Re-thinking Co-Salient Object Detection
Jul 11, 2020Deng-Ping Fan, Tengpeng Li, Zheng Lin, Ge-Peng Ji, Dingwen Zhang, Ming-Ming Cheng, Huazhu Fu, Jianbing Shen
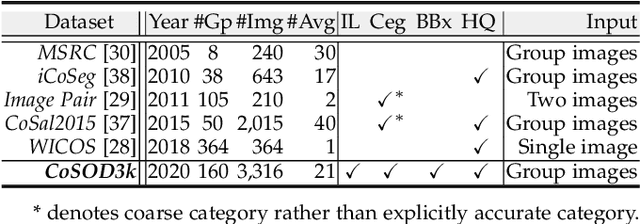


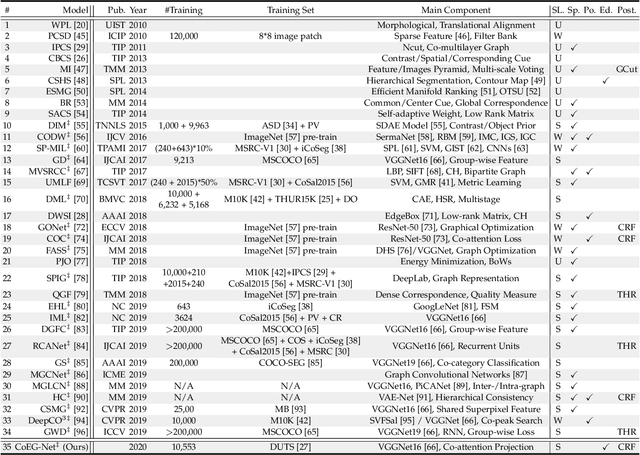

In this paper, we conduct a comprehensive study on the co-salient object detection (CoSOD) problem for images. CoSOD is an emerging and rapidly growing extension of salient object detection (SOD), which aims to detect the co-occurring salient objects in a group of images. However, existing CoSOD datasets often have a serious data bias, assuming that each group of images contains salient objects of similar visual appearances. This bias can lead to the ideal settings and effectiveness of models trained on existing datasets, being impaired in real-life situations, where similarities are usually semantic or conceptual. To tackle this issue, we first introduce a new benchmark, called CoSOD3k in the wild, which requires a large amount of semantic context, making it more challenging than existing CoSOD datasets. Our CoSOD3k consists of 3,316 high-quality, elaborately selected images divided into 160 groups with hierarchical annotations. The images span a wide range of categories, shapes, object sizes, and backgrounds. Second, we integrate the existing SOD techniques to build a unified, trainable CoSOD framework, which is long overdue in this field. Specifically, we propose a novel CoEG-Net that augments our prior model EGNet with a co-attention projection strategy to enable fast common information learning. CoEG-Net fully leverages previous large-scale SOD datasets and significantly improves the model scalability and stability. Third, we comprehensively summarize 34 cutting-edge algorithms, benchmarking 16 of them over three challenging CoSOD datasets (iCoSeg, CoSal2015, and our CoSOD3k), and reporting more detailed (i.e., group-level) performance analysis. Finally, we discuss the challenges and future works of CoSOD. We hope that our study will give a strong boost to growth in the CoSOD community
PraNet: Parallel Reverse Attention Network for Polyp Segmentation
Jul 03, 2020Deng-Ping Fan, Ge-Peng Ji, Tao Zhou, Geng Chen, Huazhu Fu, Jianbing Shen, Ling Shao
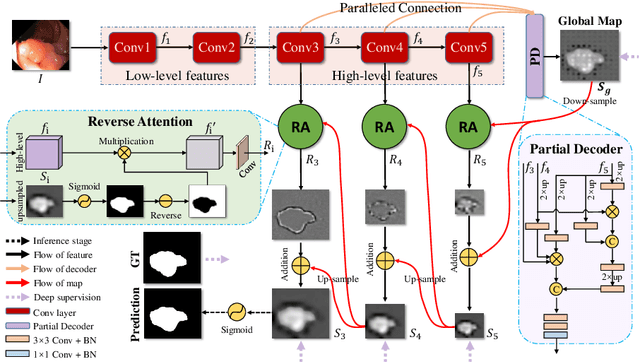
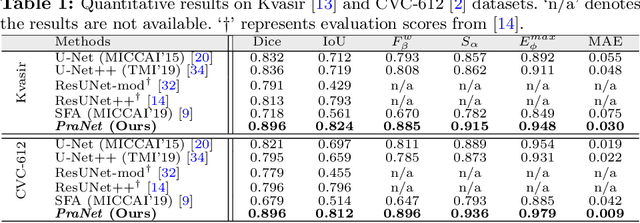
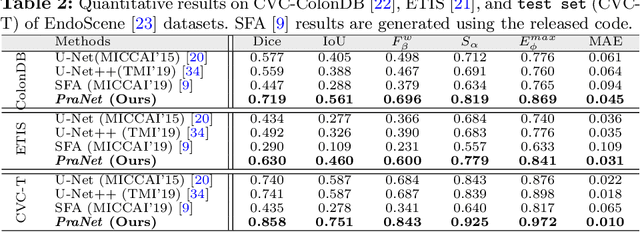

Colonoscopy is an effective technique for detecting colorectal polyps, which are highly related to colorectal cancer. In clinical practice, segmenting polyps from colonoscopy images is of great importance since it provides valuable information for diagnosis and surgery. However, accurate polyp segmentation is a challenging task, for two major reasons: (i) the same type of polyps has a diversity of size, color and texture; and (ii) the boundary between a polyp and its surrounding mucosa is not sharp. To address these challenges, we propose a parallel reverse attention network (PraNet) for accurate polyp segmentation in colonoscopy images. Specifically, we first aggregate the features in high-level layers using a parallel partial decoder (PPD). Based on the combined feature, we then generate a global map as the initial guidance area for the following components. In addition, we mine the boundary cues using a reverse attention (RA) module, which is able to establish the relationship between areas and boundary cues. Thanks to the recurrent cooperation mechanism between areas and boundaries, our PraNet is capable of calibrating any misaligned predictions, improving the segmentation accuracy. Quantitative and qualitative evaluations on five challenging datasets across six metrics show that our PraNet improves the segmentation accuracy significantly, and presents a number of advantages in terms of generalizability, and real-time segmentation efficiency.
M2Net: Multi-modal Multi-channel Network for Overall Survival Time Prediction of Brain Tumor Patients
Jun 01, 2020Tao Zhou, Huazhu Fu, Yu Zhang, Changqing Zhang, Xiankai Lu, Jianbing Shen, Ling Shao
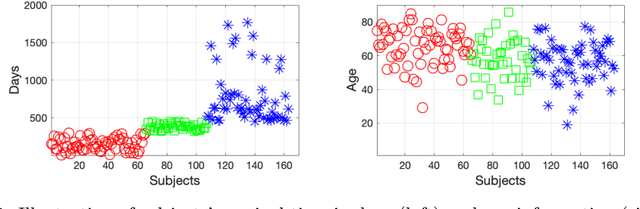
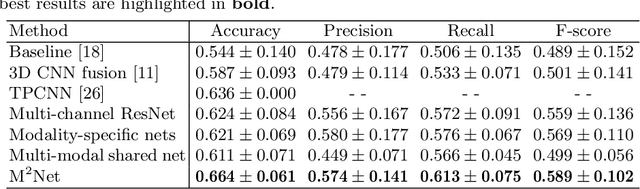
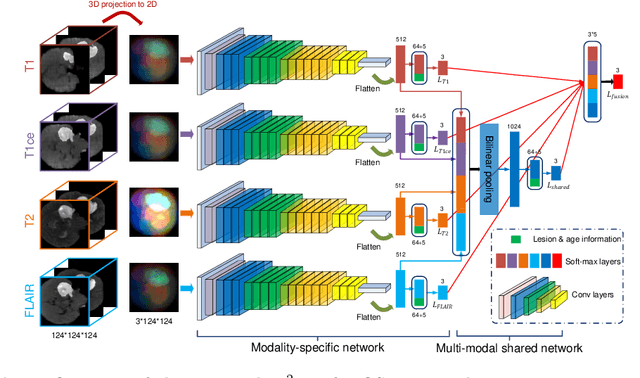
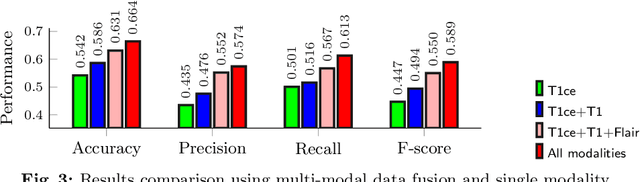
Early and accurate prediction of overall survival (OS) time can help to obtain better treatment planning for brain tumor patients. Although many OS time prediction methods have been developed and obtain promising results, there are still several issues. First, conventional prediction methods rely on radiomic features at the local lesion area of a magnetic resonance (MR) volume, which may not represent the full image or model complex tumor patterns. Second, different types of scanners (i.e., multi-modal data) are sensitive to different brain regions, which makes it challenging to effectively exploit the complementary information across multiple modalities and also preserve the modality-specific properties. Third, existing methods focus on prediction models, ignoring complex data-to-label relationships. To address the above issues, we propose an end-to-end OS time prediction model; namely, Multi-modal Multi-channel Network (M2Net). Specifically, we first project the 3D MR volume onto 2D images in different directions, which reduces computational costs, while preserving important information and enabling pre-trained models to be transferred from other tasks. Then, we use a modality-specific network to extract implicit and high-level features from different MR scans. A multi-modal shared network is built to fuse these features using a bilinear pooling model, exploiting their correlations to provide complementary information. Finally, we integrate the outputs from each modality-specific network and the multi-modal shared network to generate the final prediction result. Experimental results demonstrate the superiority of our M2Net model over other methods.
Inf-Net: Automatic COVID-19 Lung Infection Segmentation from CT Images
May 21, 2020Deng-Ping Fan, Tao Zhou, Ge-Peng Ji, Yi Zhou, Geng Chen, Huazhu Fu, Jianbing Shen, Ling Shao
Coronavirus Disease 2019 (COVID-19) spread globally in early 2020, causing the world to face an existential health crisis. Automated detection of lung infections from computed tomography (CT) images offers a great potential to augment the traditional healthcare strategy for tackling COVID-19. However, segmenting infected regions from CT slices faces several challenges, including high variation in infection characteristics, and low intensity contrast between infections and normal tissues. Further, collecting a large amount of data is impractical within a short time period, inhibiting the training of a deep model. To address these challenges, a novel COVID-19 Lung Infection Segmentation Deep Network (Inf-Net) is proposed to automatically identify infected regions from chest CT slices. In our Inf-Net, a parallel partial decoder is used to aggregate the high-level features and generate a global map. Then, the implicit reverse attention and explicit edge-attention are utilized to model the boundaries and enhance the representations. Moreover, to alleviate the shortage of labeled data, we present a semi-supervised segmentation framework based on a randomly selected propagation strategy, which only requires a few labeled images and leverages primarily unlabeled data. Our semi-supervised framework can improve the learning ability and achieve a higher performance. Extensive experiments on our COVID-SemiSeg and real CT volumes demonstrate that the proposed Inf-Net outperforms most cutting-edge segmentation models and advances the state-of-the-art performance.
Understanding and Correcting Low-quality Retinal Fundus Images for Clinical Analysis
May 12, 2020Ziyi Shen, Huazhu Fu, Jianbing Shen, Ling Shao
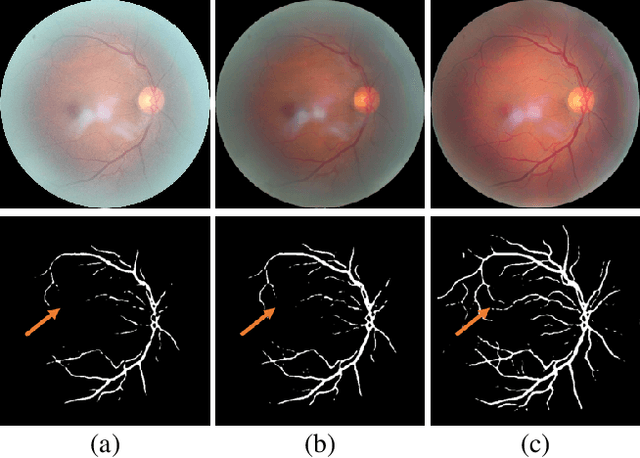
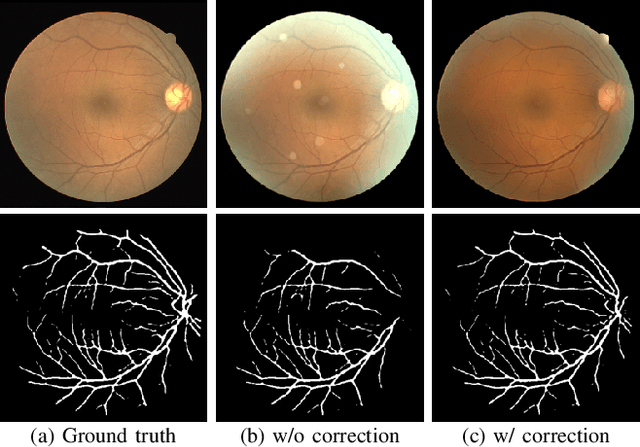
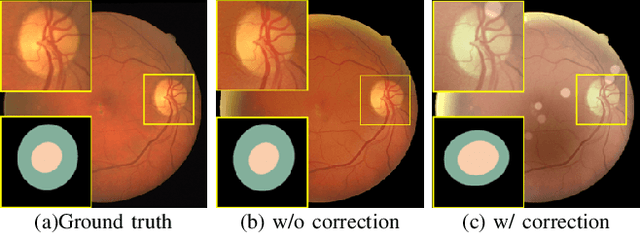
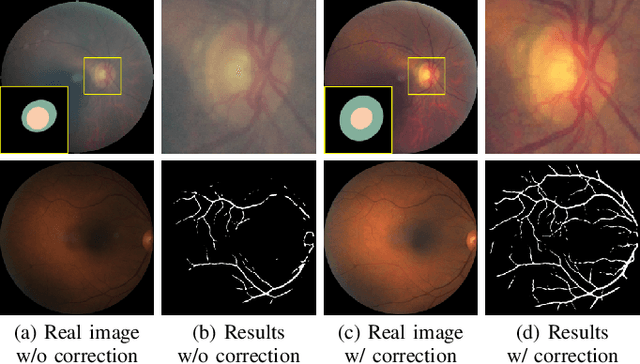
Retinal fundus images are widely used for clinical screening and diagnosis of eye diseases. However, fundus images captured by operators with various levels of experiences have a large variation in quality. Low-quality fundus images increase the uncertainty in clinical observation and lead to a risk of misdiagnosis. Due to the special optical beam of fundus imaging and retinal structure, the natural image enhancement methods cannot be utilized directly. In this paper, we first analyze the ophthalmoscope imaging system and model the reliable degradation of major inferior-quality factors, including uneven illumination, blur, and artifacts. Then, based on the degradation model, a clinical-oriented fundus enhancement network~(cofe-Net)~is proposed to suppress the global degradation factors, and simultaneously preserve anatomical retinal structures and pathological characteristics for clinical observation and analysis. Experiments on both synthetic and real fundus images demonstrate that our algorithm effectively corrects low-quality fundus images without losing retinal details. Moreover, we also show that the fundus correction method can benefit medical image analysis applications, e.g, retinal vessel segmentation and optic disc/cup detection.
Inf-Net: Automatic COVID-19 Lung Infection Segmentation from CT Scans
May 01, 2020Deng-Ping Fan, Tao Zhou, Ge-Peng Ji, Yi Zhou, Geng Chen, Huazhu Fu, Jianbing Shen, Ling Shao
Coronavirus Disease 2019 (COVID-19) spread globally in early 2020, causing the world to face an existential health crisis. Automated detection of lung infections from computed tomography (CT) images offers a great potential to augment the traditional healthcare strategy for tackling COVID-19. However, segmenting infected regions from CT scans faces several challenges, including high variation in infection characteristics, and low intensity contrast between infections and normal tissues. Further, collecting a large amount of data is impractical within a short time period, inhibiting the training of a deep model. To address these challenges, a novel COVID-19 Lung Infection Segmentation Deep Network (Inf-Net) is proposed to automatically identify infected regions from chest CT scans. In our Inf-Net, a parallel partial decoder is used to aggregate the high-level features and generate a global map. Then, the implicit reverse attention and explicit edge-attention are utilized to model the boundaries and enhance the representations. Moreover, to alleviate the shortage of labeled data, we present a semi-supervised segmentation framework based on a randomly selected propagation strategy, which only requires a few labeled images and leverages primarily unlabeled data. Our semi-supervised framework can improve the learning ability and achieve a higher performance. Extensive experiments on our COVID-SemiSeg and and real CT volumes demonstrate that the proposed Inf-Net outperforms most cutting-edge segmentation models and advances the state-of-the-art performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge