Huiyi Hu
An Incremental Reseeding Strategy for Clustering
Jun 15, 2014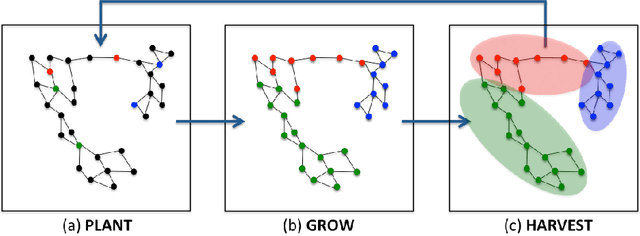
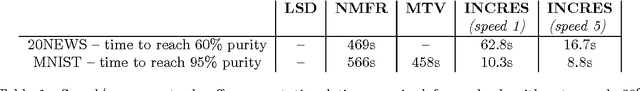
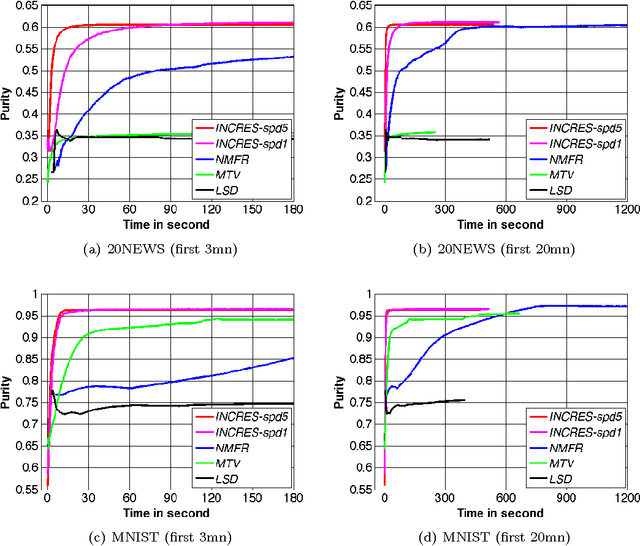
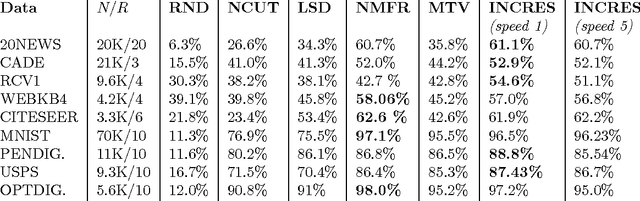
Abstract:In this work we propose a simple and easily parallelizable algorithm for multiway graph partitioning. The algorithm alternates between three basic components: diffusing seed vertices over the graph, thresholding the diffused seeds, and then randomly reseeding the thresholded clusters. We demonstrate experimentally that the proper combination of these ingredients leads to an algorithm that achieves state-of-the-art performance in terms of cluster purity on standard benchmarks datasets. Moreover, the algorithm runs an order of magnitude faster than the other algorithms that achieve comparable results in terms of accuracy. We also describe a coarsen, cluster and refine approach similar to GRACLUS and METIS that removes an additional order of magnitude from the runtime of our algorithm while still maintaining competitive accuracy.
Multislice Modularity Optimization in Community Detection and Image Segmentation
Nov 30, 2012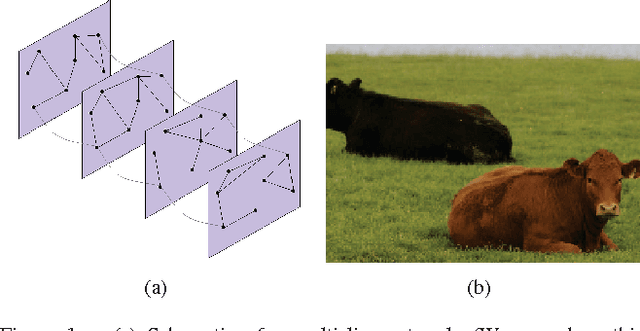
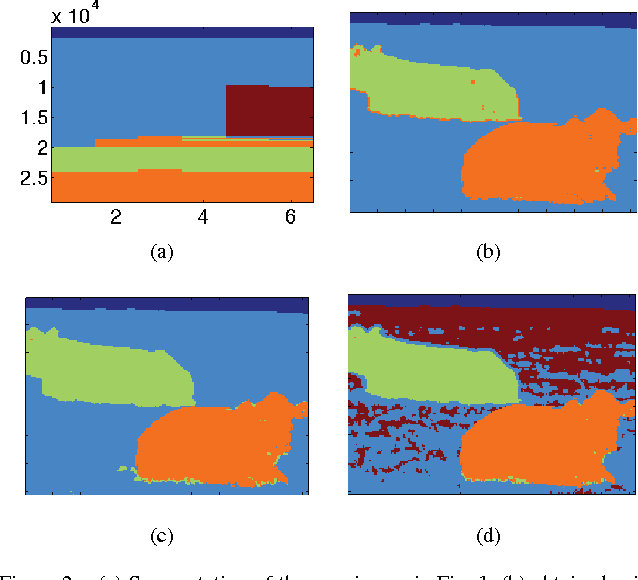
Abstract:Because networks can be used to represent many complex systems, they have attracted considerable attention in physics, computer science, sociology, and many other disciplines. One of the most important areas of network science is the algorithmic detection of cohesive groups (i.e., "communities") of nodes. In this paper, we algorithmically detect communities in social networks and image data by optimizing multislice modularity. A key advantage of modularity optimization is that it does not require prior knowledge of the number or sizes of communities, and it is capable of finding network partitions that are composed of communities of different sizes. By optimizing multislice modularity and subsequently calculating diagnostics on the resulting network partitions, it is thereby possible to obtain information about network structure across multiple system scales. We illustrate this method on data from both social networks and images, and we find that optimization of multislice modularity performs well on these two tasks without the need for extensive problem-specific adaptation. However, improving the computational speed of this method remains a challenging open problem.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge