Globus Automation Services: Research process automation across the space-time continuum
Aug 19, 2022Ryan Chard, Jim Pruyne, Kurt McKee, Josh Bryan, Brigitte Raumann, Rachana Ananthakrishnan, Kyle Chard, Ian Foster
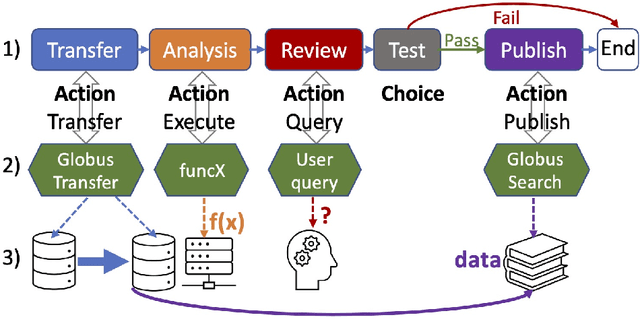
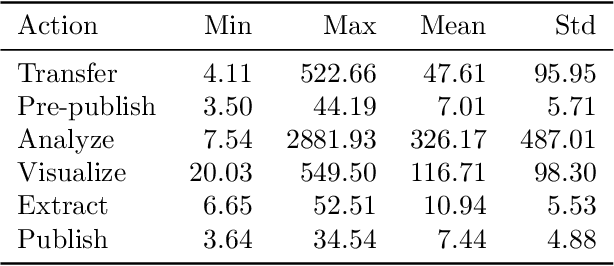
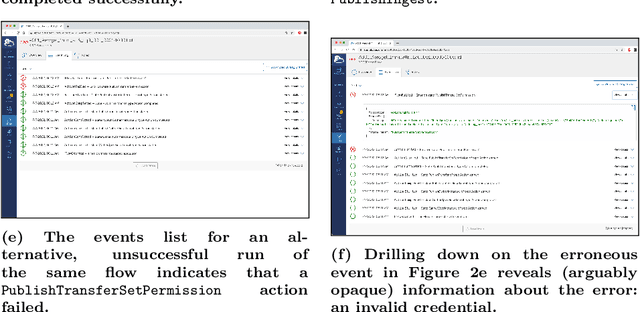
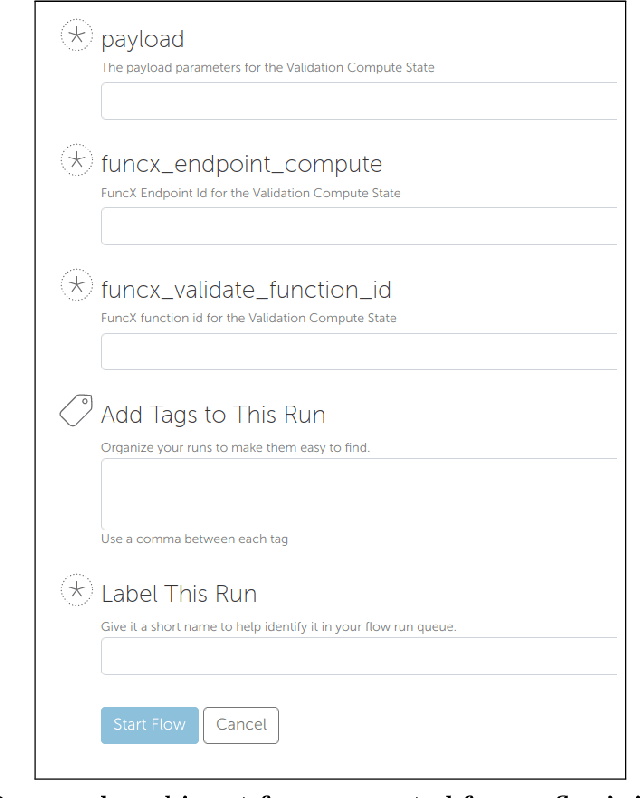
Research process automation--the reliable, efficient, and reproducible execution of linked sets of actions on scientific instruments, computers, data stores, and other resources--has emerged as an essential element of modern science. We report here on new services within the Globus research data management platform that enable the specification of diverse research processes as reusable sets of actions, flows, and the execution of such flows in heterogeneous research environments. To support flows with broad spatial extent (e.g., from scientific instrument to remote data center) and temporal extent (from seconds to weeks), these Globus automation services feature: 1) cloud hosting for reliable execution of even long-lived flows despite sporadic failures; 2) a declarative notation, and extensible asynchronous action provider API, for defining and executing a wide variety of actions and flow specifications involving arbitrary resources; 3) authorization delegation mechanisms for secure invocation of actions. These services permit researchers to outsource and automate the management of a broad range of research tasks to a reliable, scalable, and secure cloud platform. We present use cases for Globus automation services, describe the design and implementation of the services, present microbenchmark studies, and review experiences applying the services in a range of applications
FAIR principles for AI models, with a practical application for accelerated high energy diffraction microscopy
Jul 14, 2022Nikil Ravi, Pranshu Chaturvedi, E. A. Huerta, Zhengchun Liu, Ryan Chard, Aristana Scourtas, K. J. Schmidt, Kyle Chard, Ben Blaiszik, Ian Foster
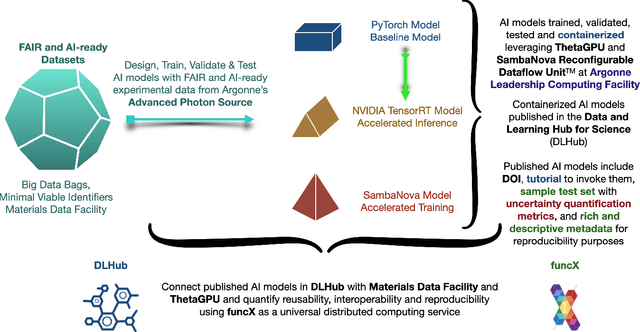
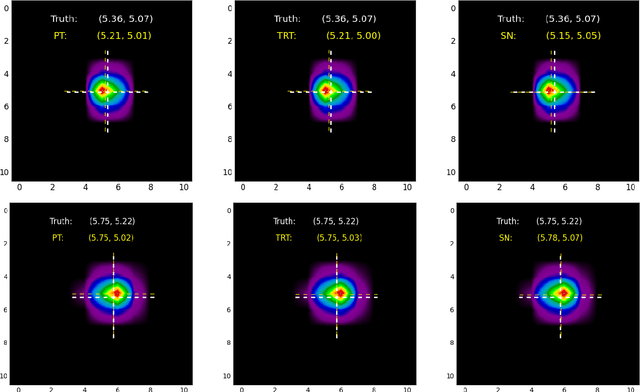
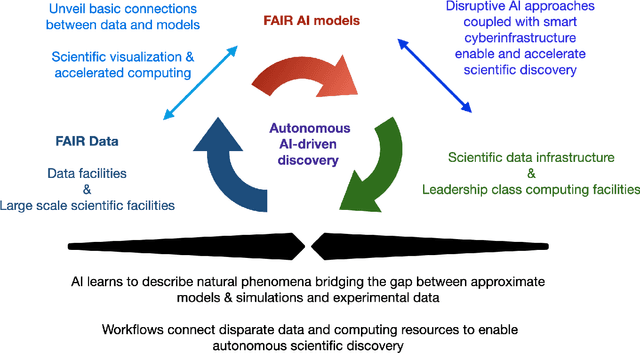
A concise and measurable set of FAIR (Findable, Accessible, Interoperable and Reusable) principles for scientific data is transforming the state-of-practice for data management and stewardship, supporting and enabling discovery and innovation. Learning from this initiative, and acknowledging the impact of artificial intelligence (AI) in the practice of science and engineering, we introduce a set of practical, concise, and measurable FAIR principles for AI models. We showcase how to create and share FAIR data and AI models within a unified computational framework combining the following elements: the Advanced Photon Source at Argonne National Laboratory, the Materials Data Facility, the Data and Learning Hub for Science, and funcX, and the Argonne Leadership Computing Facility (ALCF), in particular the ThetaGPU supercomputer and the SambaNova DataScale system at the ALCF AI Testbed. We describe how this domain-agnostic computational framework may be harnessed to enable autonomous AI-driven discovery.
ScholarBERT: Bigger is Not Always Better
May 23, 2022Zhi Hong, Aswathy Ajith, Gregory Pauloski, Eamon Duede, Carl Malamud, Roger Magoulas, Kyle Chard, Ian Foster
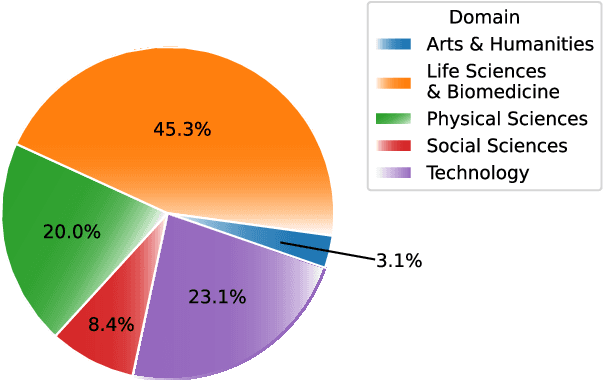
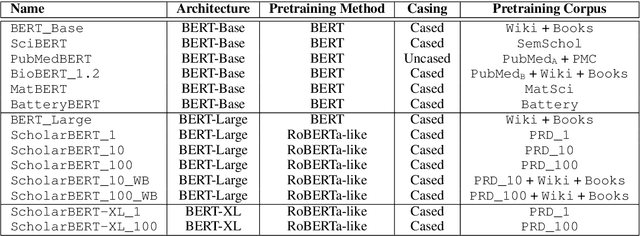

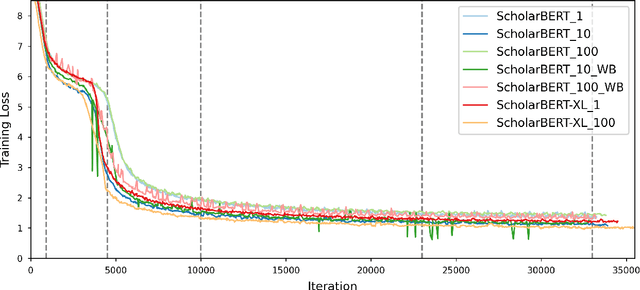
Transformer-based masked language models trained on general corpora, such as BERT and RoBERTa, have shown impressive performance on various downstream tasks. Increasingly, researchers are "finetuning" these models to improve performance on domain-specific tasks. Here, we report a broad study in which we applied 14 transformer-based models to 11 scientific tasks in order to evaluate how downstream performance is affected by changes along various dimensions (e.g., training data, model size, pretraining time, finetuning length). In this process, we created the largest and most diverse scientific language model to date, ScholarBERT, by training a 770M-parameter BERT model on an 221B token scientific literature dataset spanning many disciplines. Counterintuitively, our evaluation of the 14 BERT-based models (seven versions of ScholarBERT, five science-specific large language models from the literature, BERT-Base, and BERT-Large) reveals little difference in performance across the 11 science-focused tasks, despite major differences in model size and training data. We argue that our results establish an upper bound for the performance achievable with BERT-based architectures on tasks from the scientific domain.
Colmena: Scalable Machine-Learning-Based Steering of Ensemble Simulations for High Performance Computing
Oct 06, 2021Logan Ward, Ganesh Sivaraman, J. Gregory Pauloski, Yadu Babuji, Ryan Chard, Naveen Dandu, Paul C. Redfern, Rajeev S. Assary, Kyle Chard, Larry A. Curtiss, Rajeev Thakur, Ian Foster
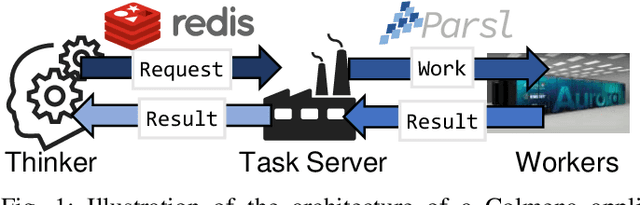
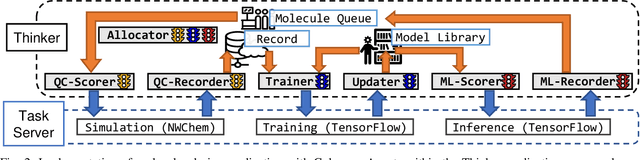
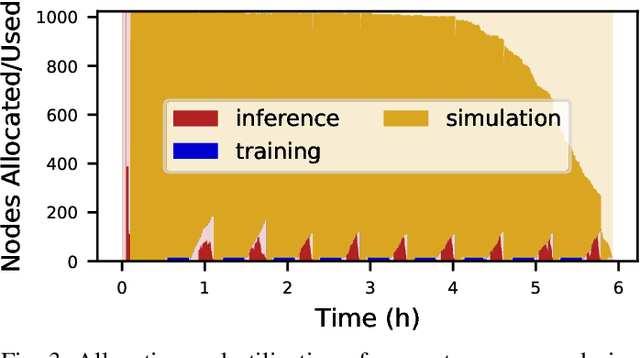
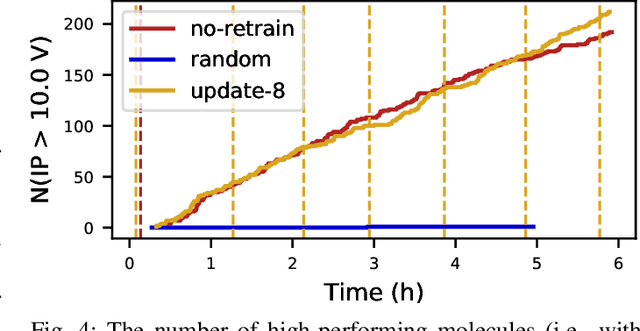
Scientific applications that involve simulation ensembles can be accelerated greatly by using experiment design methods to select the best simulations to perform. Methods that use machine learning (ML) to create proxy models of simulations show particular promise for guiding ensembles but are challenging to deploy because of the need to coordinate dynamic mixes of simulation and learning tasks. We present Colmena, an open-source Python framework that allows users to steer campaigns by providing just the implementations of individual tasks plus the logic used to choose which tasks to execute when. Colmena handles task dispatch, results collation, ML model invocation, and ML model (re)training, using Parsl to execute tasks on HPC systems. We describe the design of Colmena and illustrate its capabilities by applying it to electrolyte design, where it both scales to 65536 CPUs and accelerates the discovery rate for high-performance molecules by a factor of 100 over unguided searches.
KAISA: An Adaptive Second-order Optimizer Framework for Deep Neural Networks
Jul 04, 2021J. Gregory Pauloski, Qi Huang, Lei Huang, Shivaram Venkataraman, Kyle Chard, Ian Foster, Zhao Zhang




Kronecker-factored Approximate Curvature (K-FAC) has recently been shown to converge faster in deep neural network (DNN) training than stochastic gradient descent (SGD); however, K-FAC's larger memory footprint hinders its applicability to large models. We present KAISA, a K-FAC-enabled, Adaptable, Improved, and ScAlable second-order optimizer framework that adapts the memory footprint, communication, and computation given specific models and hardware to achieve maximized performance and enhanced scalability. We quantify the tradeoffs between memory and communication cost and evaluate KAISA on large models, including ResNet-50, Mask R-CNN, U-Net, and BERT, on up to 128 NVIDIA A100 GPUs. Compared to the original optimizers, KAISA converges 18.1-36.3% faster across applications with the same global batch size. Under a fixed memory budget, KAISA converges 32.5% and 41.6% faster in ResNet-50 and BERT-Large, respectively. KAISA can balance memory and communication to achieve scaling efficiency equal to or better than the baseline optimizers.
AI- and HPC-enabled Lead Generation for SARS-CoV-2: Models and Processes to Extract Druglike Molecules Contained in Natural Language Text
Jan 12, 2021Zhi Hong, J. Gregory Pauloski, Logan Ward, Kyle Chard, Ben Blaiszik, Ian Foster
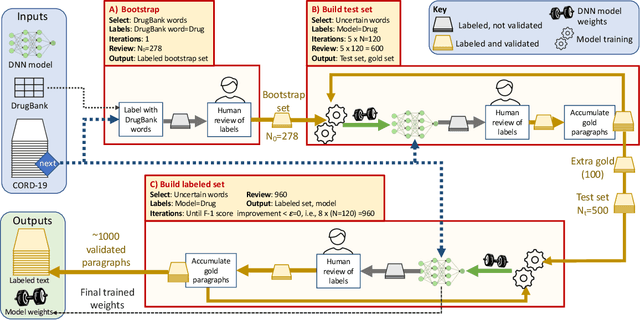
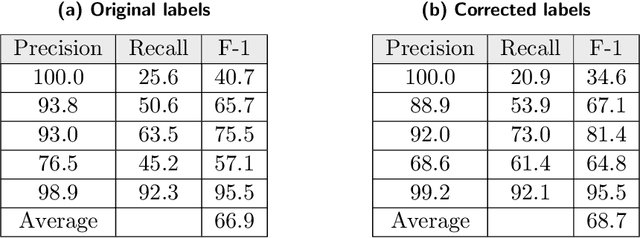
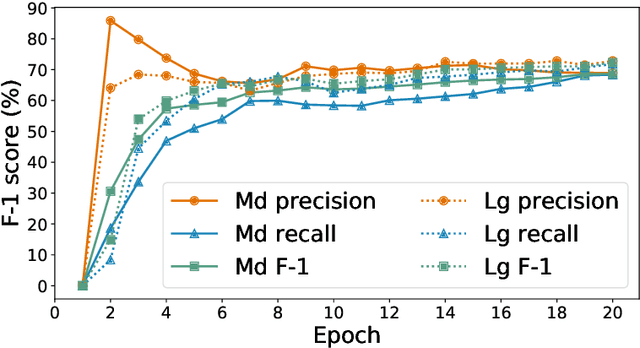
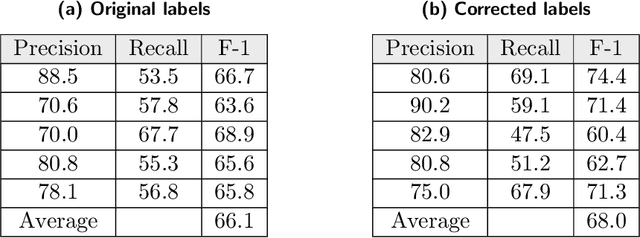
Researchers worldwide are seeking to repurpose existing drugs or discover new drugs to counter the disease caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). A promising source of candidates for such studies is molecules that have been reported in the scientific literature to be drug-like in the context of coronavirus research. We report here on a project that leverages both human and artificial intelligence to detect references to drug-like molecules in free text. We engage non-expert humans to create a corpus of labeled text, use this labeled corpus to train a named entity recognition model, and employ the trained model to extract 10912 drug-like molecules from the COVID-19 Open Research Dataset Challenge (CORD-19) corpus of 198875 papers. Performance analyses show that our automated extraction model can achieve performance on par with that of non-expert humans.
Towards Online Steering of Flame Spray Pyrolysis Nanoparticle Synthesis
Oct 16, 2020Maksim Levental, Ryan Chard, Joseph A. Libera, Kyle Chard, Aarthi Koripelly, Jakob R. Elias, Marcus Schwarting, Ben Blaiszik, Marius Stan, Santanu Chaudhuri, Ian Foster
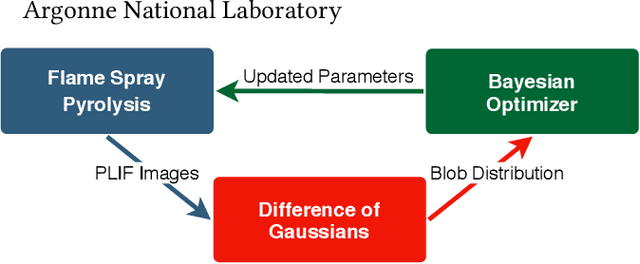
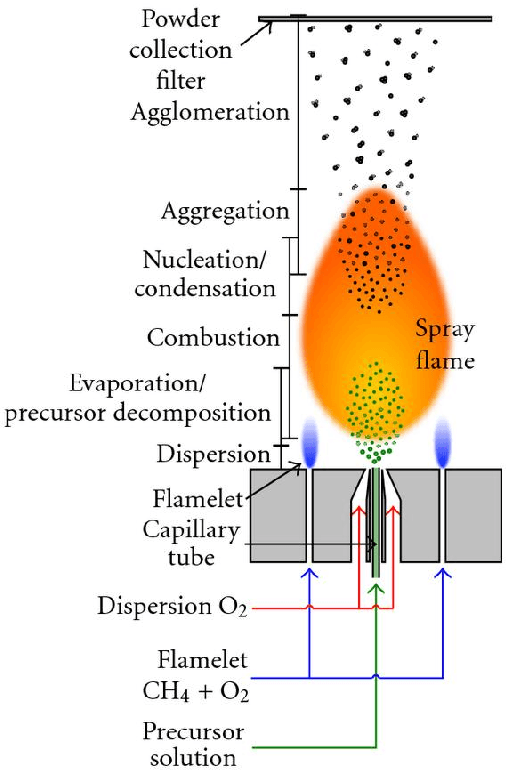

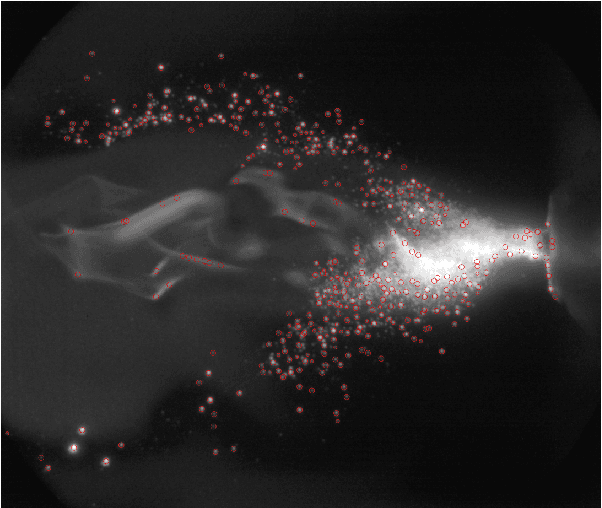
Flame Spray Pyrolysis (FSP) is a manufacturing technique to mass produce engineered nanoparticles for applications in catalysis, energy materials, composites, and more. FSP instruments are highly dependent on a number of adjustable parameters, including fuel injection rate, fuel-oxygen mixtures, and temperature, which can greatly affect the quality, quantity, and properties of the yielded nanoparticles. Optimizing FSP synthesis requires monitoring, analyzing, characterizing, and modifying experimental conditions.Here, we propose a hybrid CPU-GPU Difference of Gaussians (DoG)method for characterizing the volume distribution of unburnt solution, so as to enable near-real-time optimization and steering of FSP experiments. Comparisons against standard implementations show our method to be an order of magnitude more efficient. This surrogate signal can be deployed as a component of an online end-to-end pipeline that maximizes the synthesis yield.
Targeting SARS-CoV-2 with AI- and HPC-enabled Lead Generation: A First Data Release
May 28, 2020Yadu Babuji, Ben Blaiszik, Tom Brettin, Kyle Chard, Ryan Chard, Austin Clyde, Ian Foster, Zhi Hong, Shantenu Jha, Zhuozhao Li, Xuefeng Liu, Arvind Ramanathan, Yi Ren, Nicholaus Saint, Marcus Schwarting, Rick Stevens, Hubertus van Dam, Rick Wagner
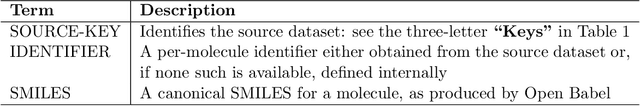
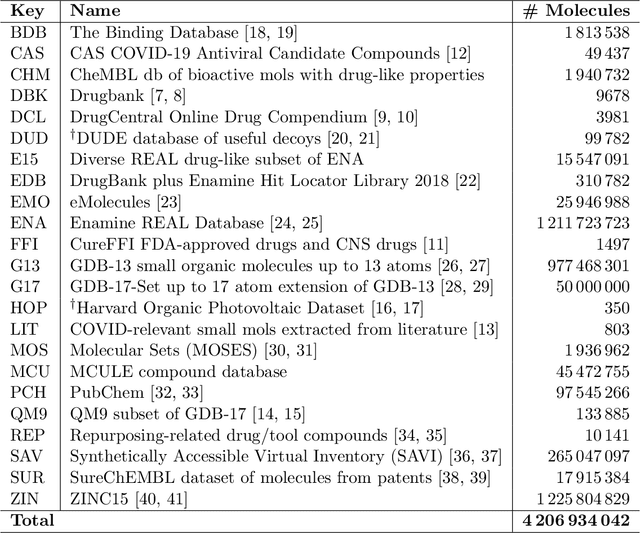
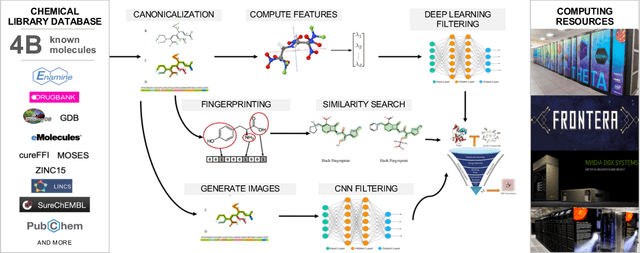

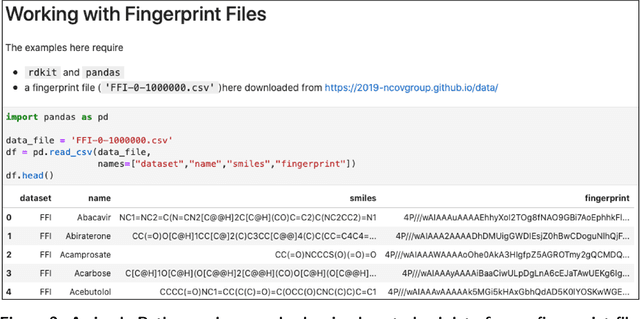
Researchers across the globe are seeking to rapidly repurpose existing drugs or discover new drugs to counter the the novel coronavirus disease (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). One promising approach is to train machine learning (ML) and artificial intelligence (AI) tools to screen large numbers of small molecules. As a contribution to that effort, we are aggregating numerous small molecules from a variety of sources, using high-performance computing (HPC) to computer diverse properties of those molecules, using the computed properties to train ML/AI models, and then using the resulting models for screening. In this first data release, we make available 23 datasets collected from community sources representing over 4.2 B molecules enriched with pre-computed: 1) molecular fingerprints to aid similarity searches, 2) 2D images of molecules to enable exploration and application of image-based deep learning methods, and 3) 2D and 3D molecular descriptors to speed development of machine learning models. This data release encompasses structural information on the 4.2 B molecules and 60 TB of pre-computed data. Future releases will expand the data to include more detailed molecular simulations, computed models, and other products.
Enabling real-time multi-messenger astrophysics discoveries with deep learning
Nov 26, 2019E. A. Huerta, Gabrielle Allen, Igor Andreoni, Javier M. Antelis, Etienne Bachelet, Bruce Berriman, Federica Bianco, Rahul Biswas, Matias Carrasco, Kyle Chard, Minsik Cho, Philip S. Cowperthwaite, Zachariah B. Etienne, Maya Fishbach, Francisco Förster, Daniel George, Tom Gibbs, Matthew Graham, William Gropp, Robert Gruendl, Anushri Gupta, Roland Haas, Sarah Habib, Elise Jennings, Margaret W. G. Johnson, Erik Katsavounidis, Daniel S. Katz, Asad Khan, Volodymyr Kindratenko, William T. C. Kramer, Xin Liu, Ashish Mahabal, Zsuzsa Marka, Kenton McHenry, Jonah Miller, Claudia Moreno, Mark Neubauer, Steve Oberlin, Alexander R. Olivas, Donald Petravick, Adam Rebei, Shawn Rosofsky, Milton Ruiz, Aaron Saxton, Bernard F. Schutz, Alex Schwing, Ed Seidel, Stuart L. Shapiro, Hongyu Shen, Yue Shen, Leo Singer, Brigitta M. Sipőcz, Lunan Sun, John Towns, Antonios Tsokaros, Wei Wei, Jack Wells, Timothy J. Williams, Jinjun Xiong, Zhizhen Zhao
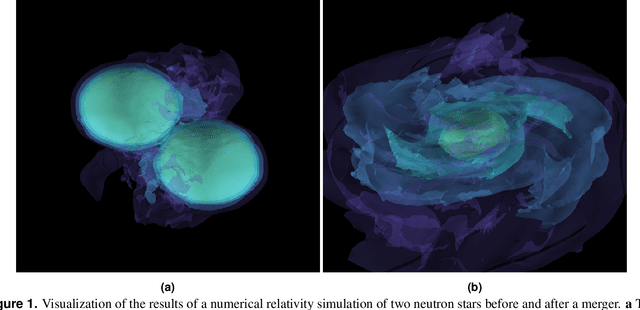
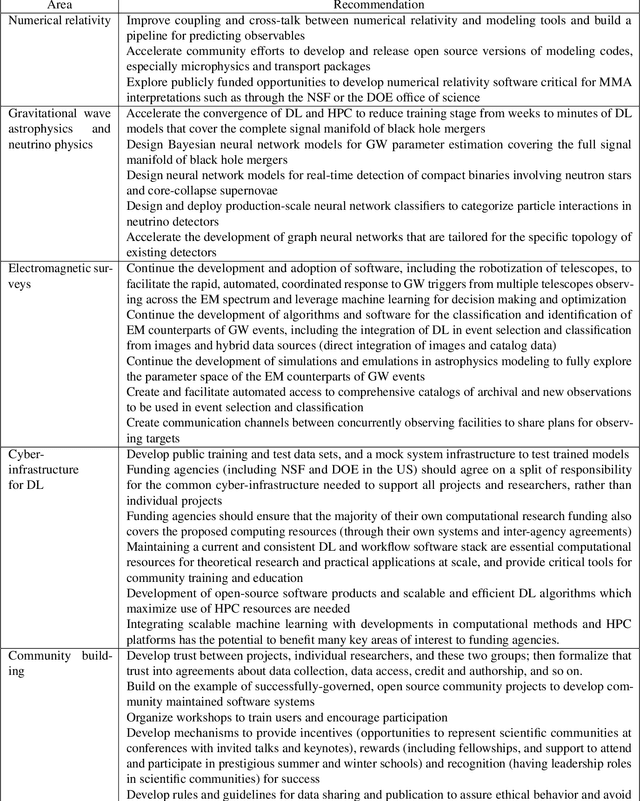
Multi-messenger astrophysics is a fast-growing, interdisciplinary field that combines data, which vary in volume and speed of data processing, from many different instruments that probe the Universe using different cosmic messengers: electromagnetic waves, cosmic rays, gravitational waves and neutrinos. In this Expert Recommendation, we review the key challenges of real-time observations of gravitational wave sources and their electromagnetic and astroparticle counterparts, and make a number of recommendations to maximize their potential for scientific discovery. These recommendations refer to the design of scalable and computationally efficient machine learning algorithms; the cyber-infrastructure to numerically simulate astrophysical sources, and to process and interpret multi-messenger astrophysics data; the management of gravitational wave detections to trigger real-time alerts for electromagnetic and astroparticle follow-ups; a vision to harness future developments of machine learning and cyber-infrastructure resources to cope with the big-data requirements; and the need to build a community of experts to realize the goals of multi-messenger astrophysics.
* Invited Expert Recommendation for Nature Reviews Physics. The art work produced by E. A. Huerta and Shawn Rosofsky for this article was used by Carl Conway to design the cover of the October 2019 issue of Nature Reviews Physics
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge