Jared Dunnmon
Predicting Food Security Outcomes Using Convolutional Neural Networks (CNNs) for Satellite Tasking
Feb 13, 2019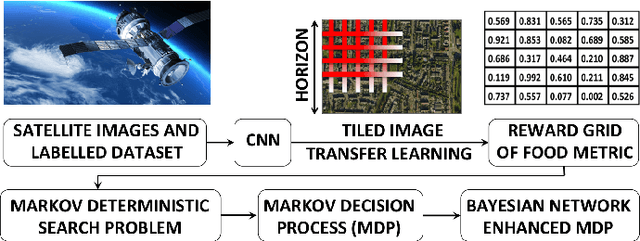
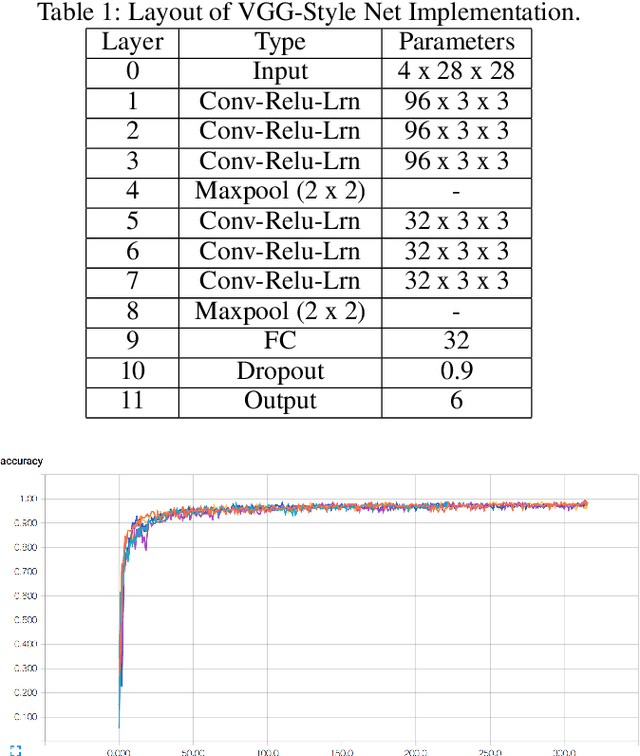
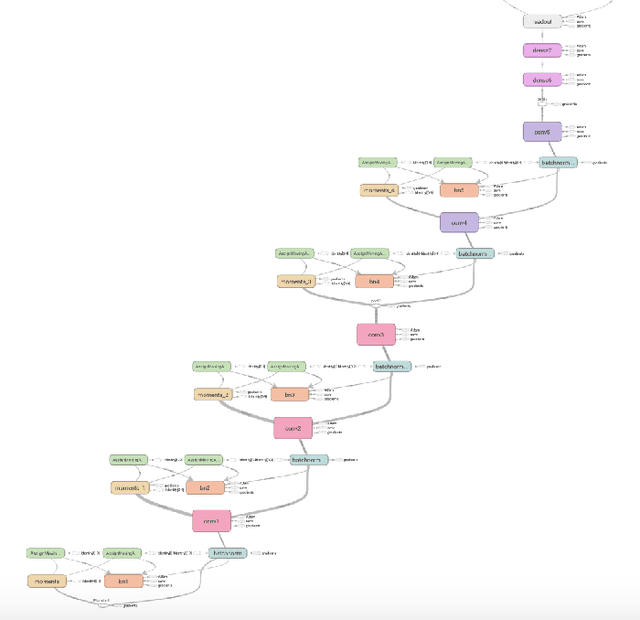
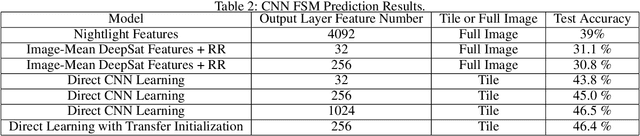
Abstract:Obtaining reliable data describing local Food Security Metrics (FSM) at a granularity that is informative to policy-makers requires expensive and logistically difficult surveys, particularly in the developing world. We train a CNN on publicly available satellite data describing land cover classification and use both transfer learning and direct training to build a model for FSM prediction purely from satellite imagery data. We then propose efficient tasking algorithms for high resolution satellite assets via transfer learning, Markovian search algorithms, and Bayesian networks.
Learning to Compose Domain-Specific Transformations for Data Augmentation
Sep 30, 2017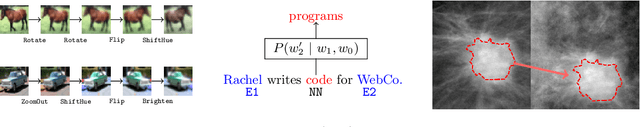
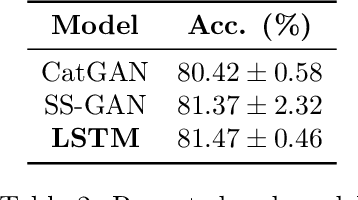
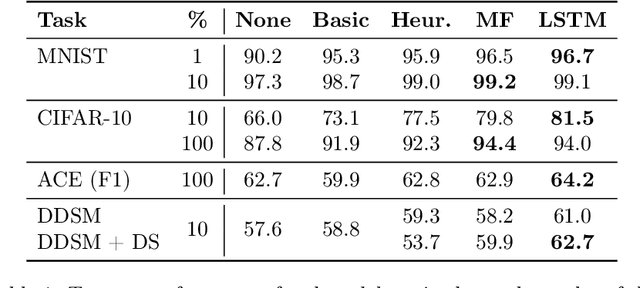
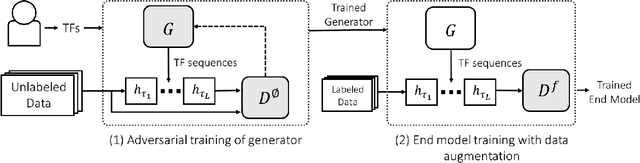

Abstract:Data augmentation is a ubiquitous technique for increasing the size of labeled training sets by leveraging task-specific data transformations that preserve class labels. While it is often easy for domain experts to specify individual transformations, constructing and tuning the more sophisticated compositions typically needed to achieve state-of-the-art results is a time-consuming manual task in practice. We propose a method for automating this process by learning a generative sequence model over user-specified transformation functions using a generative adversarial approach. Our method can make use of arbitrary, non-deterministic transformation functions, is robust to misspecified user input, and is trained on unlabeled data. The learned transformation model can then be used to perform data augmentation for any end discriminative model. In our experiments, we show the efficacy of our approach on both image and text datasets, achieving improvements of 4.0 accuracy points on CIFAR-10, 1.4 F1 points on the ACE relation extraction task, and 3.4 accuracy points when using domain-specific transformation operations on a medical imaging dataset as compared to standard heuristic augmentation approaches.
Optimizing and Visualizing Deep Learning for Benign/Malignant Classification in Breast Tumors
May 17, 2017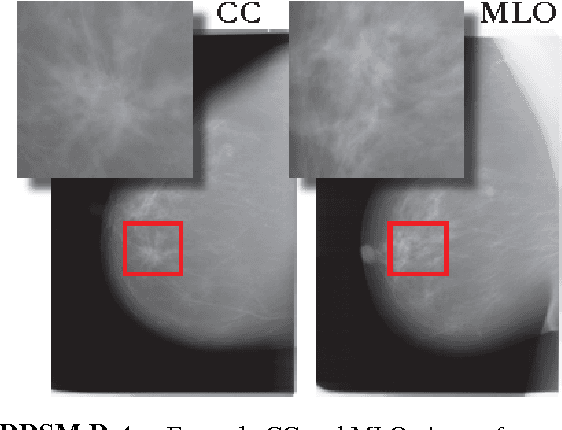
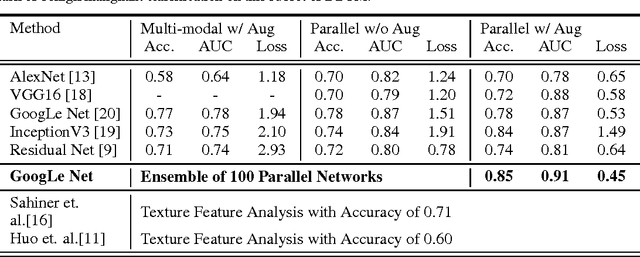
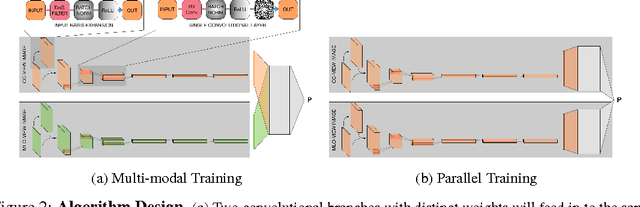
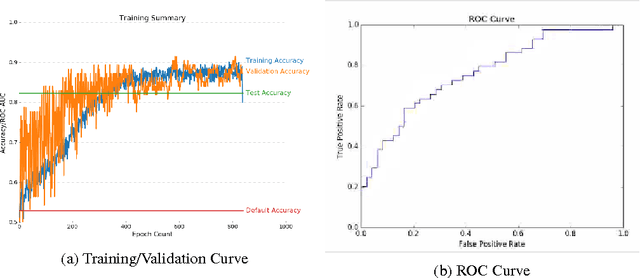
Abstract:Breast cancer has the highest incidence and second highest mortality rate for women in the US. Our study aims to utilize deep learning for benign/malignant classification of mammogram tumors using a subset of cases from the Digital Database of Screening Mammography (DDSM). Though it was a small dataset from the view of Deep Learning (about 1000 patients), we show that currently state of the art architectures of deep learning can find a robust signal, even when trained from scratch. Using convolutional neural networks (CNNs), we are able to achieve an accuracy of 85% and an ROC AUC of 0.91, while leading hand-crafted feature based methods are only able to achieve an accuracy of 71%. We investigate an amalgamation of architectures to show that our best result is reached with an ensemble of the lightweight GoogLe Nets tasked with interpreting both the coronal caudal view and the mediolateral oblique view, simply averaging the probability scores of both views to make the final prediction. In addition, we have created a novel method to visualize what features the neural network detects for the benign/malignant classification, and have correlated those features with well known radiological features, such as spiculation. Our algorithm significantly improves existing classification methods for mammography lesions and identifies features that correlate with established clinical markers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge