Kay Gregor Hartmann
EEG-GAN: Generative adversarial networks for electroencephalograhic (EEG) brain signals
Jun 05, 2018
Abstract:Generative adversarial networks (GANs) are recently highly successful in generative applications involving images and start being applied to time series data. Here we describe EEG-GAN as a framework to generate electroencephalographic (EEG) brain signals. We introduce a modification to the improved training of Wasserstein GANs to stabilize training and investigate a range of architectural choices critical for time series generation (most notably up- and down-sampling). For evaluation we consider and compare different metrics such as Inception score, Frechet inception distance and sliced Wasserstein distance, together showing that our EEG-GAN framework generated naturalistic EEG examples. It thus opens up a range of new generative application scenarios in the neuroscientific and neurological context, such as data augmentation in brain-computer interfacing tasks, EEG super-sampling, or restoration of corrupted data segments. The possibility to generate signals of a certain class and/or with specific properties may also open a new avenue for research into the underlying structure of brain signals.
Hierarchical internal representation of spectral features in deep convolutional networks trained for EEG decoding
Dec 15, 2017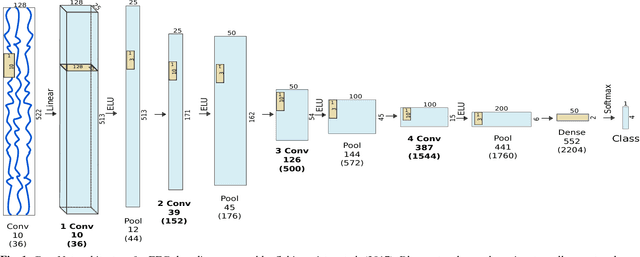
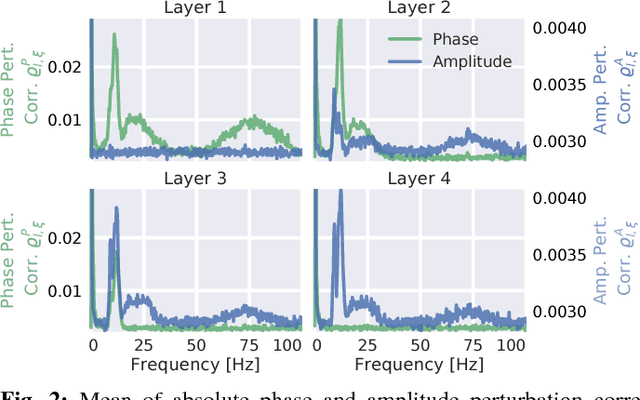
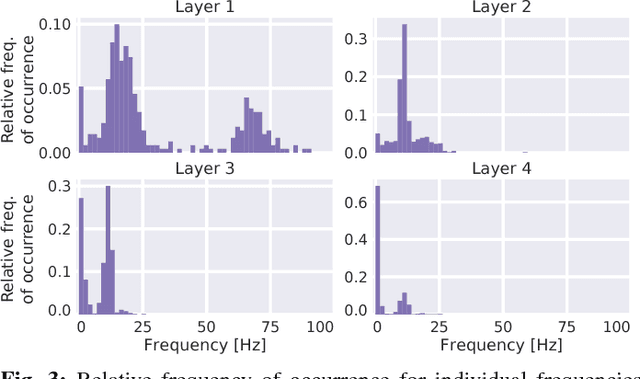
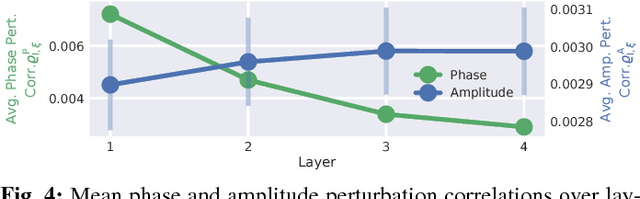
Abstract:Recently, there is increasing interest and research on the interpretability of machine learning models, for example how they transform and internally represent EEG signals in Brain-Computer Interface (BCI) applications. This can help to understand the limits of the model and how it may be improved, in addition to possibly provide insight about the data itself. Schirrmeister et al. (2017) have recently reported promising results for EEG decoding with deep convolutional neural networks (ConvNets) trained in an end-to-end manner and, with a causal visualization approach, showed that they learn to use spectral amplitude changes in the input. In this study, we investigate how ConvNets represent spectral features through the sequence of intermediate stages of the network. We show higher sensitivity to EEG phase features at earlier stages and higher sensitivity to EEG amplitude features at later stages. Intriguingly, we observed a specialization of individual stages of the network to the classical EEG frequency bands alpha, beta, and high gamma. Furthermore, we find first evidence that particularly in the last convolutional layer, the network learns to detect more complex oscillatory patterns beyond spectral phase and amplitude, reminiscent of the representation of complex visual features in later layers of ConvNets in computer vision tasks. Our findings thus provide insights into how ConvNets hierarchically represent spectral EEG features in their intermediate layers and suggest that ConvNets can exploit and might help to better understand the compositional structure of EEG time series.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge