Image Compositing for Segmentation of Surgical Tools without Manual Annotations
Feb 18, 2021Luis C. Garcia-Peraza-Herrera, Lucas Fidon, Claudia D'Ettorre, Danail Stoyanov, Tom Vercauteren, Sebastien Ourselin
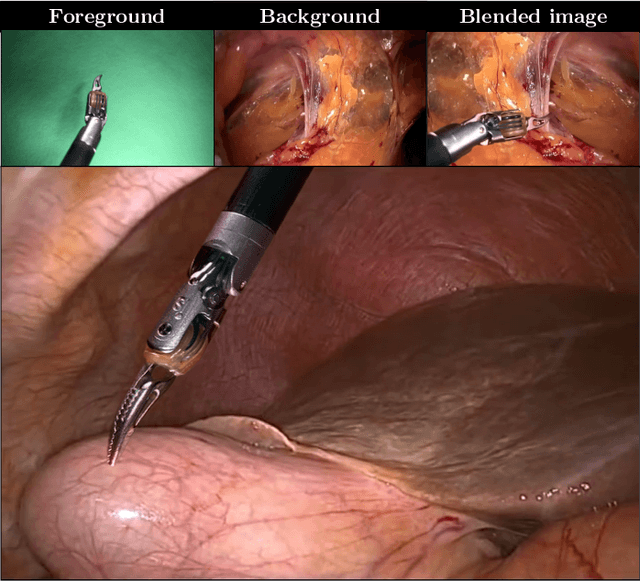
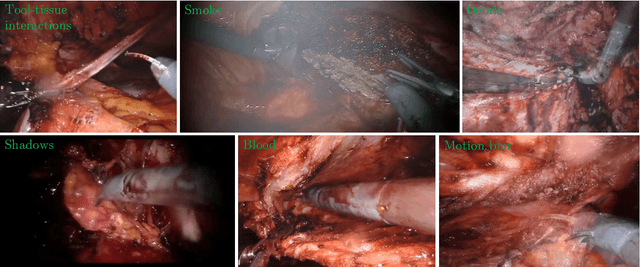
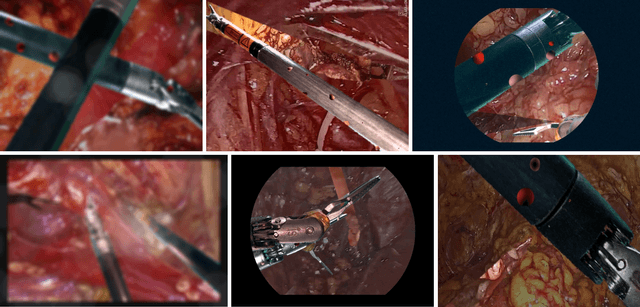
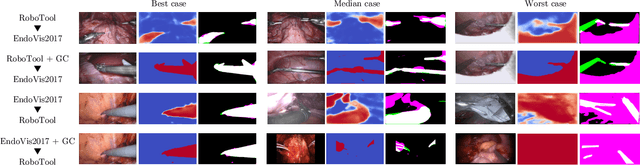
Producing manual, pixel-accurate, image segmentation labels is tedious and time-consuming. This is often a rate-limiting factor when large amounts of labeled images are required, such as for training deep convolutional networks for instrument-background segmentation in surgical scenes. No large datasets comparable to industry standards in the computer vision community are available for this task. To circumvent this problem, we propose to automate the creation of a realistic training dataset by exploiting techniques stemming from special effects and harnessing them to target training performance rather than visual appeal. Foreground data is captured by placing sample surgical instruments over a chroma key (a.k.a. green screen) in a controlled environment, thereby making extraction of the relevant image segment straightforward. Multiple lighting conditions and viewpoints can be captured and introduced in the simulation by moving the instruments and camera and modulating the light source. Background data is captured by collecting videos that do not contain instruments. In the absence of pre-existing instrument-free background videos, minimal labeling effort is required, just to select frames that do not contain surgical instruments from videos of surgical interventions freely available online. We compare different methods to blend instruments over tissue and propose a novel data augmentation approach that takes advantage of the plurality of options. We show that by training a vanilla U-Net on semi-synthetic data only and applying a simple post-processing, we are able to match the results of the same network trained on a publicly available manually labeled real dataset.
Gesture Recognition in Robotic Surgery: a Review
Jan 29, 2021Beatrice van Amsterdam, Matthew J. Clarkson, Danail Stoyanov
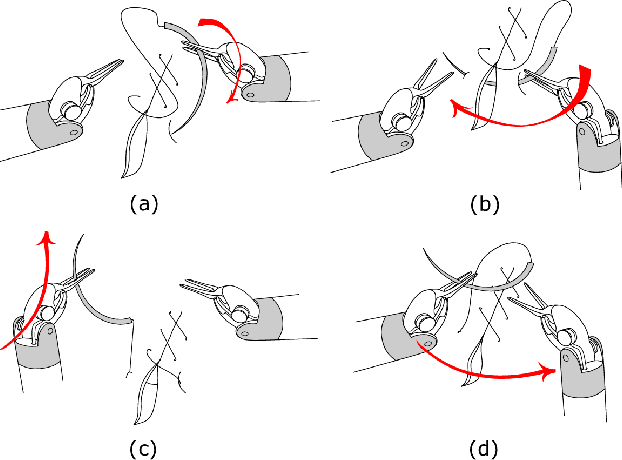
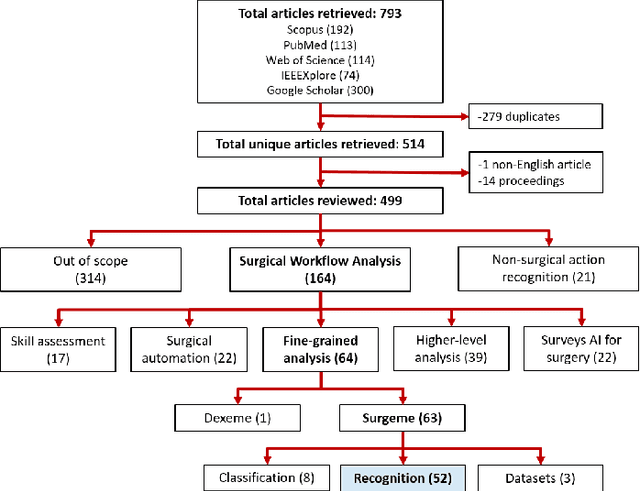
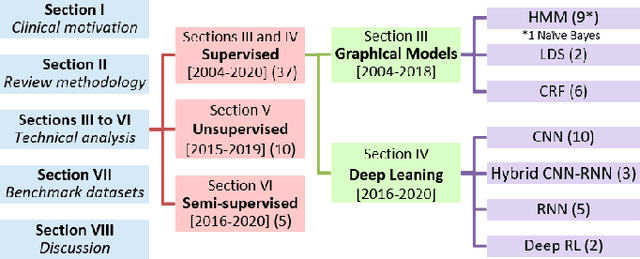
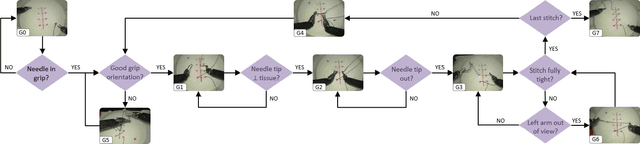
Objective: Surgical activity recognition is a fundamental step in computer-assisted interventions. This paper reviews the state-of-the-art in methods for automatic recognition of fine-grained gestures in robotic surgery focusing on recent data-driven approaches and outlines the open questions and future research directions. Methods: An article search was performed on 5 bibliographic databases with the following search terms: robotic, robot-assisted, JIGSAWS, surgery, surgical, gesture, fine-grained, surgeme, action, trajectory, segmentation, recognition, parsing. Selected articles were classified based on the level of supervision required for training and divided into different groups representing major frameworks for time series analysis and data modelling. Results: A total of 52 articles were reviewed. The research field is showing rapid expansion, with the majority of articles published in the last 4 years. Deep-learning-based temporal models with discriminative feature extraction and multi-modal data integration have demonstrated promising results on small surgical datasets. Currently, unsupervised methods perform significantly less well than the supervised approaches. Conclusion: The development of large and diverse open-source datasets of annotated demonstrations is essential for development and validation of robust solutions for surgical gesture recognition. While new strategies for discriminative feature extraction and knowledge transfer, or unsupervised and semi-supervised approaches, can mitigate the need for data and labels, they have not yet been demonstrated to achieve comparable performance. Important future research directions include detection and forecast of gesture-specific errors and anomalies. Significance: This paper is a comprehensive and structured analysis of surgical gesture recognition methods aiming to summarize the status of this rapidly evolving field.
Stereo Correspondence and Reconstruction of Endoscopic Data Challenge
Jan 28, 2021Max Allan, Jonathan Mcleod, Congcong Wang, Jean Claude Rosenthal, Zhenglei Hu, Niklas Gard, Peter Eisert, Ke Xue Fu, Trevor Zeffiro, Wenyao Xia, Zhanshi Zhu, Huoling Luo, Fucang Jia, Xiran Zhang, Xiaohong Li, Lalith Sharan, Tom Kurmann, Sebastian Schmid, Raphael Sznitman, Dimitris Psychogyios, Mahdi Azizian, Danail Stoyanov, Lena Maier-Hein, Stefanie Speidel
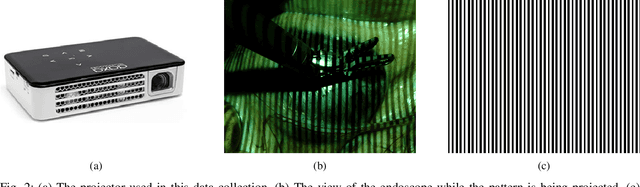


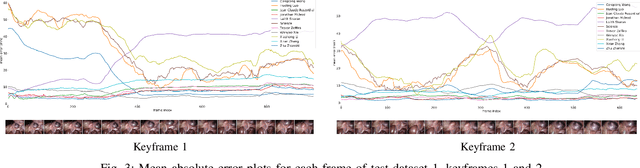
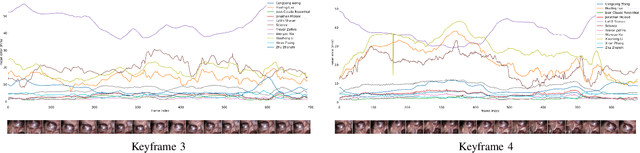
The stereo correspondence and reconstruction of endoscopic data sub-challenge was organized during the Endovis challenge at MICCAI 2019 in Shenzhen, China. The task was to perform dense depth estimation using 7 training datasets and 2 test sets of structured light data captured using porcine cadavers. These were provided by a team at Intuitive Surgical. 10 teams participated in the challenge day. This paper contains 3 additional methods which were submitted after the challenge finished as well as a supplemental section from these teams on issues they found with the dataset.
Towards a Computed-Aided Diagnosis System in Colonoscopy: Automatic Polyp Segmentation Using Convolution Neural Networks
Jan 15, 2021Patrick Brandao, Odysseas Zisimopoulos, Evangelos Mazomenos, Gastone Ciuti, Jorge Bernal, Marco Visentini-Scarzanella, Arianna Menciassi, Paolo Dario, Anastasios Koulaouzidis, Alberto Arezzo, David J Hawkes, Danail Stoyanov
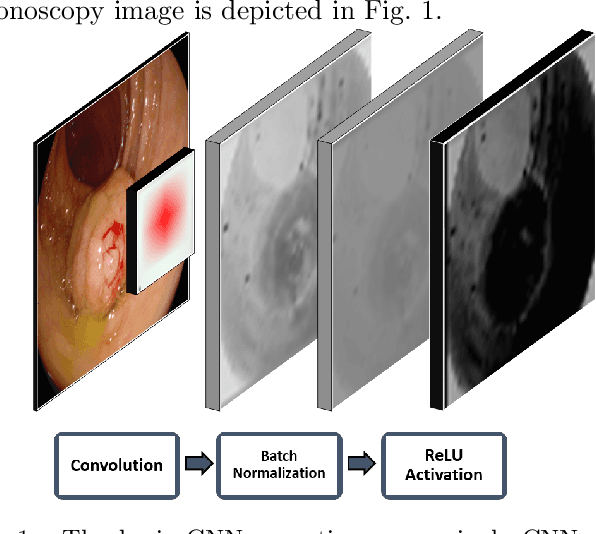
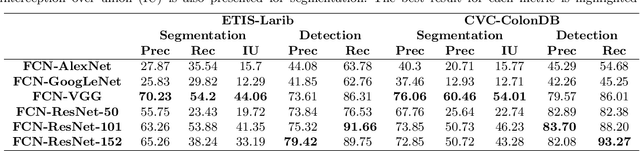
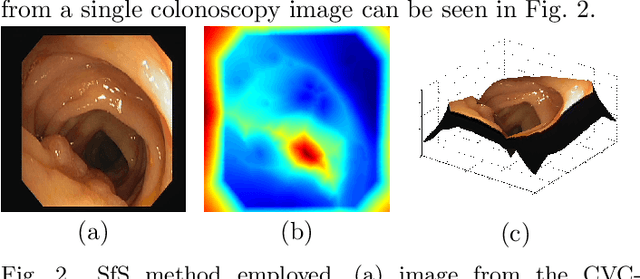
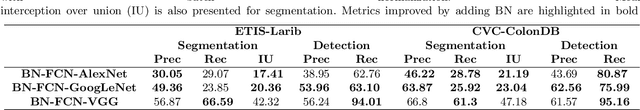
Early diagnosis is essential for the successful treatment of bowel cancers including colorectal cancer (CRC) and capsule endoscopic imaging with robotic actuation can be a valuable diagnostic tool when combined with automated image analysis. We present a deep learning rooted detection and segmentation framework for recognizing lesions in colonoscopy and capsule endoscopy images. We restructure established convolution architectures, such as VGG and ResNets, by converting them into fully-connected convolution networks (FCNs), fine-tune them and study their capabilities for polyp segmentation and detection. We additionally use Shape from-Shading (SfS) to recover depth and provide a richer representation of the tissue's structure in colonoscopy images. Depth is incorporated into our network models as an additional input channel to the RGB information and we demonstrate that the resulting network yields improved performance. Our networks are tested on publicly available datasets and the most accurate segmentation model achieved a mean segmentation IU of 47.78% and 56.95% on the ETIS-Larib and CVC-Colon datasets, respectively. For polyp detection, the top performing models we propose surpass the current state of the art with detection recalls superior to 90% for all datasets tested. To our knowledge, we present the first work to use FCNs for polyp segmentation in addition to proposing a novel combination of SfS and RGB that boosts performance
* 10 pages, 6 figures
SERV-CT: A disparity dataset from CT for validation of endoscopic 3D reconstruction
Dec 22, 2020P. J. "Eddie'' Edwards, Dimitris Psychogyios, Stefanie Speidel, Lena Maier-Hein, Danail Stoyanov
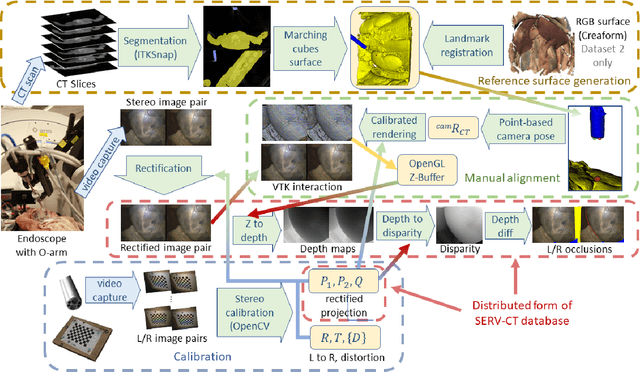
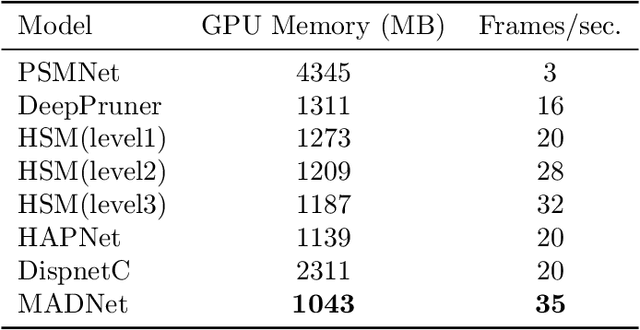
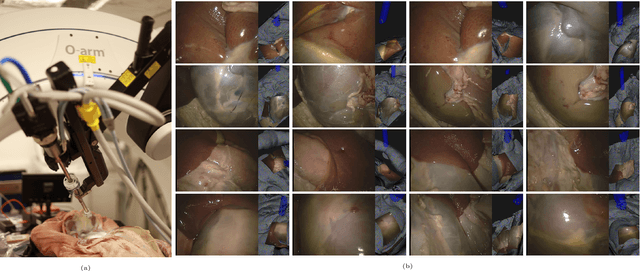
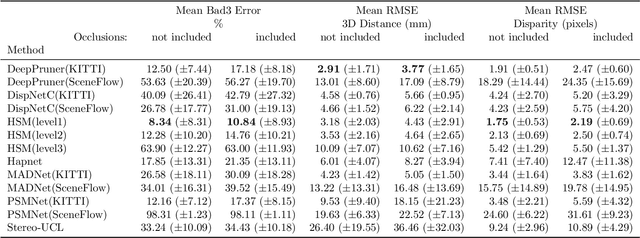
In computer vision, reference datasets have been highly successful in promoting algorithmic development in stereo reconstruction. Surgical scenes gives rise to specific problems, including the lack of clear corner features, highly specular surfaces and the presence of blood and smoke. Publicly available datasets have been produced using CT and either phantom images or biological tissue samples covering a relatively small region of the endoscope field-of-view. We present a stereo-endoscopic reconstruction validation dataset based on CT (SERV-CT). Two {\it ex vivo} small porcine full torso cadavers were placed within the view of the endoscope with both the endoscope and target anatomy visible in the CT scan. Orientation of the endoscope was manually aligned to the stereoscopic view. Reference disparities and occlusions were calculated for 8 stereo pairs from each sample. For the second sample an RGB surface was acquired to aid alignment of smooth, featureless surfaces. Repeated manual alignments showed an RMS disparity accuracy of ~2 pixels and a depth accuracy of ~2mm. The reference dataset includes endoscope image pairs with corresponding calibration, disparities, depths and occlusions covering the majority of the endoscopic image and a range of tissue types. Smooth specular surfaces and images with significant variation of depth are included. We assessed the performance of various stereo algorithms from online available repositories. There is a significant variation between algorithms, highlighting some of the challenges of surgical endoscopic images. The SERV-CT dataset provides an easy to use stereoscopic validation for surgical applications with smooth reference disparities and depths with coverage over the majority of the endoscopic images. This complements existing resources well and we hope will aid the development of surgical endoscopic anatomical reconstruction algorithms.
Surgical Data Science -- from Concepts to Clinical Translation
Oct 30, 2020Lena Maier-Hein, Matthias Eisenmann, Duygu Sarikaya, Keno März, Toby Collins, Anand Malpani, Johannes Fallert, Hubertus Feussner, Stamatia Giannarou, Pietro Mascagni, Hirenkumar Nakawala, Adrian Park, Carla Pugh, Danail Stoyanov, Swaroop S. Vedula, Beat Peter Müller, Kevin Cleary, Gabor Fichtinger, Germain Forestier, Bernard Gibaud, Teodor Grantcharov, Makoto Hashizume, Hannes Kenngott, Ron Kikinis, Lars Mündermann, Nassir Navab, Sinan Onogur, Raphael Sznitman, Russell Taylor, Minu Dietlinde Tizabi, Martin Wagner, Gregory D. Hager, Thomas Neumuth, Nicolas Padoy, Pierre Jannin, Stefanie Speidel
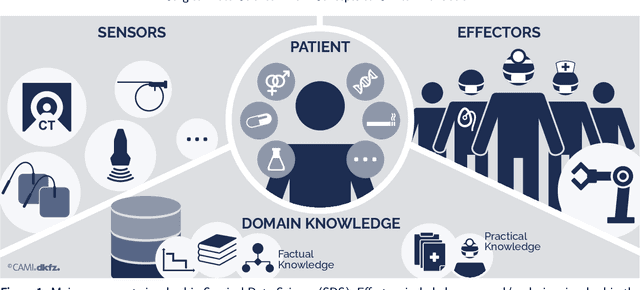
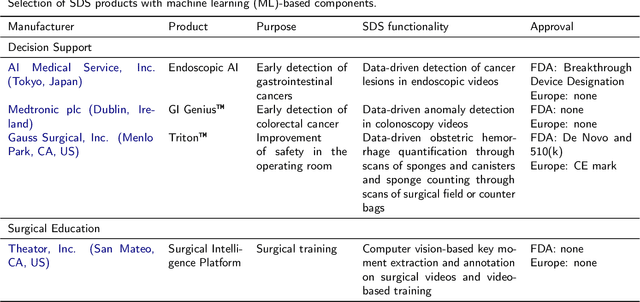
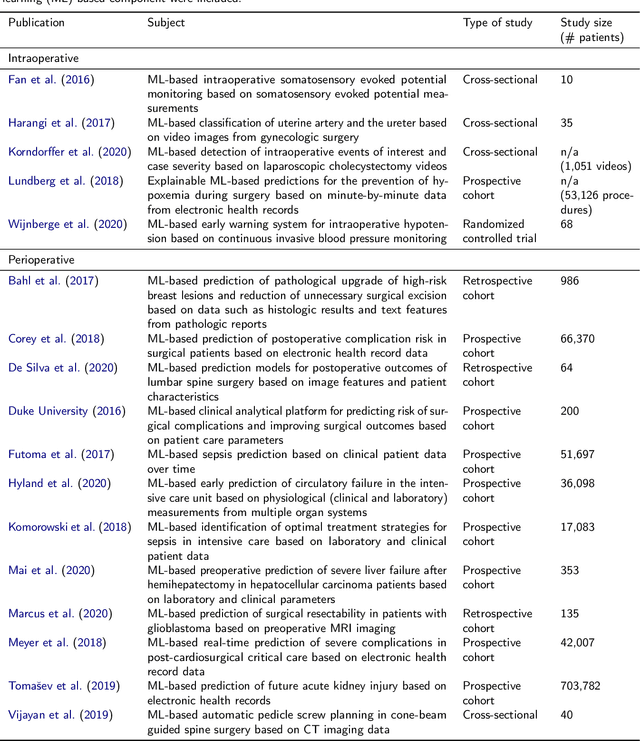
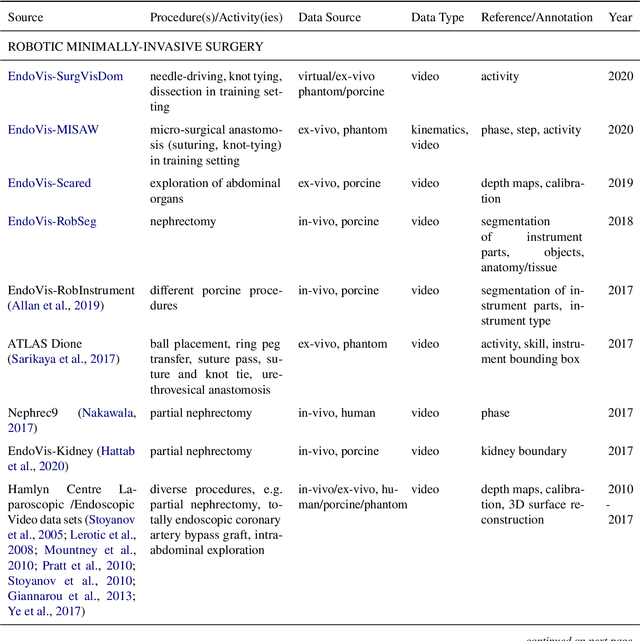
Recent developments in data science in general and machine learning in particular have transformed the way experts envision the future of surgery. Surgical data science is a new research field that aims to improve the quality of interventional healthcare through the capture, organization, analysis and modeling of data. While an increasing number of data-driven approaches and clinical applications have been studied in the fields of radiological and clinical data science, translational success stories are still lacking in surgery. In this publication, we shed light on the underlying reasons and provide a roadmap for future advances in the field. Based on an international workshop involving leading researchers in the field of surgical data science, we review current practice, key achievements and initiatives as well as available standards and tools for a number of topics relevant to the field, namely (1) technical infrastructure for data acquisition, storage and access in the presence of regulatory constraints, (2) data annotation and sharing and (3) data analytics. Drawing from this extensive review, we present current challenges for technology development and (4) describe a roadmap for faster clinical translation and exploitation of the full potential of surgical data science.
Detecting small polyps using a Dynamic SSD-GAN
Oct 29, 2020Daniel C. Ohrenstein, Patrick Brandao, Daniel Toth, Laurence Lovat, Danail Stoyanov, Peter Mountney
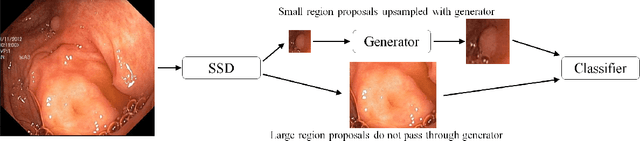
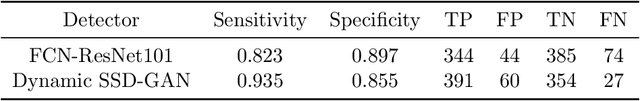
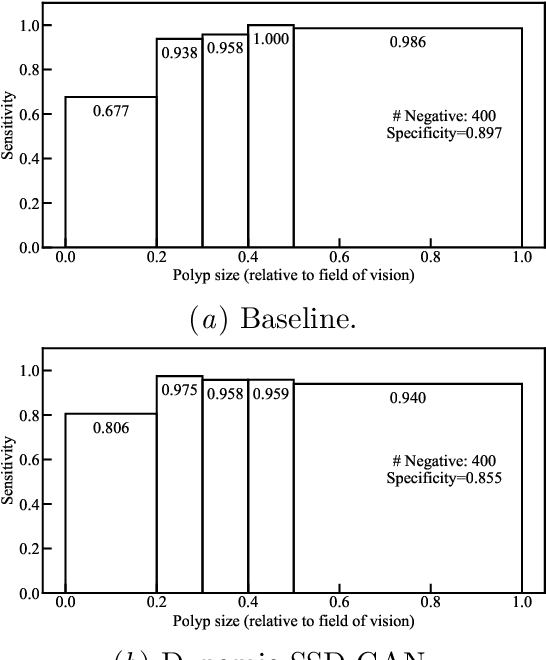
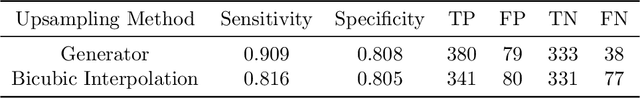
Endoscopic examinations are used to inspect the throat, stomach and bowel for polyps which could develop into cancer. Machine learning systems can be trained to process colonoscopy images and detect polyps. However, these systems tend to perform poorly on objects which appear visually small in the images. It is shown here that combining the single-shot detector as a region proposal network with an adversarially-trained generator to upsample small region proposals can significantly improve the detection of visually-small polyps. The Dynamic SSD-GAN pipeline introduced in this paper achieved a 12% increase in sensitivity on visually-small polyps compared to a conventional FCN baseline.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge