Denis Newman-Griffis
Classifying the reported ability in clinical mobility descriptions
Jun 07, 2019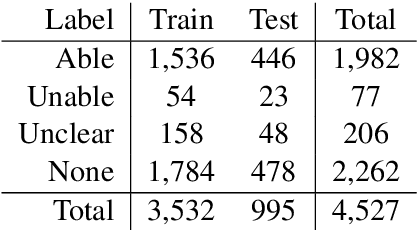
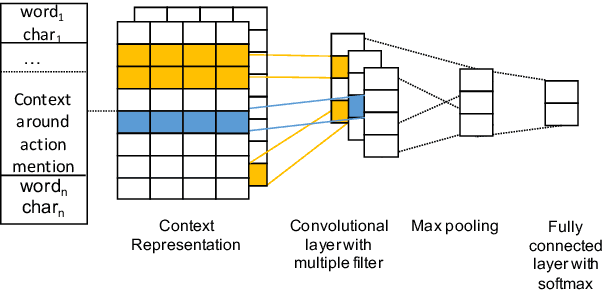
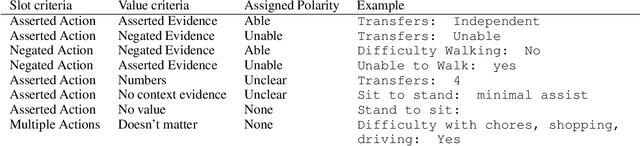
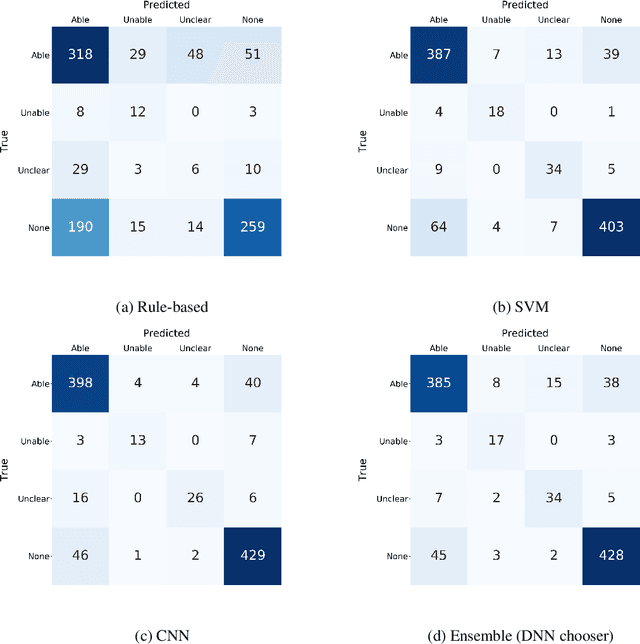
Abstract:Assessing how individuals perform different activities is key information for modeling health states of individuals and populations. Descriptions of activity performance in clinical free text are complex, including syntactic negation and similarities to textual entailment tasks. We explore a variety of methods for the novel task of classifying four types of assertions about activity performance: Able, Unable, Unclear, and None (no information). We find that ensembling an SVM trained with lexical features and a CNN achieves 77.9% macro F1 score on our task, and yields nearly 80% recall on the rare Unclear and Unable samples. Finally, we highlight several challenges in classifying performance assertions, including capturing information about sources of assistance, incorporating syntactic structure and negation scope, and handling new modalities at test time. Our findings establish a strong baseline for this novel task, and identify intriguing areas for further research.
Characterizing the impact of geometric properties of word embeddings on task performance
Apr 09, 2019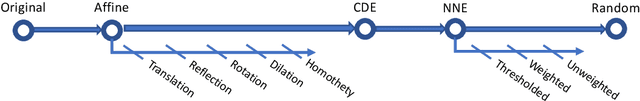
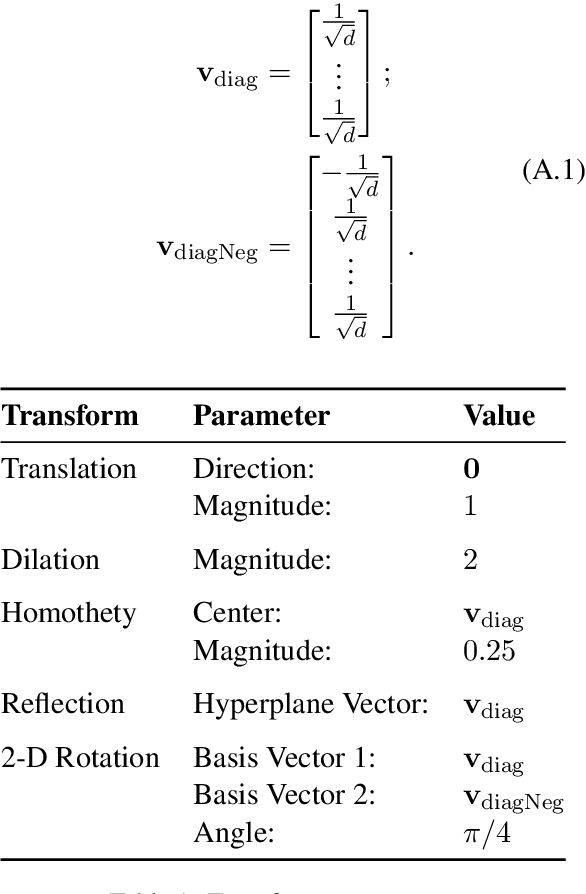
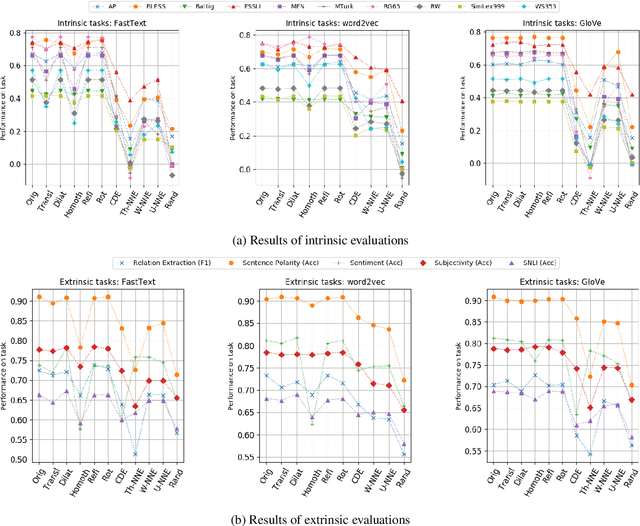
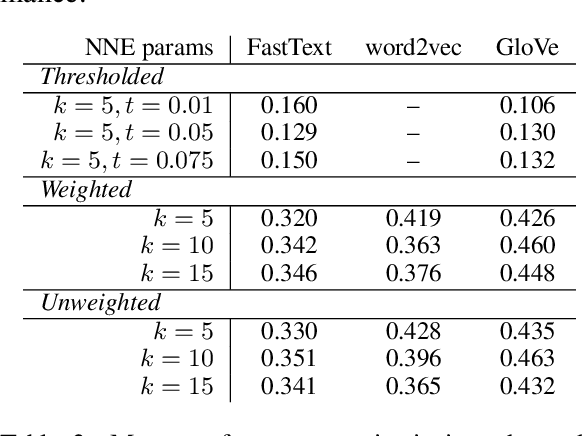
Abstract:Analysis of word embedding properties to inform their use in downstream NLP tasks has largely been studied by assessing nearest neighbors. However, geometric properties of the continuous feature space contribute directly to the use of embedding features in downstream models, and are largely unexplored. We consider four properties of word embedding geometry, namely: position relative to the origin, distribution of features in the vector space, global pairwise distances, and local pairwise distances. We define a sequence of transformations to generate new embeddings that expose subsets of these properties to downstream models and evaluate change in task performance to understand the contribution of each property to NLP models. We transform publicly available pretrained embeddings from three popular toolkits (word2vec, GloVe, and FastText) and evaluate on a variety of intrinsic tasks, which model linguistic information in the vector space, and extrinsic tasks, which use vectors as input to machine learning models. We find that intrinsic evaluations are highly sensitive to absolute position, while extrinsic tasks rely primarily on local similarity. Our findings suggest that future embedding models and post-processing techniques should focus primarily on similarity to nearby points in vector space.
Jointly Embedding Entities and Text with Distant Supervision
Jul 09, 2018
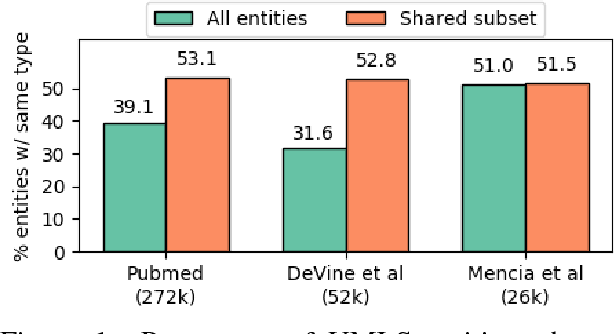
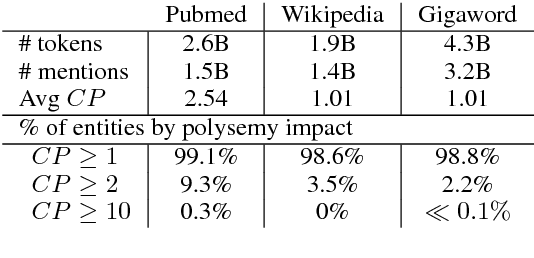
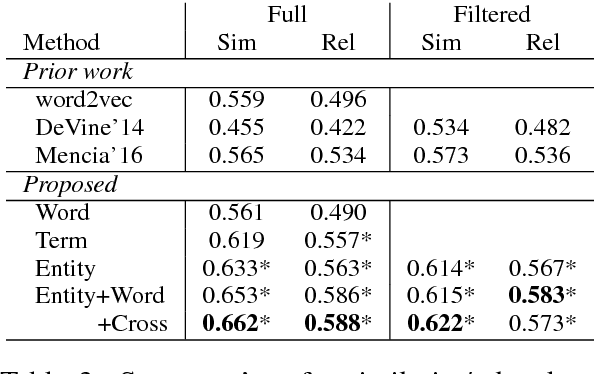
Abstract:Learning representations for knowledge base entities and concepts is becoming increasingly important for NLP applications. However, recent entity embedding methods have relied on structured resources that are expensive to create for new domains and corpora. We present a distantly-supervised method for jointly learning embeddings of entities and text from an unnanotated corpus, using only a list of mappings between entities and surface forms. We learn embeddings from open-domain and biomedical corpora, and compare against prior methods that rely on human-annotated text or large knowledge graph structure. Our embeddings capture entity similarity and relatedness better than prior work, both in existing biomedical datasets and a new Wikipedia-based dataset that we release to the community. Results on analogy completion and entity sense disambiguation indicate that entities and words capture complementary information that can be effectively combined for downstream use.
Embedding Transfer for Low-Resource Medical Named Entity Recognition: A Case Study on Patient Mobility
Jun 07, 2018
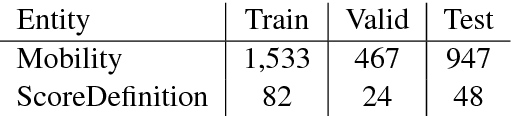

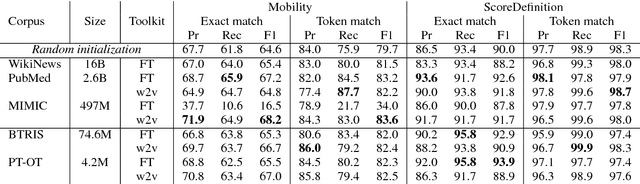
Abstract:Functioning is gaining recognition as an important indicator of global health, but remains under-studied in medical natural language processing research. We present the first analysis of automatically extracting descriptions of patient mobility, using a recently-developed dataset of free text electronic health records. We frame the task as a named entity recognition (NER) problem, and investigate the applicability of NER techniques to mobility extraction. As text corpora focused on patient functioning are scarce, we explore domain adaptation of word embeddings for use in a recurrent neural network NER system. We find that embeddings trained on a small in-domain corpus perform nearly as well as those learned from large out-of-domain corpora, and that domain adaptation techniques yield additional improvements in both precision and recall. Our analysis identifies several significant challenges in extracting descriptions of patient mobility, including the length and complexity of annotated entities and high linguistic variability in mobility descriptions.
Insights into Analogy Completion from the Biomedical Domain
Jun 07, 2017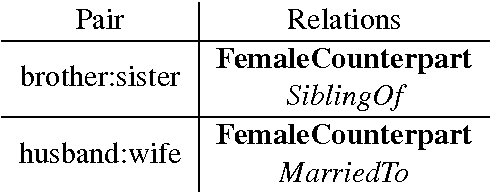
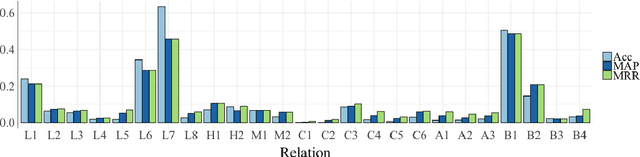
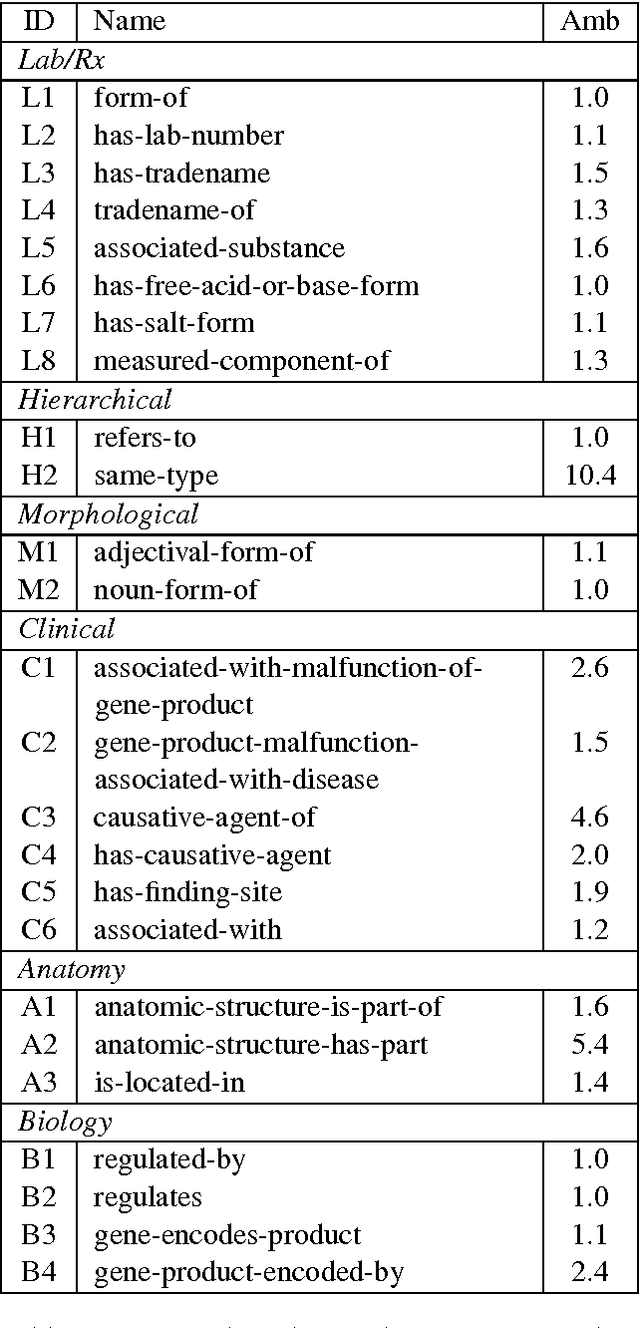
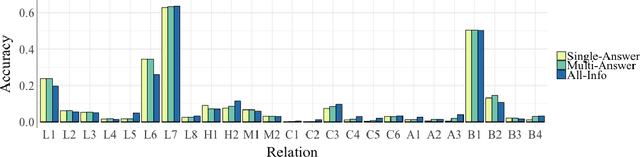
Abstract:Analogy completion has been a popular task in recent years for evaluating the semantic properties of word embeddings, but the standard methodology makes a number of assumptions about analogies that do not always hold, either in recent benchmark datasets or when expanding into other domains. Through an analysis of analogies in the biomedical domain, we identify three assumptions: that of a Single Answer for any given analogy, that the pairs involved describe the Same Relationship, and that each pair is Informative with respect to the other. We propose modifying the standard methodology to relax these assumptions by allowing for multiple correct answers, reporting MAP and MRR in addition to accuracy, and using multiple example pairs. We further present BMASS, a novel dataset for evaluating linguistic regularities in biomedical embeddings, and demonstrate that the relationships described in the dataset pose significant semantic challenges to current word embedding methods.
Second-Order Word Embeddings from Nearest Neighbor Topological Features
May 23, 2017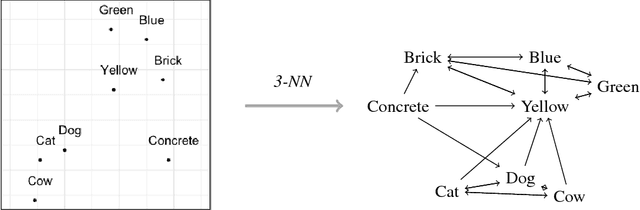
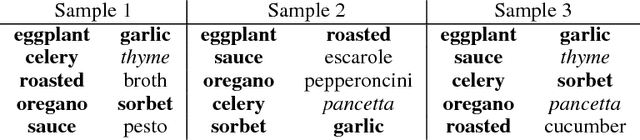

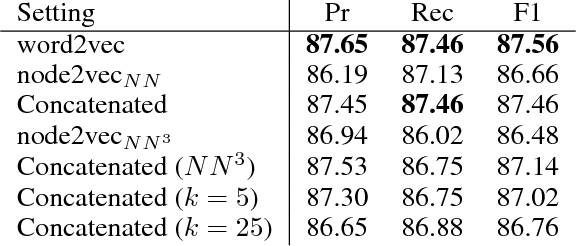
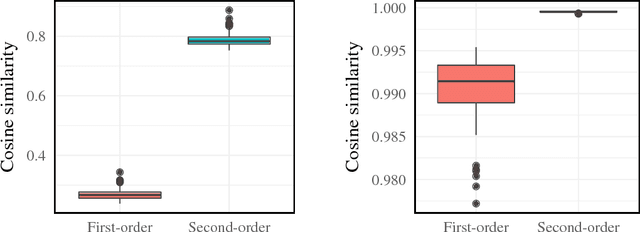
Abstract:We introduce second-order vector representations of words, induced from nearest neighborhood topological features in pre-trained contextual word embeddings. We then analyze the effects of using second-order embeddings as input features in two deep natural language processing models, for named entity recognition and recognizing textual entailment, as well as a linear model for paraphrase recognition. Surprisingly, we find that nearest neighbor information alone is sufficient to capture most of the performance benefits derived from using pre-trained word embeddings. Furthermore, second-order embeddings are able to handle highly heterogeneous data better than first-order representations, though at the cost of some specificity. Additionally, augmenting contextual embeddings with second-order information further improves model performance in some cases. Due to variance in the random initializations of word embeddings, utilizing nearest neighbor features from multiple first-order embedding samples can also contribute to downstream performance gains. Finally, we identify intriguing characteristics of second-order embedding spaces for further research, including much higher density and different semantic interpretations of cosine similarity.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge