Jaime Roquero Gimenez
A Unified f-divergence Framework Generalizing VAE and GAN
May 11, 2022
Abstract:Developing deep generative models that flexibly incorporate diverse measures of probability distance is an important area of research. Here we develop an unified mathematical framework of f-divergence generative model, f-GM, that incorporates both VAE and f-GAN, and enables tractable learning with general f-divergences. f-GM allows the experimenter to flexibly design the f-divergence function without changing the structure of the networks or the learning procedure. f-GM jointly models three components: a generator, a inference network and a density estimator. Therefore it simultaneously enables sampling, posterior inference of the latent variable as well as evaluation of the likelihood of an arbitrary datum. f-GM belongs to the class of encoder-decoder GANs: our density estimator can be interpreted as playing the role of a discriminator between samples in the joint space of latent code and observed space. We prove that f-GM naturally simplifies to the standard VAE and to f-GAN as special cases, and illustrates the connections between different encoder-decoder GAN architectures. f-GM is compatible with general network architecture and optimizer. We leverage it to experimentally explore the effects -- e.g. mode collapse and image sharpness -- of different choices of f-divergence.
Discovering Conditionally Salient Features with Statistical Guarantees
May 29, 2019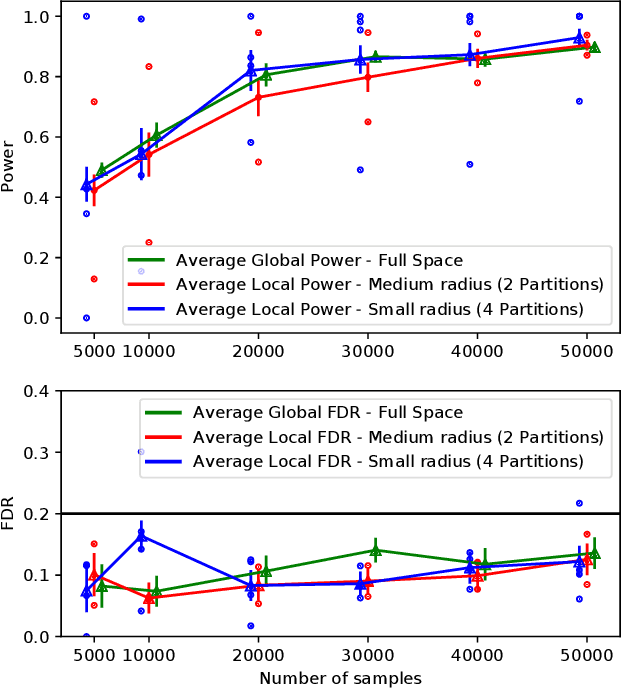
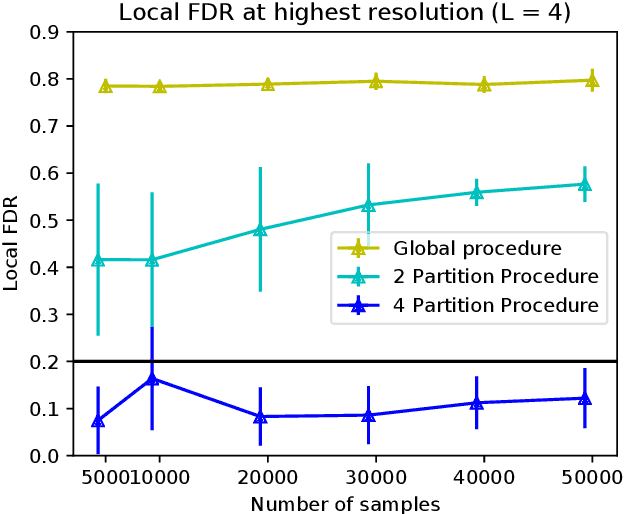
Abstract:The goal of feature selection is to identify important features that are relevant to explain an outcome variable. Most of the work in this domain has focused on identifying globally relevant features, which are features that are related to the outcome using evidence across the entire dataset. We study a more fine-grained statistical problem: conditional feature selection, where a feature may be relevant depending on the values of the other features. For example in genetic association studies, variant $A$ could be associated with the phenotype in the entire dataset, but conditioned on variant $B$ being present it might be independent of the phenotype. In this sense, variant $A$ is globally relevant, but conditioned on $B$ it is no longer locally relevant in that region of the feature space. We present a generalization of the knockoff procedure that performs conditional feature selection while controlling a generalization of the false discovery rate (FDR) to the conditional setting. By exploiting the feature/response model-free framework of the knockoffs, the quality of the statistical FDR guarantee is not degraded even when we perform conditional feature selections. We implement this method and present an algorithm that automatically partitions the feature space such that it enhances the differences between selected sets in different regions, and validate the statistical theoretical results with experiments.
Improving the Stability of the Knockoff Procedure: Multiple Simultaneous Knockoffs and Entropy Maximization
Oct 26, 2018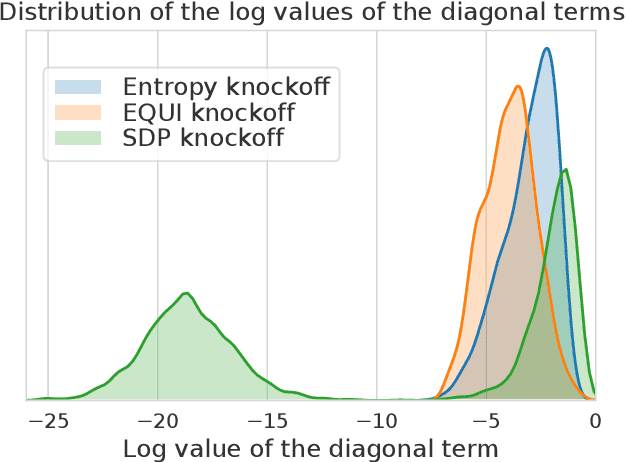
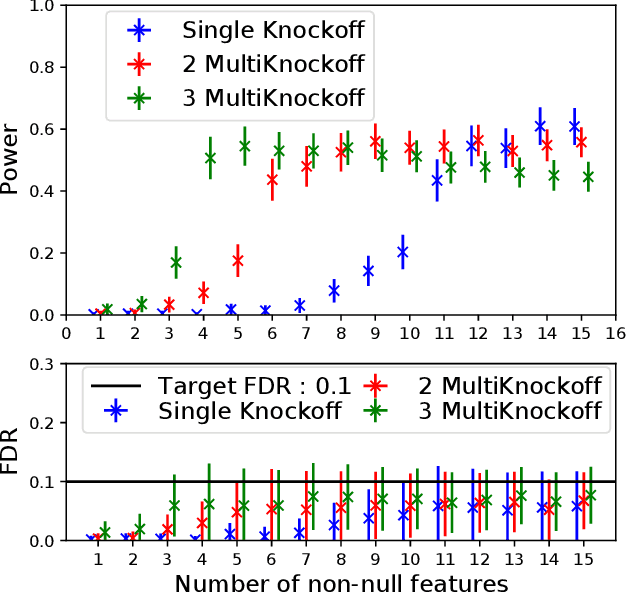
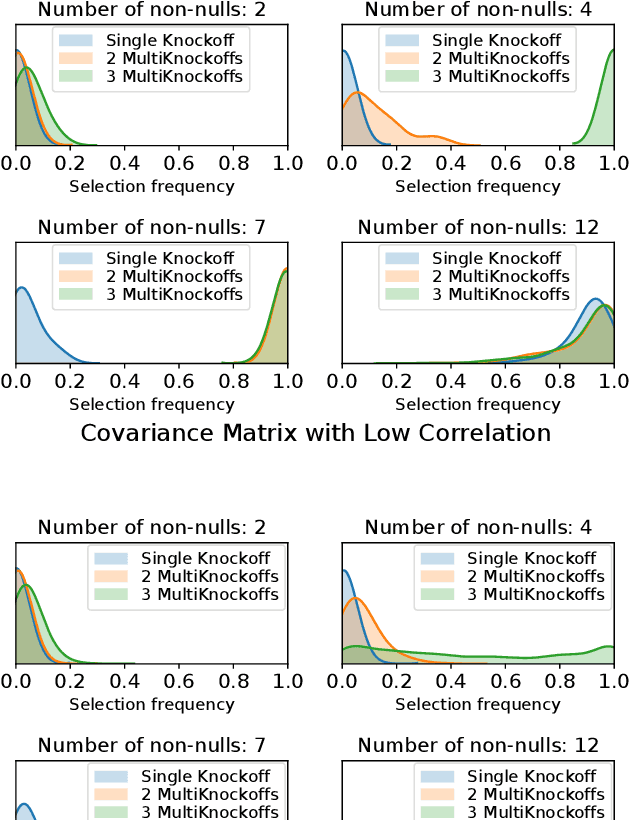
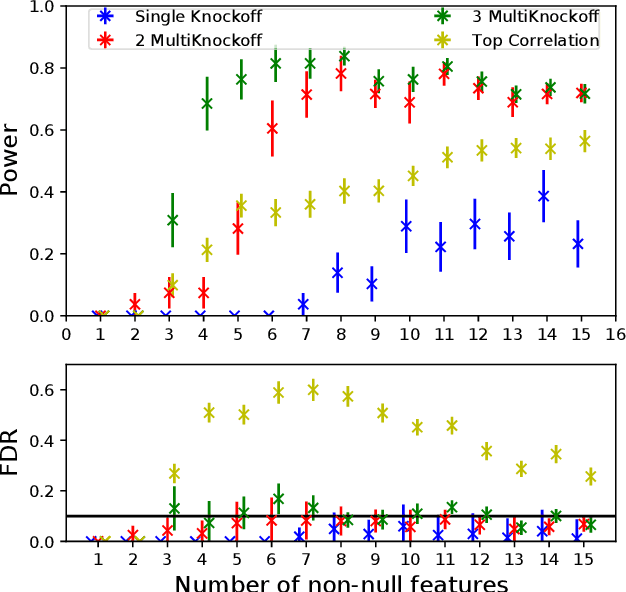
Abstract:The Model-X knockoff procedure has recently emerged as a powerful approach for feature selection with statistical guarantees. The advantage of knockoff is that if we have a good model of the features X, then we can identify salient features without knowing anything about how the outcome Y depends on X. An important drawback of knockoffs is its instability: running the procedure twice can result in very different selected features, potentially leading to different conclusions. Addressing this instability is critical for obtaining reproducible and robust results. Here we present a generalization of the knockoff procedure that we call simultaneous multi-knockoffs. We show that multi-knockoff guarantees false discovery rate (FDR) control, and is substantially more stable and powerful compared to the standard (single) knockoff. Moreover we propose a new algorithm based on entropy maximization for generating Gaussian multi-knockoffs. We validate the improved stability and power of multi-knockoffs in systematic experiments. We also illustrate how multi-knockoffs can improve the accuracy of detecting genetic mutations that are causally linked to phenotypes.
Knockoffs for the mass: new feature importance statistics with false discovery guarantees
Jul 17, 2018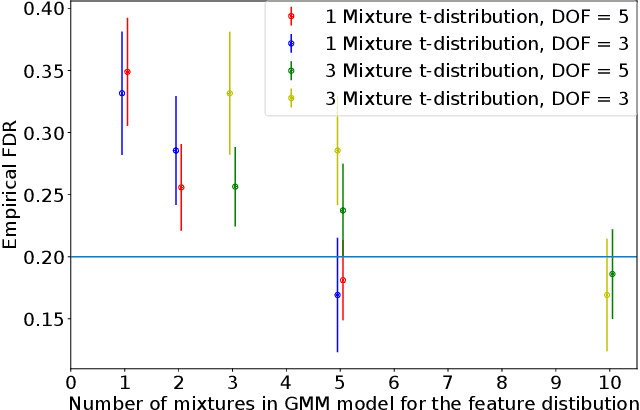
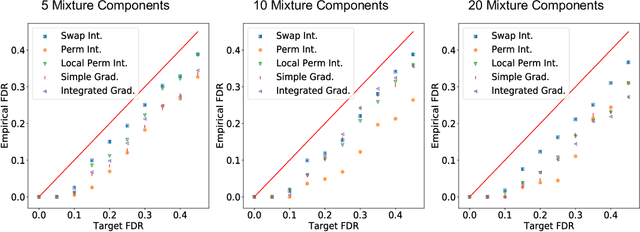
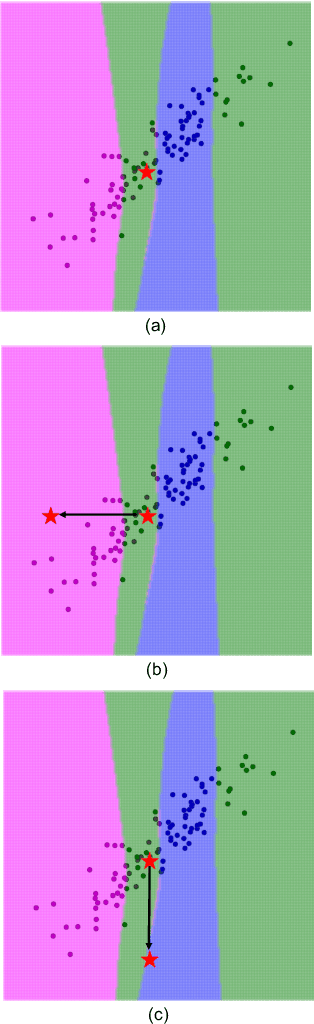
Abstract:An important problem in machine learning and statistics is to identify features that causally affect the outcome. This is often impossible to do from purely observational data, and a natural relaxation is to identify features that are correlated with the outcome even conditioned on all other observed features. For example, we want to identify that smoking really is correlated with cancer conditioned on demographics. The knockoff procedure is a recent breakthrough in statistics that, in theory, can identify truly correlated features while guaranteeing that the false discovery is limited. The idea is to create synthetic data -knockoffs- that captures correlations amongst the features. However there are substantial computational and practical challenges to generating and using knockoffs. This paper makes several key advances that enable knockoff application to be more efficient and powerful. We develop an efficient algorithm to generate valid knockoffs from Bayesian Networks. Then we systematically evaluate knockoff test statistics and develop new statistics with improved power. The paper combines new mathematical guarantees with systematic experiments on real and synthetic data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge