim$8.16-fold decrease in clusters for KITTI) is particularly useful for accelerated performance on parallel computing hardware architectures such as those in GPUs and deep neural network accelerator chips.
Discovery Radiomics for Pathologically-Proven Computed Tomography Lung Cancer Prediction
Mar 28, 2017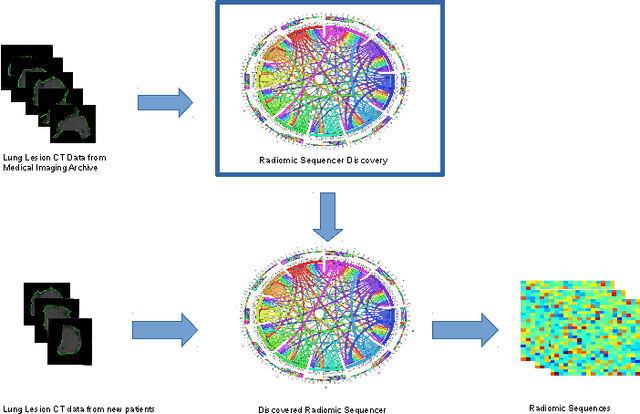
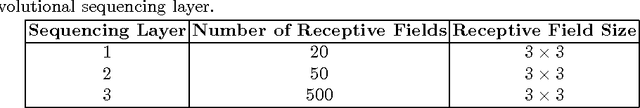
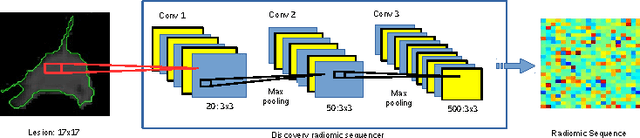
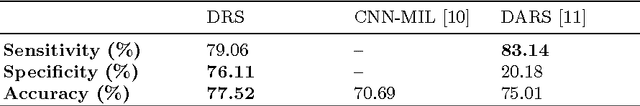
Abstract:Lung cancer is the leading cause for cancer related deaths. As such, there is an urgent need for a streamlined process that can allow radiologists to provide diagnosis with greater efficiency and accuracy. A powerful tool to do this is radiomics: a high-dimension imaging feature set. In this study, we take the idea of radiomics one step further by introducing the concept of discovery radiomics for lung cancer prediction using CT imaging data. In this study, we realize these custom radiomic sequencers as deep convolutional sequencers using a deep convolutional neural network learning architecture. To illustrate the prognostic power and effectiveness of the radiomic sequences produced by the discovered sequencer, we perform cancer prediction between malignant and benign lesions from 97 patients using the pathologically-proven diagnostic data from the LIDC-IDRI dataset. Using the clinically provided pathologically-proven data as ground truth, the proposed framework provided an average accuracy of 77.52% via 10-fold cross-validation with a sensitivity of 79.06% and specificity of 76.11%, surpassing the state-of-the art method.
Deep Learning with Darwin: Evolutionary Synthesis of Deep Neural Networks
Feb 06, 2017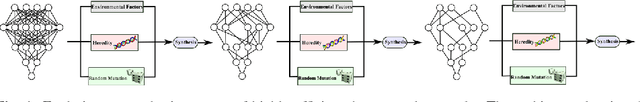
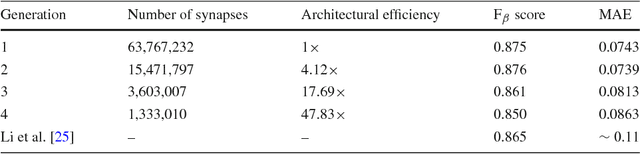

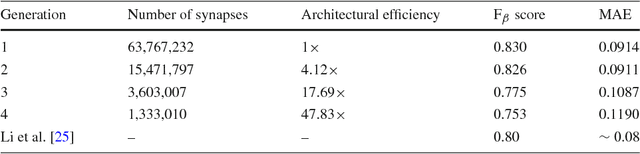
Abstract:Taking inspiration from biological evolution, we explore the idea of "Can deep neural networks evolve naturally over successive generations into highly efficient deep neural networks?" by introducing the notion of synthesizing new highly efficient, yet powerful deep neural networks over successive generations via an evolutionary process from ancestor deep neural networks. The architectural traits of ancestor deep neural networks are encoded using synaptic probability models, which can be viewed as the `DNA' of these networks. New descendant networks with differing network architectures are synthesized based on these synaptic probability models from the ancestor networks and computational environmental factor models, in a random manner to mimic heredity, natural selection, and random mutation. These offspring networks are then trained into fully functional networks, like one would train a newborn, and have more efficient, more diverse network architectures than their ancestor networks, while achieving powerful modeling capabilities. Experimental results for the task of visual saliency demonstrated that the synthesized `evolved' offspring networks can achieve state-of-the-art performance while having network architectures that are significantly more efficient (with a staggering $\sim$48-fold decrease in synapses by the fourth generation) compared to the original ancestor network.
Evolutionary Synthesis of Deep Neural Networks via Synaptic Cluster-driven Genetic Encoding
Nov 22, 2016
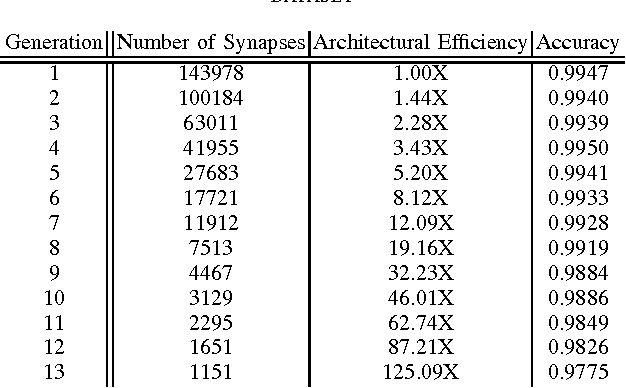
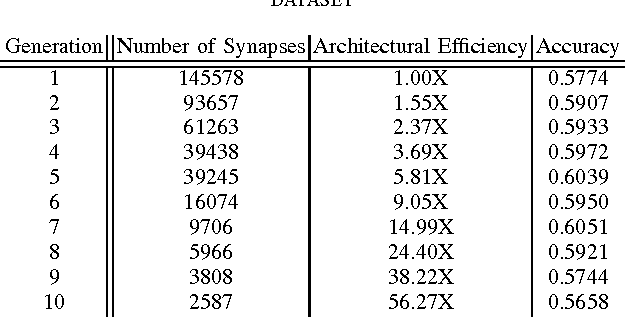
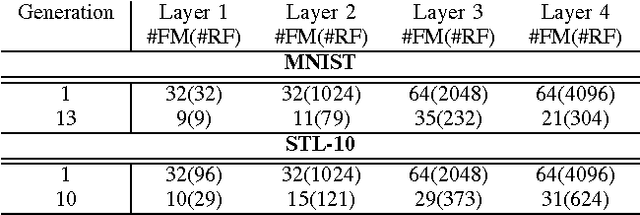
Abstract:There has been significant recent interest towards achieving highly efficient deep neural network architectures. A promising paradigm for achieving this is the concept of evolutionary deep intelligence, which attempts to mimic biological evolution processes to synthesize highly-efficient deep neural networks over successive generations. An important aspect of evolutionary deep intelligence is the genetic encoding scheme used to mimic heredity, which can have a significant impact on the quality of offspring deep neural networks. Motivated by the neurobiological phenomenon of synaptic clustering, we introduce a new genetic encoding scheme where synaptic probability is driven towards the formation of a highly sparse set of synaptic clusters. Experimental results for the task of image classification demonstrated that the synthesized offspring networks using this synaptic cluster-driven genetic encoding scheme can achieve state-of-the-art performance while having network architectures that are not only significantly more efficient (with a ~125-fold decrease in synapses for MNIST) compared to the original ancestor network, but also tailored for GPU-accelerated machine learning applications.
Noise-Compensated, Bias-Corrected Diffusion Weighted Endorectal Magnetic Resonance Imaging via a Stochastically Fully-Connected Joint Conditional Random Field Model
Jul 05, 2016
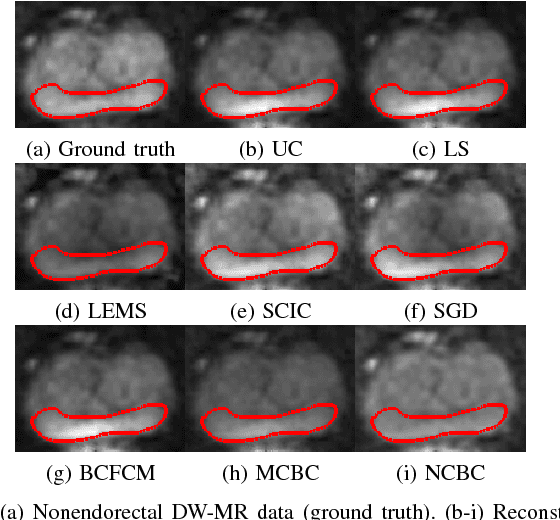
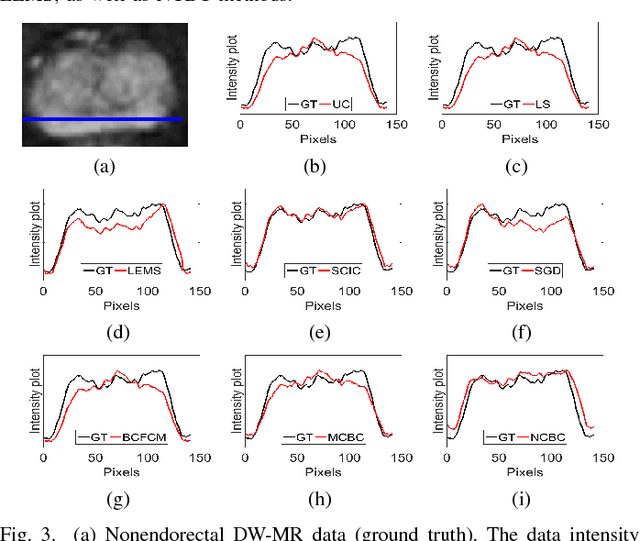
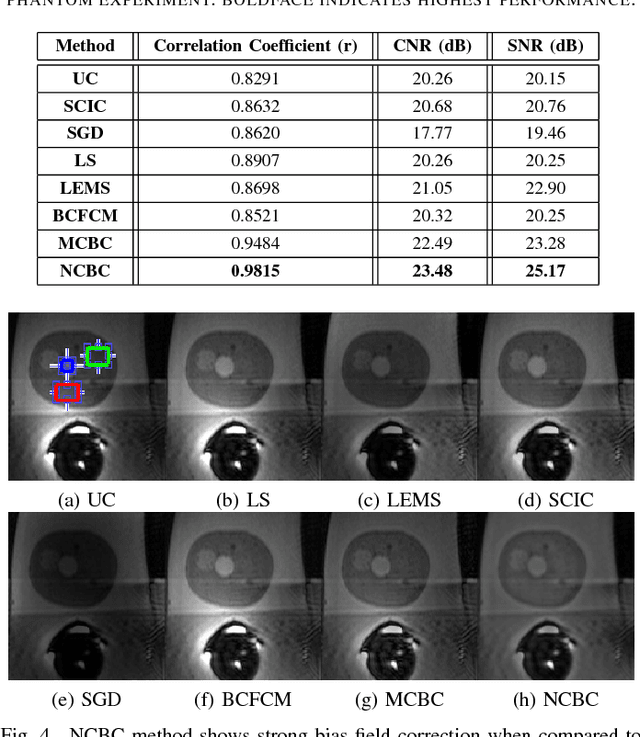
Abstract:Diffusion weighted magnetic resonance imaging (DW-MR) is a powerful tool in imaging-based prostate cancer screening and detection. Endorectal coils are commonly used in DW-MR imaging to improve the signal-to-noise ratio (SNR) of the acquisition, at the expense of significant intensity inhomogeneities (bias field) that worsens as we move away from the endorectal coil. The presence of bias field can have a significant negative impact on the accuracy of different image analysis tasks, as well as prostate tumor localization, thus leading to increased inter- and intra-observer variability. Retrospective bias correction approaches are introduced as a more efficient way of bias correction compared to the prospective methods such that they correct for both of the scanner and anatomy-related bias fields in MR imaging. Previously proposed retrospective bias field correction methods suffer from undesired noise amplification that can reduce the quality of bias-corrected DW-MR image. Here, we propose a unified data reconstruction approach that enables joint compensation of bias field as well as data noise in DW-MR imaging. The proposed noise-compensated, bias-corrected (NCBC) data reconstruction method takes advantage of a novel stochastically fully connected joint conditional random field (SFC-JCRF) model to mitigate the effects of data noise and bias field in the reconstructed MR data. The proposed NCBC reconstruction method was tested on synthetic DW-MR data, physical DW-phantom as well as real DW-MR data all acquired using endorectal MR coil. Both qualitative and quantitative analysis illustrated that the proposed NCBC method can achieve improved image quality when compared to other tested bias correction methods. As such, the proposed NCBC method may have potential as a useful retrospective approach for improving the consistency of image interpretations.
Scene Invariant Crowd Segmentation and Counting Using Scale-Normalized Histogram of Moving Gradients (HoMG)
Feb 01, 2016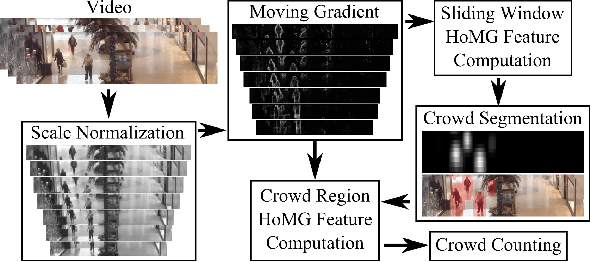
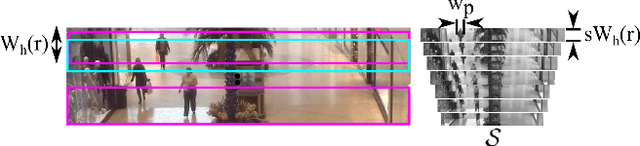
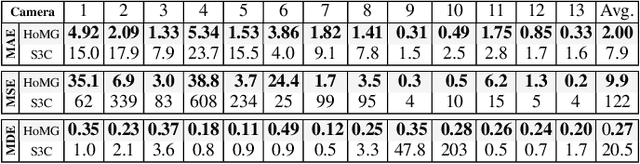

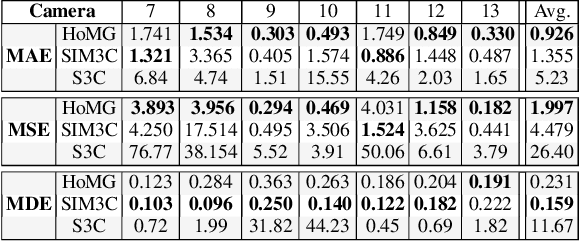
Abstract:The problem of automated crowd segmentation and counting has garnered significant interest in the field of video surveillance. This paper proposes a novel scene invariant crowd segmentation and counting algorithm designed with high accuracy yet low computational complexity in mind, which is key for widespread industrial adoption. A novel low-complexity, scale-normalized feature called Histogram of Moving Gradients (HoMG) is introduced for highly effective spatiotemporal representation of individuals and crowds within a video. Real-time crowd segmentation is achieved via boosted cascade of weak classifiers based on sliding-window HoMG features, while linear SVM regression of crowd-region HoMG features is employed for real-time crowd counting. Experimental results using multi-camera crowd datasets show that the proposed algorithm significantly outperform state-of-the-art crowd counting algorithms, as well as achieve very promising crowd segmentation results, thus demonstrating the efficacy of the proposed method for highly-accurate, real-time video-driven crowd analysis.
Sparse Reconstruction of Compressive Sensing MRI using Cross-Domain Stochastically Fully Connected Conditional Random Fields
Dec 25, 2015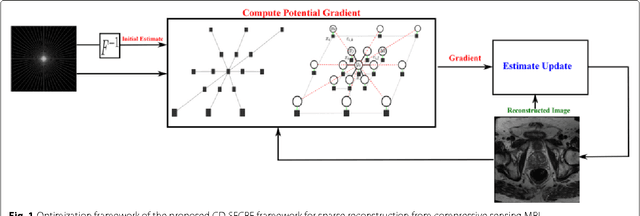
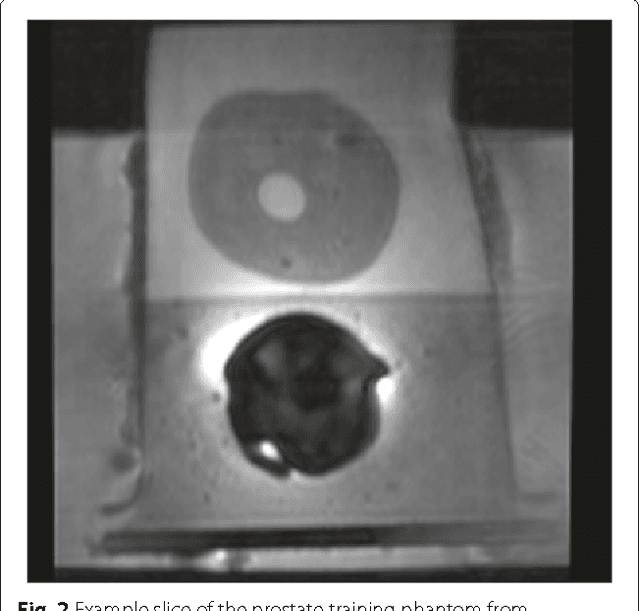


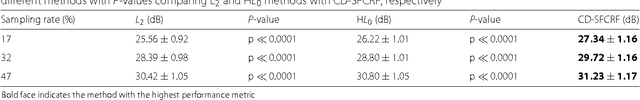
Abstract:Magnetic Resonance Imaging (MRI) is a crucial medical imaging technology for the screening and diagnosis of frequently occurring cancers. However image quality may suffer by long acquisition times for MRIs due to patient motion, as well as result in great patient discomfort. Reducing MRI acquisition time can reduce patient discomfort and as a result reduces motion artifacts from the acquisition process. Compressive sensing strategies, when applied to MRI, have been demonstrated to be effective at decreasing acquisition times significantly by sparsely sampling the \emph{k}-space during the acquisition process. However, such a strategy requires advanced reconstruction algorithms to produce high quality and reliable images from compressive sensing MRI. This paper proposes a new reconstruction approach based on cross-domain stochastically fully connected conditional random fields (CD-SFCRF) for compressive sensing MRI. The CD-SFCRF introduces constraints in both \emph{k}-space and spatial domains within a stochastically fully connected graphical model to produce improved MRI reconstruction. Experimental results using T2-weighted (T2w) imaging and diffusion-weighted imaging (DWI) of the prostate show strong performance in preserving fine details and tissue structures in the reconstructed images when compared to other tested methods even at low sampling rates.
Domain Adaptation and Transfer Learning in StochasticNets
Dec 18, 2015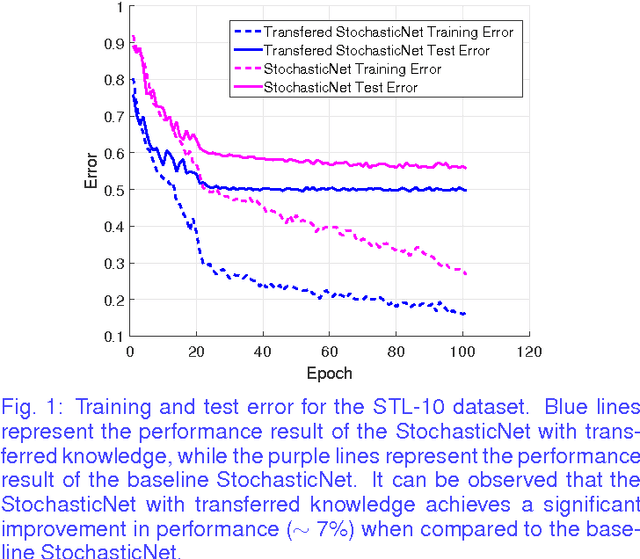
Abstract:Transfer learning is a recent field of machine learning research that aims to resolve the challenge of dealing with insufficient training data in the domain of interest. This is a particular issue with traditional deep neural networks where a large amount of training data is needed. Recently, StochasticNets was proposed to take advantage of sparse connectivity in order to decrease the number of parameters that needs to be learned, which in turn may relax training data size requirements. In this paper, we study the efficacy of transfer learning on StochasticNet frameworks. Experimental results show ~7% improvement on StochasticNet performance when the transfer learning is applied in training step.
Efficient Deep Feature Learning and Extraction via StochasticNets
Dec 11, 2015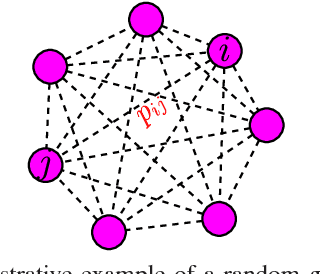
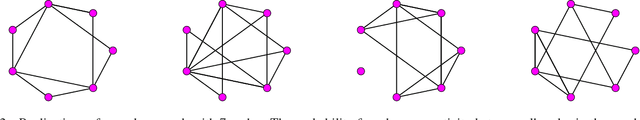
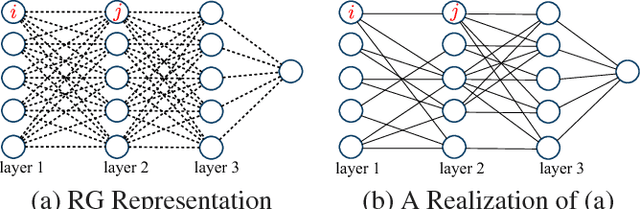
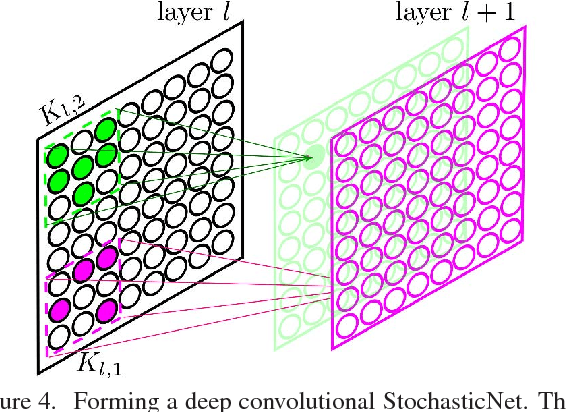
Abstract:Deep neural networks are a powerful tool for feature learning and extraction given their ability to model high-level abstractions in highly complex data. One area worth exploring in feature learning and extraction using deep neural networks is efficient neural connectivity formation for faster feature learning and extraction. Motivated by findings of stochastic synaptic connectivity formation in the brain as well as the brain's uncanny ability to efficiently represent information, we propose the efficient learning and extraction of features via StochasticNets, where sparsely-connected deep neural networks can be formed via stochastic connectivity between neurons. To evaluate the feasibility of such a deep neural network architecture for feature learning and extraction, we train deep convolutional StochasticNets to learn abstract features using the CIFAR-10 dataset, and extract the learned features from images to perform classification on the SVHN and STL-10 datasets. Experimental results show that features learned using deep convolutional StochasticNets, with fewer neural connections than conventional deep convolutional neural networks, can allow for better or comparable classification accuracy than conventional deep neural networks: relative test error decrease of ~4.5% for classification on the STL-10 dataset and ~1% for classification on the SVHN dataset. Furthermore, it was shown that the deep features extracted using deep convolutional StochasticNets can provide comparable classification accuracy even when only 10% of the training data is used for feature learning. Finally, it was also shown that significant gains in feature extraction speed can be achieved in embedded applications using StochasticNets. As such, StochasticNets allow for faster feature learning and extraction performance while facilitate for better or comparable accuracy performances.
Discovery Radiomics via StochasticNet Sequencers for Cancer Detection
Nov 11, 2015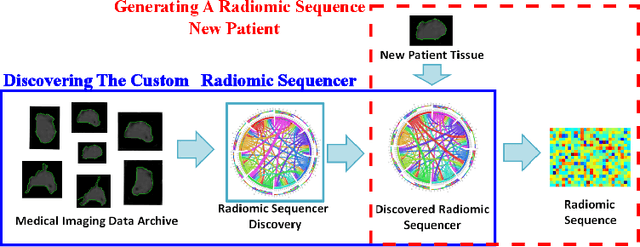
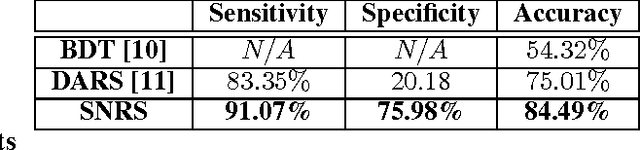
Abstract:Radiomics has proven to be a powerful prognostic tool for cancer detection, and has previously been applied in lung, breast, prostate, and head-and-neck cancer studies with great success. However, these radiomics-driven methods rely on pre-defined, hand-crafted radiomic feature sets that can limit their ability to characterize unique cancer traits. In this study, we introduce a novel discovery radiomics framework where we directly discover custom radiomic features from the wealth of available medical imaging data. In particular, we leverage novel StochasticNet radiomic sequencers for extracting custom radiomic features tailored for characterizing unique cancer tissue phenotype. Using StochasticNet radiomic sequencers discovered using a wealth of lung CT data, we perform binary classification on 42,340 lung lesions obtained from the CT scans of 93 patients in the LIDC-IDRI dataset. Preliminary results show significant improvement over previous state-of-the-art methods, indicating the potential of the proposed discovery radiomics framework for improving cancer screening and diagnosis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge