Joseph Futoma
Scalable Modeling of Multivariate Longitudinal Data for Prediction of Chronic Kidney Disease Progression
Aug 16, 2016
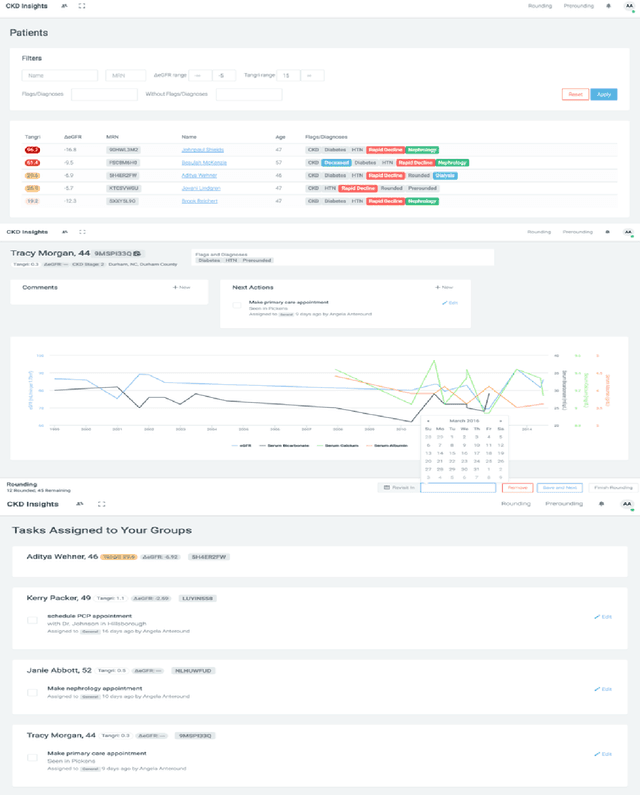
Abstract:Prediction of the future trajectory of a disease is an important challenge for personalized medicine and population health management. However, many complex chronic diseases exhibit large degrees of heterogeneity, and furthermore there is not always a single readily available biomarker to quantify disease severity. Even when such a clinical variable exists, there are often additional related biomarkers routinely measured for patients that may better inform the predictions of their future disease state. To this end, we propose a novel probabilistic generative model for multivariate longitudinal data that captures dependencies between multivariate trajectories. We use a Gaussian process based regression model for each individual trajectory, and build off ideas from latent class models to induce dependence between their mean functions. We fit our method using a scalable variational inference algorithm to a large dataset of longitudinal electronic patient health records, and find that it improves dynamic predictions compared to a recent state of the art method. Our local accountable care organization then uses the model predictions during chart reviews of high risk patients with chronic kidney disease.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge