Why Deep Surgical Models Fail?: Revisiting Surgical Action Triplet Recognition through the Lens of Robustness
Sep 18, 2022Yanqi Cheng, Lihao Liu, Shujun Wang, Yueming Jin, Carola-Bibiane Schönlieb, Angelica I. Aviles-Rivero
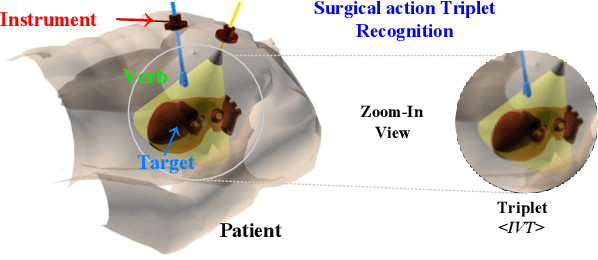
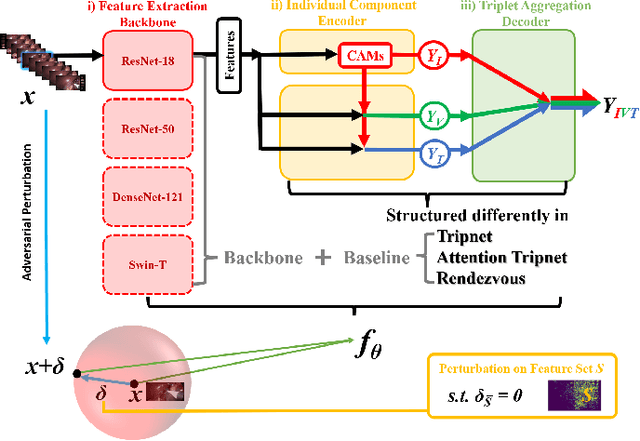
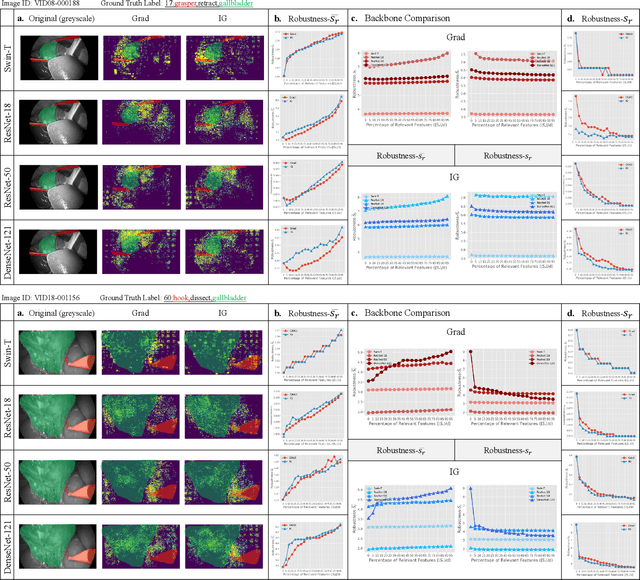
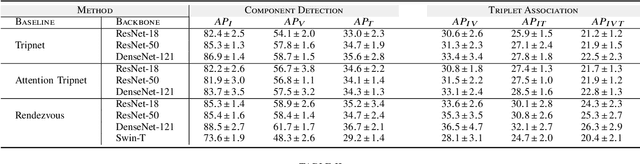
Surgical action triplet recognition provides a better understanding of the surgical scene. This task is of high relevance as it provides to the surgeon with context-aware support and safety. The current go-to strategy for improving performance is the development of new network mechanisms. However, the performance of current state-of-the-art techniques is substantially lower than other surgical tasks. Why is this happening? This is the question that we address in this work. We present the first study to understand the failure of existing deep learning models through the lens of robustness and explainabilty. Firstly, we study current existing models under weak and strong $\delta-$perturbations via adversarial optimisation scheme. We then provide the failure modes via feature based explanations. Our study revels that the key for improving performance and increasing reliability is in the core and spurious attributes. Our work opens the door to more trustworthiness and reliability deep learning models in surgical science.
LAMBDA: Covering the Solution Set of Black-Box Inequality by Search Space Quantization
Mar 25, 2022Lihao Liu, Tianyue Feng, Xingyu Xing, Junyi Chen
Black-box functions are broadly used to model complex problems that provide no explicit information but the input and output. Despite existing studies of black-box function optimization, the solution set satisfying an inequality with a black-box function plays a more significant role than only one optimum in many practical situations. Covering as much as possible of the solution set through limited evaluations to the black-box objective function is defined as the Black-Box Coverage (BBC) problem in this paper. We formalized this problem in a sample-based search paradigm and constructed a coverage criterion with Confusion Matrix Analysis. Further, we propose LAMBDA (Latent-Action Monte-Carlo Beam Search with Density Adaption) to solve BBC problems. LAMBDA can focus around the solution set quickly by recursively partitioning the search space into accepted and rejected sub-spaces. Compared with La-MCTS, LAMBDA introduces density information to overcome the sampling bias of optimization and obtain more exploration. Benchmarking shows, LAMBDA achieved state-of-the-art performance among all baselines and was at most 33x faster to get 95% coverage than Random Search. Experiments also demonstrate that LAMBDA has a promising future in the verification of autonomous systems in virtual tests.
PC-SwinMorph: Patch Representation for Unsupervised Medical Image Registration and Segmentation
Mar 10, 2022Lihao Liu, Zhening Huang, Pietro Liò, Carola-Bibiane Schönlieb, Angelica I. Aviles-Rivero
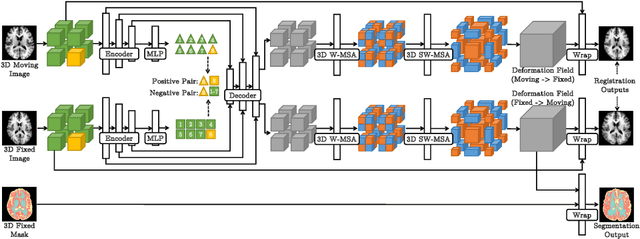
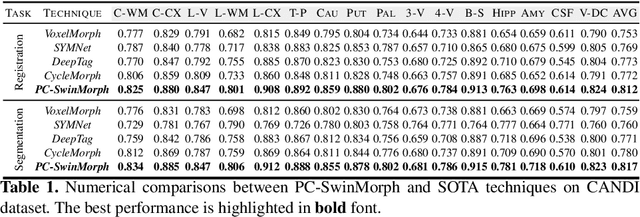
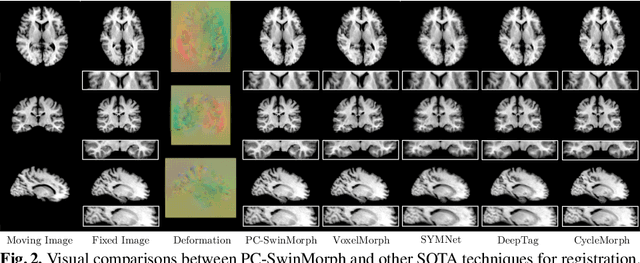
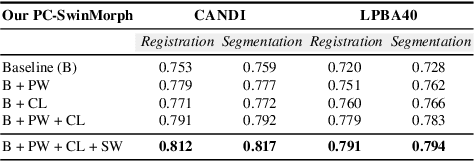
Medical image registration and segmentation are critical tasks for several clinical procedures. Manual realisation of those tasks is time-consuming and the quality is highly dependent on the level of expertise of the physician. To mitigate that laborious task, automatic tools have been developed where the majority of solutions are supervised techniques. However, in medical domain, the strong assumption of having a well-representative ground truth is far from being realistic. To overcome this challenge, unsupervised techniques have been investigated. However, they are still limited in performance and they fail to produce plausible results. In this work, we propose a novel unified unsupervised framework for image registration and segmentation that we called PC-SwinMorph. The core of our framework is two patch-based strategies, where we demonstrate that patch representation is key for performance gain. We first introduce a patch-based contrastive strategy that enforces locality conditions and richer feature representation. Secondly, we utilise a 3D window/shifted-window multi-head self-attention module as a patch stitching strategy to eliminate artifacts from the patch splitting. We demonstrate, through a set of numerical and visual results, that our technique outperforms current state-of-the-art unsupervised techniques.
Simultaneous Semantic and Instance Segmentation for Colon Nuclei Identification and Counting
Mar 01, 2022Lihao Liu, Chenyang Hong, Angelica I. Aviles-Rivero, Carola-Bibiane Schönlieb
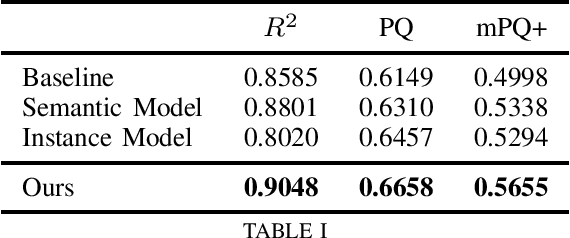
We address the problem of automated nuclear segmentation, classification, and quantification from Haematoxylin and Eosin stained histology images, which is of great relevance for several downstream computational pathology applications. In this work, we present a solution framed as a simultaneous semantic and instance segmentation framework. Our solution is part of the Colon Nuclei Identification and Counting (CoNIC) Challenge. We first train a semantic and instance segmentation model separately. Our framework uses as backbone HoverNet and Cascade Mask-RCNN models. We then ensemble the results with a custom Non-Maximum Suppression embedding (NMS). In our framework, the semantic model computes a class prediction for the cells whilst the instance model provides a refined segmentation. We demonstrate, through our experimental results, that our model outperforms the provided baselines by a large margin.
Contrastive Registration for Unsupervised Medical Image Segmentation
Nov 17, 2020Lihao Liu, Angelica I Aviles-Rivero, Carola-Bibiane Schönlieb
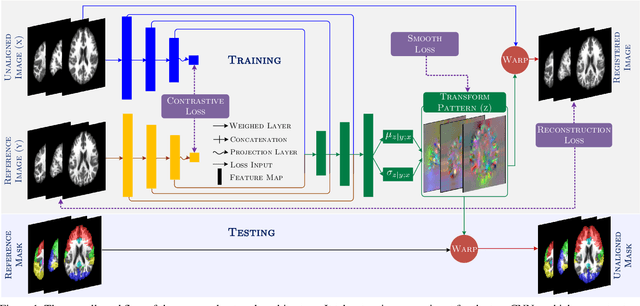
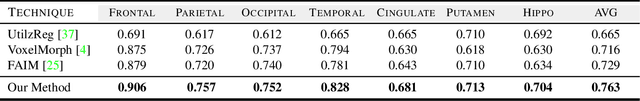
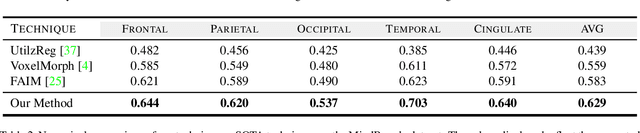
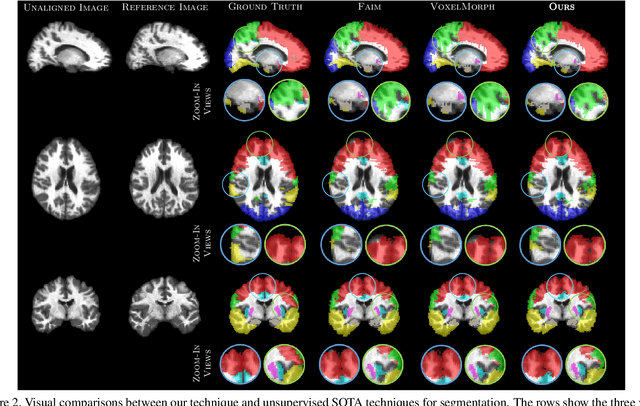
Medical image segmentation is a relevant task as it serves as the first step for several diagnosis processes, thus it is indispensable in clinical usage. Whilst major success has been reported using supervised techniques, they assume a large and well-representative labelled set. This is a strong assumption in the medical domain where annotations are expensive, time-consuming, and inherent to human bias. To address this problem, unsupervised techniques have been proposed in the literature yet it is still an open problem due to the difficulty of learning any transformation pattern. In this work, we present a novel optimisation model framed into a new CNN-based contrastive registration architecture for unsupervised medical image segmentation. The core of our approach is to exploit image-level registration and feature-level from a contrastive learning mechanism, to perform registration-based segmentation. Firstly, we propose an architecture to capture the image-to-image transformation pattern via registration for unsupervised medical image segmentation. Secondly, we embed a contrastive learning mechanism into the registration architecture to enhance the discriminating capacity of the network in the feature-level. We show that our proposed technique mitigates the major drawbacks of existing unsupervised techniques. We demonstrate, through numerical and visual experiments, that our technique substantially outperforms the current state-of-the-art unsupervised segmentation methods on two major medical image datasets.
Probabilistic Multilayer Regularization Network for Unsupervised 3D Brain Image Registration
Jul 03, 2019Lihao Liu, Xiaowei Hu, Lei Zhu, Pheng-Ann Heng
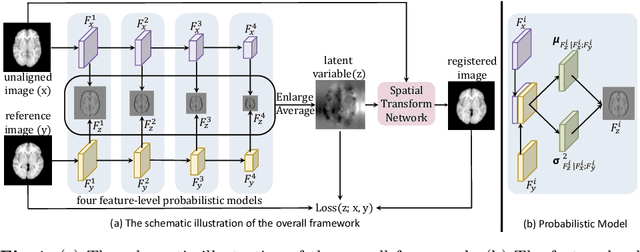
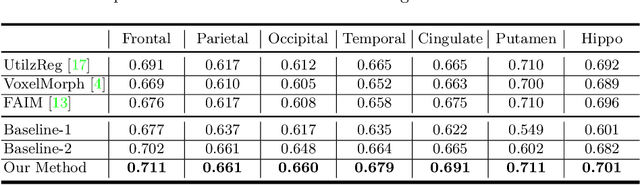
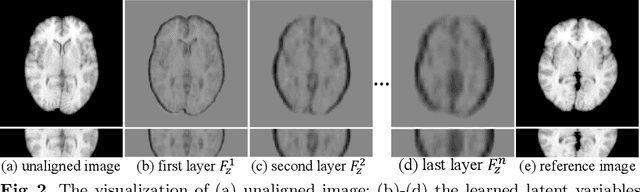
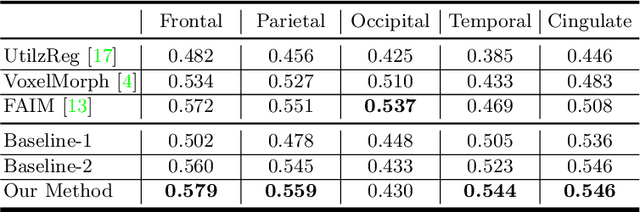
Brain image registration transforms a pair of images into one system with the matched imaging contents, which is of essential importance for brain image analysis. This paper presents a novel framework for unsupervised 3D brain image registration by capturing the feature-level transformation relationships between the unaligned image and reference image. To achieve this, we develop a feature-level probabilistic model to provide the direct regularization to the hidden layers of two deep convolutional neural networks, which are constructed from two input images. This model design is developed into multiple layers of these two networks to capture the transformation relationships at different levels. We employ two common benchmark datasets for 3D brain image registration and perform various experiments to evaluate our method. Experimental results show that our method clearly outperforms state-of-the-art methods on both benchmark datasets by a large margin.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge