Dirk Weissenborn
Neural Question Answering at BioASQ 5B
Jun 26, 2017
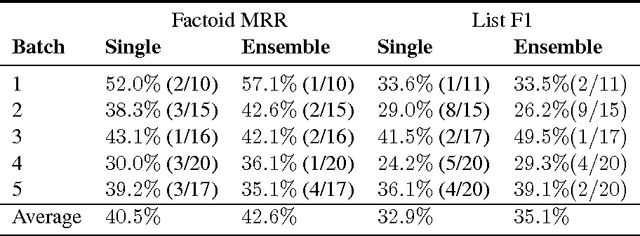
Abstract:This paper describes our submission to the 2017 BioASQ challenge. We participated in Task B, Phase B which is concerned with biomedical question answering (QA). We focus on factoid and list question, using an extractive QA model, that is, we restrict our system to output substrings of the provided text snippets. At the core of our system, we use FastQA, a state-of-the-art neural QA system. We extended it with biomedical word embeddings and changed its answer layer to be able to answer list questions in addition to factoid questions. We pre-trained the model on a large-scale open-domain QA dataset, SQuAD, and then fine-tuned the parameters on the BioASQ training set. With our approach, we achieve state-of-the-art results on factoid questions and competitive results on list questions.
Neural Domain Adaptation for Biomedical Question Answering
Jun 15, 2017
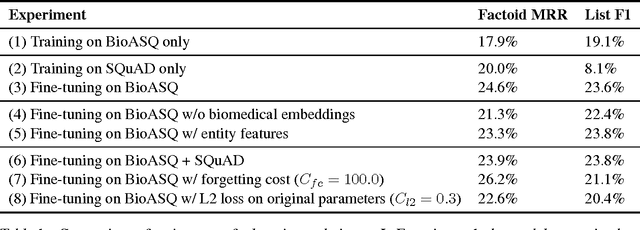

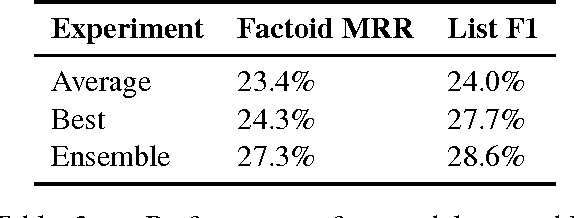
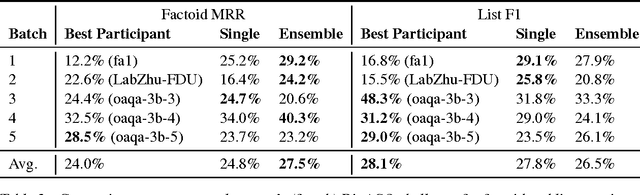
Abstract:Factoid question answering (QA) has recently benefited from the development of deep learning (DL) systems. Neural network models outperform traditional approaches in domains where large datasets exist, such as SQuAD (ca. 100,000 questions) for Wikipedia articles. However, these systems have not yet been applied to QA in more specific domains, such as biomedicine, because datasets are generally too small to train a DL system from scratch. For example, the BioASQ dataset for biomedical QA comprises less then 900 factoid (single answer) and list (multiple answers) QA instances. In this work, we adapt a neural QA system trained on a large open-domain dataset (SQuAD, source) to a biomedical dataset (BioASQ, target) by employing various transfer learning techniques. Our network architecture is based on a state-of-the-art QA system, extended with biomedical word embeddings and a novel mechanism to answer list questions. In contrast to existing biomedical QA systems, our system does not rely on domain-specific ontologies, parsers or entity taggers, which are expensive to create. Despite this fact, our systems achieve state-of-the-art results on factoid questions and competitive results on list questions.
Making Neural QA as Simple as Possible but not Simpler
Jun 08, 2017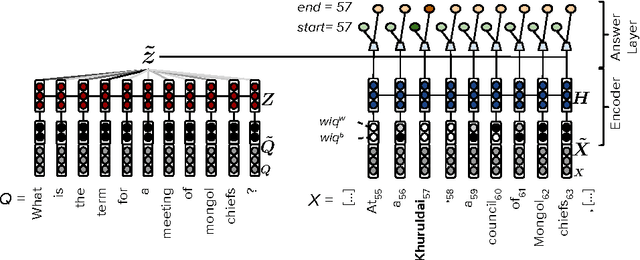
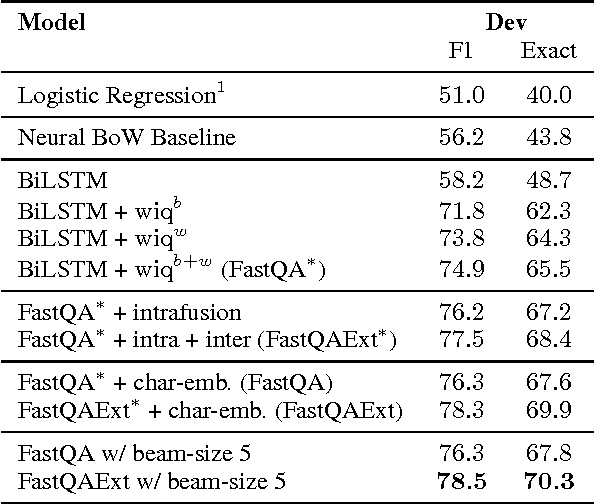
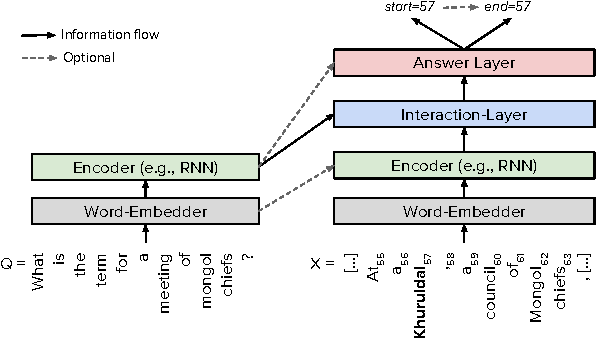
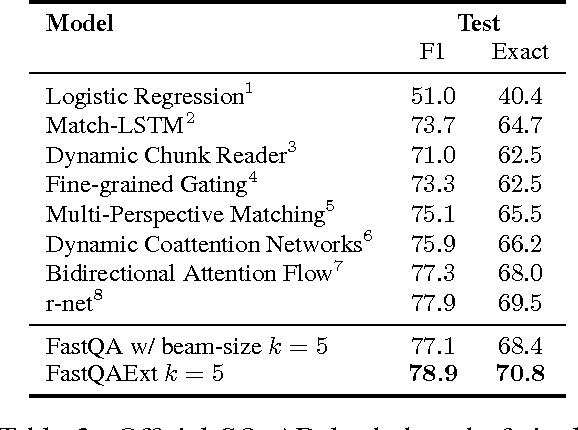
Abstract:Recent development of large-scale question answering (QA) datasets triggered a substantial amount of research into end-to-end neural architectures for QA. Increasingly complex systems have been conceived without comparison to simpler neural baseline systems that would justify their complexity. In this work, we propose a simple heuristic that guides the development of neural baseline systems for the extractive QA task. We find that there are two ingredients necessary for building a high-performing neural QA system: first, the awareness of question words while processing the context and second, a composition function that goes beyond simple bag-of-words modeling, such as recurrent neural networks. Our results show that FastQA, a system that meets these two requirements, can achieve very competitive performance compared with existing models. We argue that this surprising finding puts results of previous systems and the complexity of recent QA datasets into perspective.
Separating Answers from Queries for Neural Reading Comprehension
Sep 27, 2016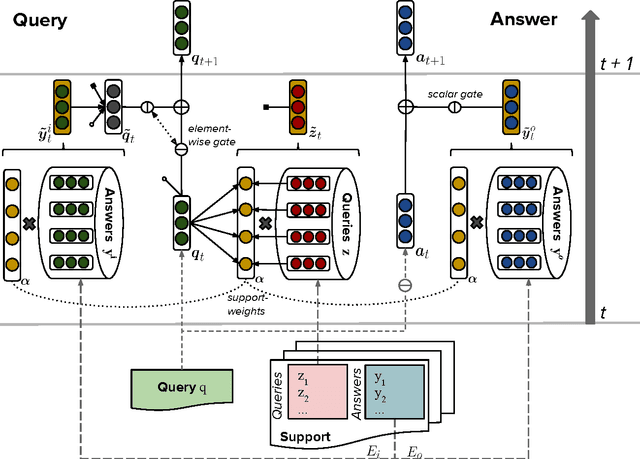
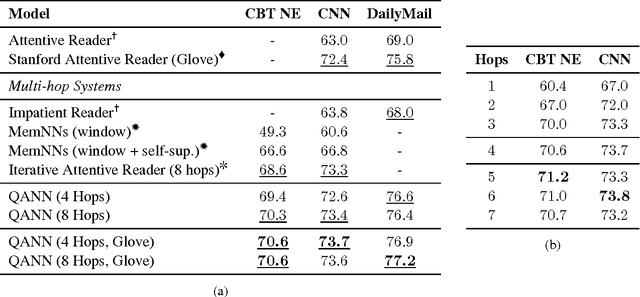
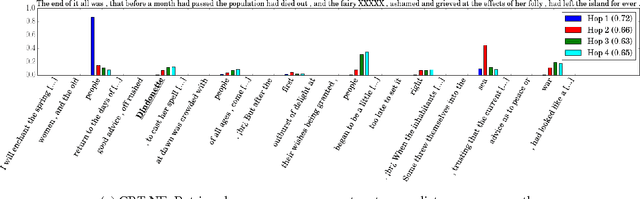
Abstract:We present a novel neural architecture for answering queries, designed to optimally leverage explicit support in the form of query-answer memories. Our model is able to refine and update a given query while separately accumulating evidence for predicting the answer. Its architecture reflects this separation with dedicated embedding matrices and loosely connected information pathways (modules) for updating the query and accumulating evidence. This separation of responsibilities effectively decouples the search for query related support and the prediction of the answer. On recent benchmark datasets for reading comprehension, our model achieves state-of-the-art results. A qualitative analysis reveals that the model effectively accumulates weighted evidence from the query and over multiple support retrieval cycles which results in a robust answer prediction.
SynsetRank: Degree-adjusted Random Walk for Relation Identification
Sep 15, 2016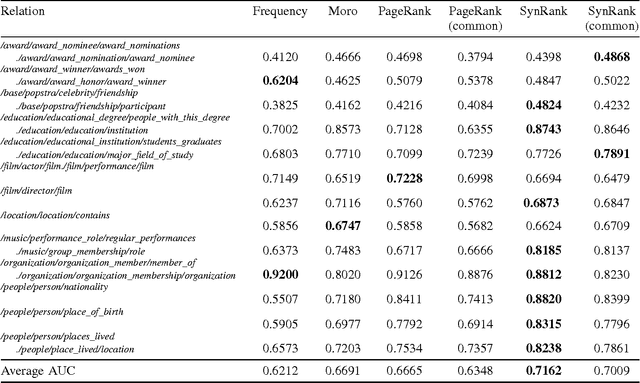

Abstract:In relation extraction, a key process is to obtain good detectors that find relevant sentences describing the target relation. To minimize the necessity of labeled data for refining detectors, previous work successfully made use of BabelNet, a semantic graph structure expressing relationships between synsets, as side information or prior knowledge. The goal of this paper is to enhance the use of graph structure in the framework of random walk with a few adjustable parameters. Actually, a straightforward application of random walk degrades the performance even after parameter optimization. With the insight from this unsuccessful trial, we propose SynsetRank, which adjusts the initial probability so that high degree nodes influence the neighbors as strong as low degree nodes. In our experiment on 13 relations in the FB15K-237 dataset, SynsetRank significantly outperforms baselines and the plain random walk approach.
Neural Associative Memory for Dual-Sequence Modeling
Jun 14, 2016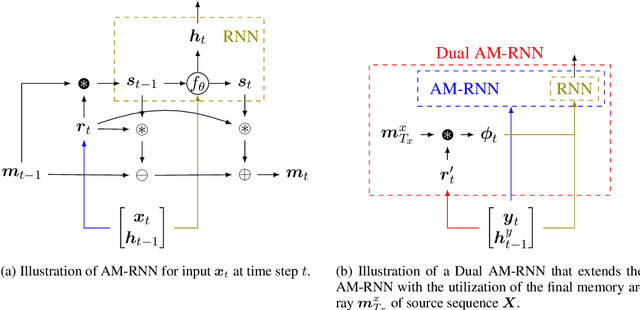
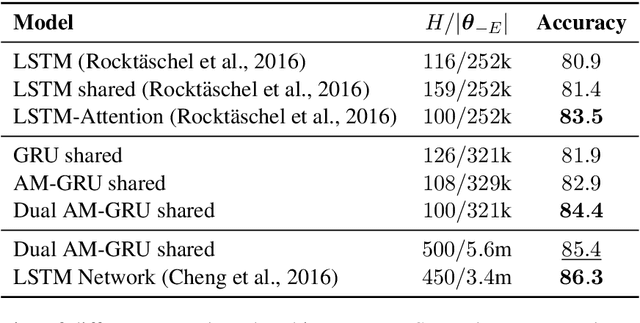
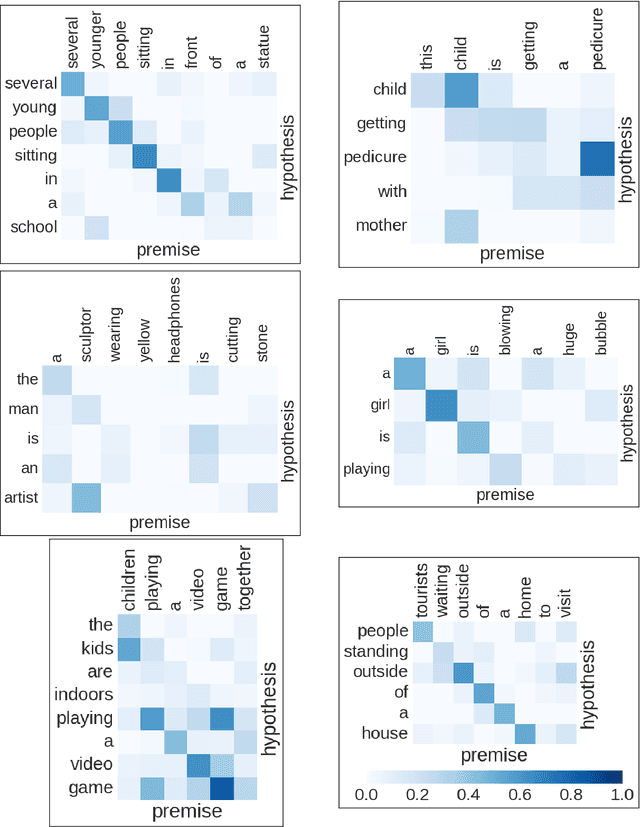
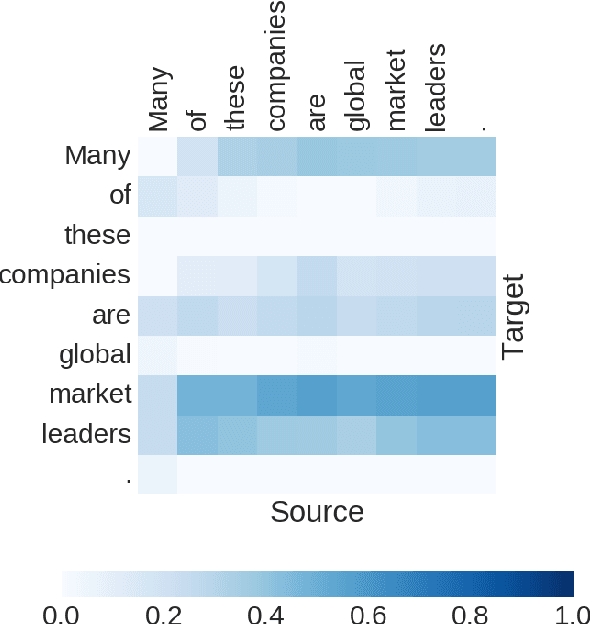
Abstract:Many important NLP problems can be posed as dual-sequence or sequence-to-sequence modeling tasks. Recent advances in building end-to-end neural architectures have been highly successful in solving such tasks. In this work we propose a new architecture for dual-sequence modeling that is based on associative memory. We derive AM-RNNs, a recurrent associative memory (AM) which augments generic recurrent neural networks (RNN). This architecture is extended to the Dual AM-RNN which operates on two AMs at once. Our models achieve very competitive results on textual entailment. A qualitative analysis demonstrates that long range dependencies between source and target-sequence can be bridged effectively using Dual AM-RNNs. However, an initial experiment on auto-encoding reveals that these benefits are not exploited by the system when learning to solve sequence-to-sequence tasks which indicates that additional supervision or regularization is needed.
MuFuRU: The Multi-Function Recurrent Unit
Jun 09, 2016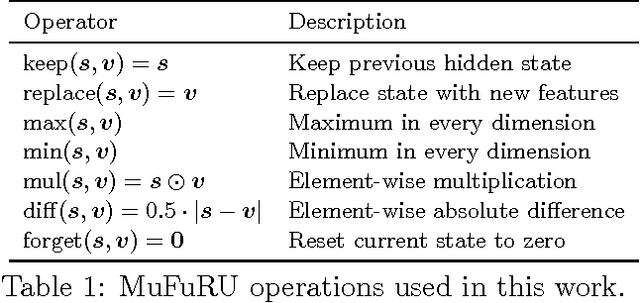
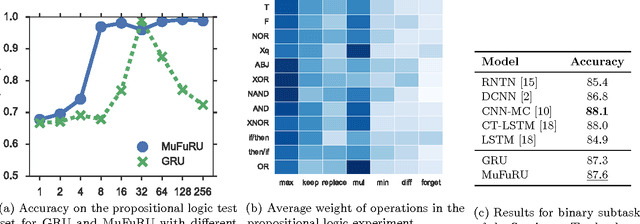
Abstract:Recurrent neural networks such as the GRU and LSTM found wide adoption in natural language processing and achieve state-of-the-art results for many tasks. These models are characterized by a memory state that can be written to and read from by applying gated composition operations to the current input and the previous state. However, they only cover a small subset of potentially useful compositions. We propose Multi-Function Recurrent Units (MuFuRUs) that allow for arbitrary differentiable functions as composition operations. Furthermore, MuFuRUs allow for an input- and state-dependent choice of these composition operations that is learned. Our experiments demonstrate that the additional functionality helps in different sequence modeling tasks, including the evaluation of propositional logic formulae, language modeling and sentiment analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge