Benjamin M. Marlin
Challenges and Opportunities in Approximate Bayesian Deep Learning for Intelligent IoT Systems
Dec 03, 2021Abstract:Approximate Bayesian deep learning methods hold significant promise for addressing several issues that occur when deploying deep learning components in intelligent systems, including mitigating the occurrence of over-confident errors and providing enhanced robustness to out of distribution examples. However, the computational requirements of existing approximate Bayesian inference methods can make them ill-suited for deployment in intelligent IoT systems that include lower-powered edge devices. In this paper, we present a range of approximate Bayesian inference methods for supervised deep learning and highlight the challenges and opportunities when applying these methods on current edge hardware. We highlight several potential solutions to decreasing model storage requirements and improving computational scalability, including model pruning and distillation methods.
Heteroscedastic Temporal Variational Autoencoder For Irregularly Sampled Time Series
Jul 23, 2021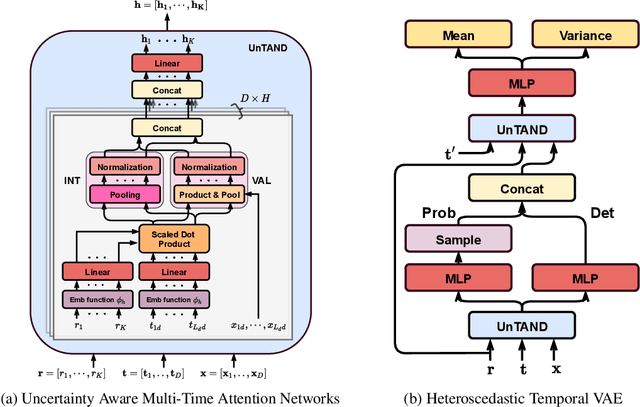
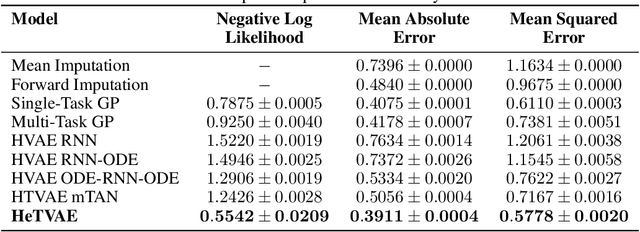
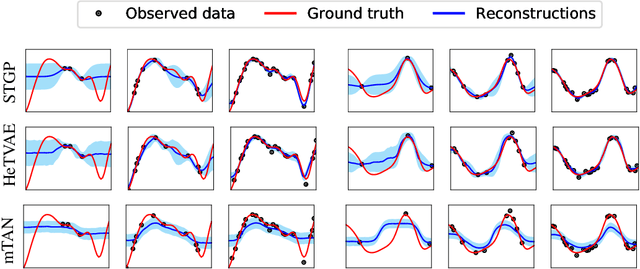
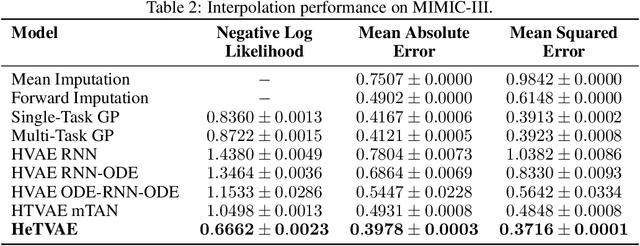
Abstract:Irregularly sampled time series commonly occur in several domains where they present a significant challenge to standard deep learning models. In this paper, we propose a new deep learning framework for probabilistic interpolation of irregularly sampled time series that we call the Heteroscedastic Temporal Variational Autoencoder (HeTVAE). HeTVAE includes a novel input layer to encode information about input observation sparsity, a temporal VAE architecture to propagate uncertainty due to input sparsity, and a heteroscedastic output layer to enable variable uncertainty in output interpolations. Our results show that the proposed architecture is better able to reflect variable uncertainty through time due to sparse and irregular sampling than a range of baseline and traditional models, as well as recently proposed deep latent variable models that use homoscedastic output layers.
Post-hoc loss-calibration for Bayesian neural networks
Jun 13, 2021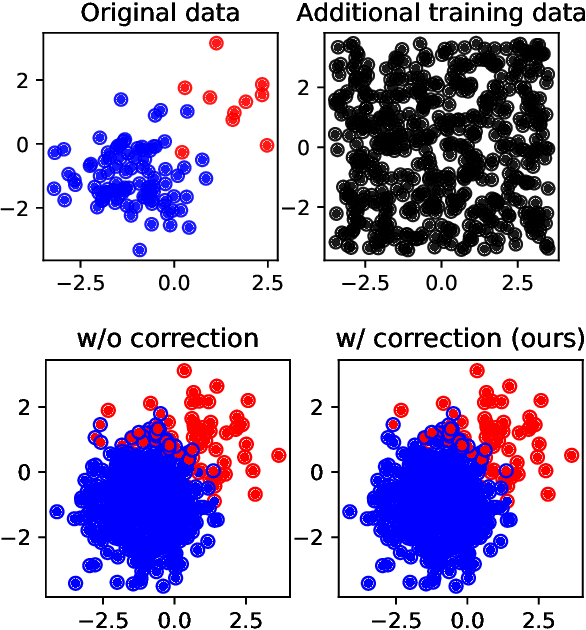
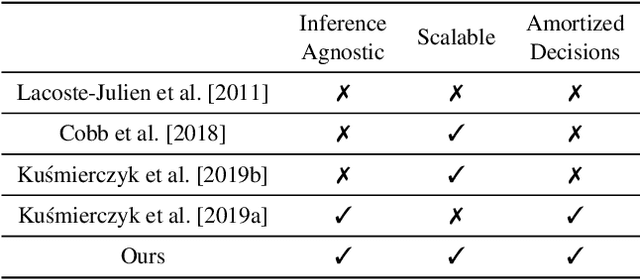
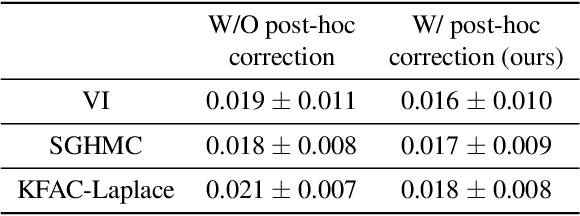
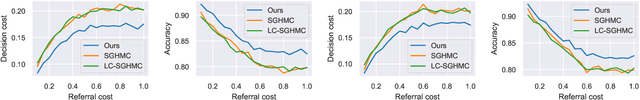
Abstract:Bayesian decision theory provides an elegant framework for acting optimally under uncertainty when tractable posterior distributions are available. Modern Bayesian models, however, typically involve intractable posteriors that are approximated with, potentially crude, surrogates. This difficulty has engendered loss-calibrated techniques that aim to learn posterior approximations that favor high-utility decisions. In this paper, focusing on Bayesian neural networks, we develop methods for correcting approximate posterior predictive distributions encouraging them to prefer high-utility decisions. In contrast to previous work, our approach is agnostic to the choice of the approximate inference algorithm, allows for efficient test time decision making through amortization, and empirically produces higher quality decisions. We demonstrate the effectiveness of our approach through controlled experiments spanning a diversity of tasks and datasets.
Multi-Time Attention Networks for Irregularly Sampled Time Series
Jan 25, 2021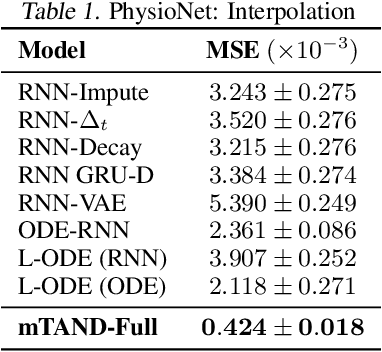
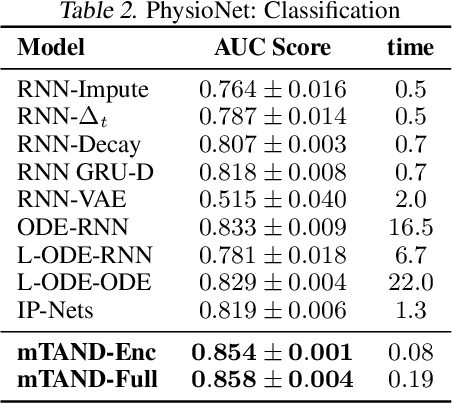
Abstract:Irregular sampling occurs in many time series modeling applications where it presents a significant challenge to standard deep learning models. This work is motivated by the analysis of physiological time series data in electronic health records, which are sparse, irregularly sampled, and multivariate. In this paper, we propose a new deep learning framework for this setting that we call Multi-Time Attention Networks. Multi-Time Attention Networks learn an embedding of continuous-time values and use an attention mechanism to produce a fixed-length representation of a time series containing a variable number of observations. We investigate the performance of our framework on interpolation and classification tasks using multiple datasets. Our results show that our approach performs as well or better than a range of baseline and recently proposed models while offering significantly faster training times than current state-of-the-art methods.
A Survey on Principles, Models and Methods for Learning from Irregularly Sampled Time Series
Jan 05, 2021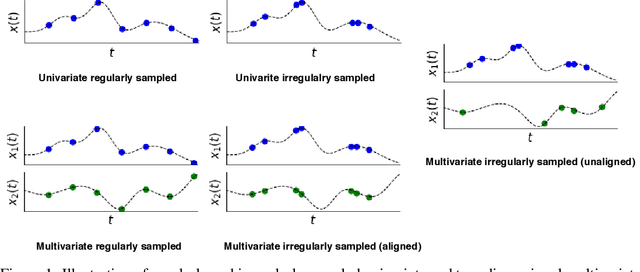
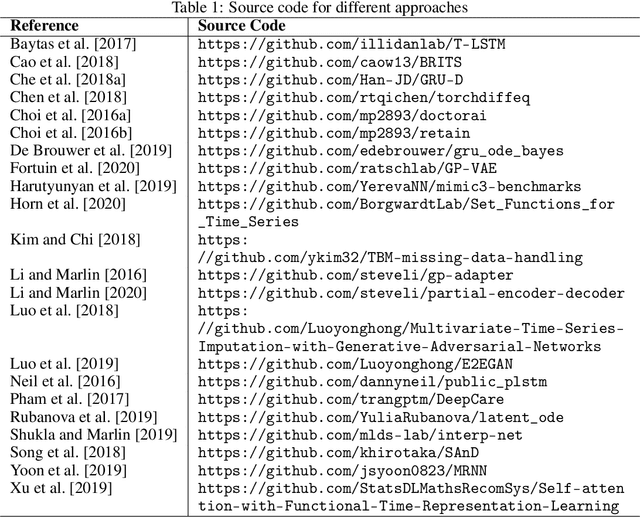
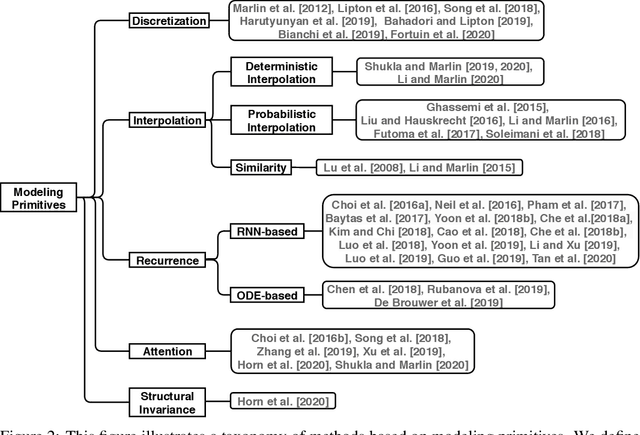
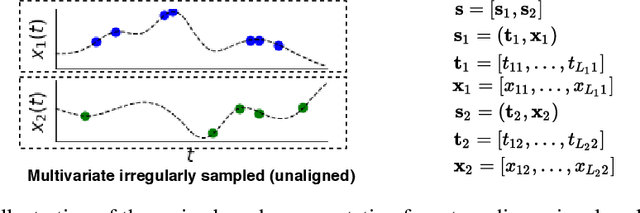
Abstract:Irregularly sampled time series data arise naturally in many application domains including biology, ecology, climate science, astronomy, and health. Such data represent fundamental challenges to many classical models from machine learning and statistics due to the presence of non-uniform intervals between observations. However, there has been significant progress within the machine learning community over the last decade on developing specialized models and architectures for learning from irregularly sampled univariate and multivariate time series data. In this survey, we first describe several axes along which approaches to learning from irregularly sampled time series differ including what data representations they are based on, what modeling primitives they leverage to deal with the fundamental problem of irregular sampling, and what inference tasks they are designed to perform. We then survey the recent literature organized primarily along the axis of modeling primitives. We describe approaches based on temporal discretization, interpolation, recurrence, attention and structural invariance. We discuss similarities and differences between approaches and highlight primary strengths and weaknesses.
Learning from Irregularly-Sampled Time Series: A Missing Data Perspective
Aug 17, 2020
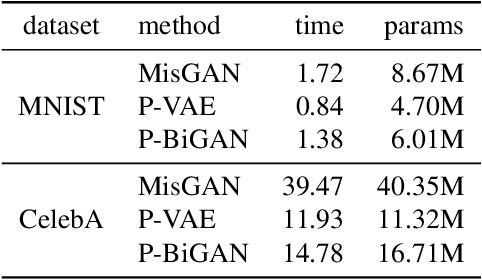
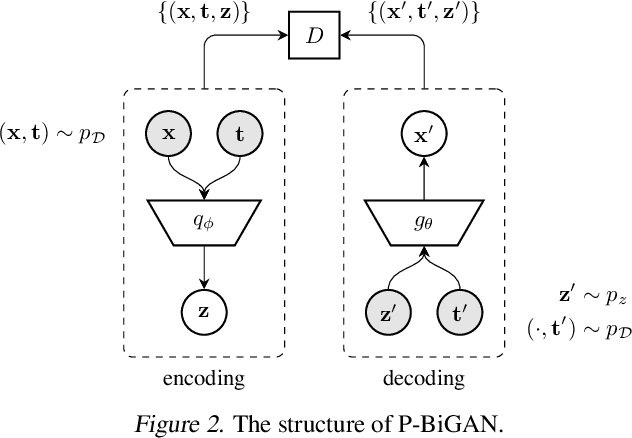
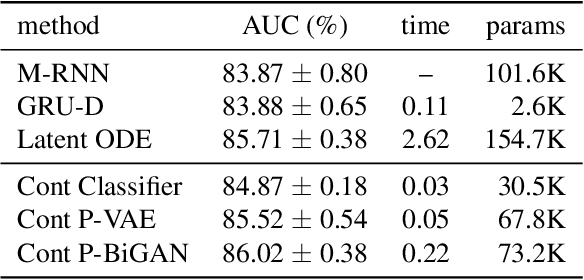
Abstract:Irregularly-sampled time series occur in many domains including healthcare. They can be challenging to model because they do not naturally yield a fixed-dimensional representation as required by many standard machine learning models. In this paper, we consider irregular sampling from the perspective of missing data. We model observed irregularly-sampled time series data as a sequence of index-value pairs sampled from a continuous but unobserved function. We introduce an encoder-decoder framework for learning from such generic indexed sequences. We propose learning methods for this framework based on variational autoencoders and generative adversarial networks. For continuous irregularly-sampled time series, we introduce continuous convolutional layers that can efficiently interface with existing neural network architectures. Experiments show that our models are able to achieve competitive or better classification results on irregularly-sampled multivariate time series compared to recent RNN models while offering significantly faster training times.
URSABench: Comprehensive Benchmarking of Approximate Bayesian Inference Methods for Deep Neural Networks
Jul 08, 2020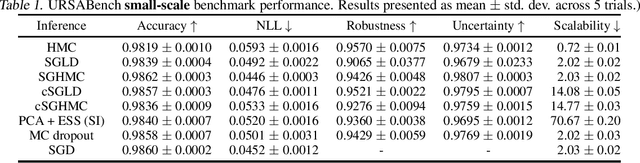
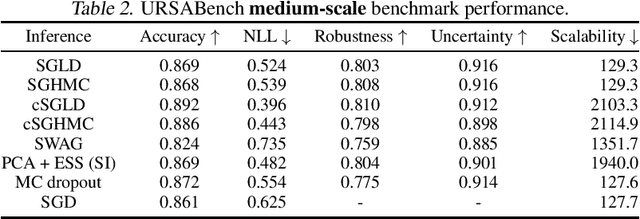
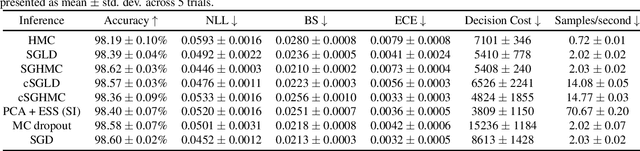
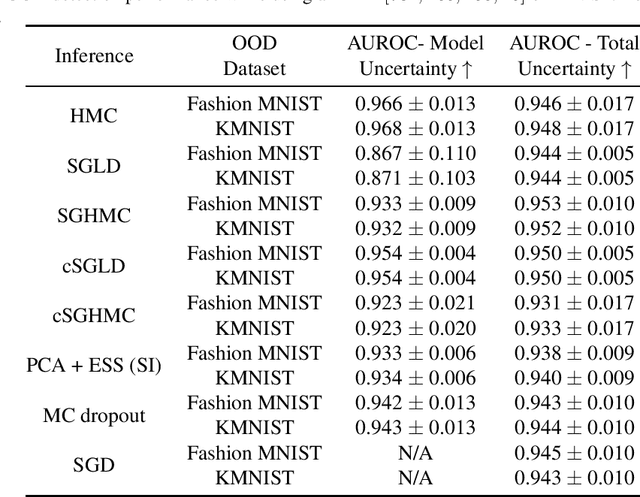
Abstract:While deep learning methods continue to improve in predictive accuracy on a wide range of application domains, significant issues remain with other aspects of their performance including their ability to quantify uncertainty and their robustness. Recent advances in approximate Bayesian inference hold significant promise for addressing these concerns, but the computational scalability of these methods can be problematic when applied to large-scale models. In this paper, we describe initial work on the development ofURSABench(the Uncertainty, Robustness, Scalability, and Accu-racy Benchmark), an open-source suite of bench-marking tools for comprehensive assessment of approximate Bayesian inference methods with a focus on deep learning-based classification tasks
Generalized Bayesian Posterior Expectation Distillation for Deep Neural Networks
May 16, 2020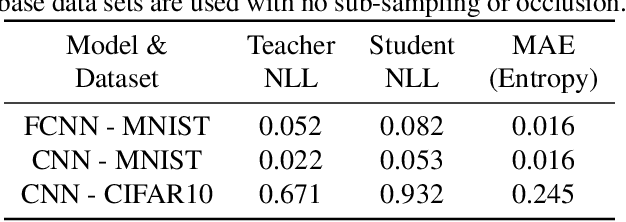
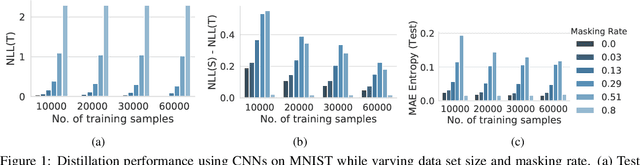
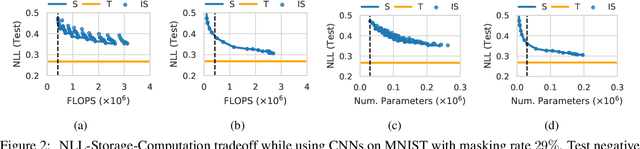
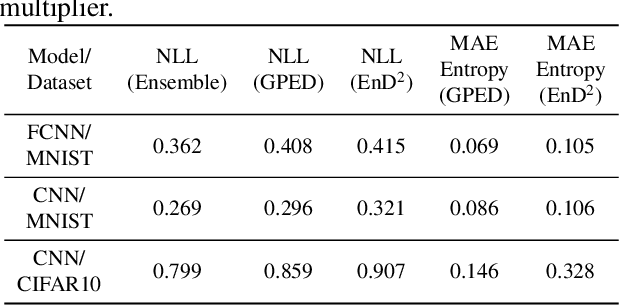
Abstract:In this paper, we present a general framework for distilling expectations with respect to the Bayesian posterior distribution of a deep neural network classifier, extending prior work on the Bayesian Dark Knowledge framework. The proposed framework takes as input "teacher" and student model architectures and a general posterior expectation of interest. The distillation method performs an online compression of the selected posterior expectation using iteratively generated Monte Carlo samples. We focus on the posterior predictive distribution and expected entropy as distillation targets. We investigate several aspects of this framework including the impact of uncertainty and the choice of student model architecture. We study methods for student model architecture search from a speed-storage-accuracy perspective and evaluate down-stream tasks leveraging entropy distillation including uncertainty ranking and out-of-distribution detection.
Integrating Physiological Time Series and Clinical Notes with Deep Learning for Improved ICU Mortality Prediction
Mar 24, 2020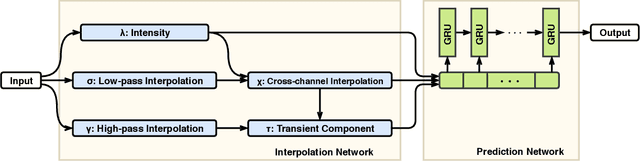
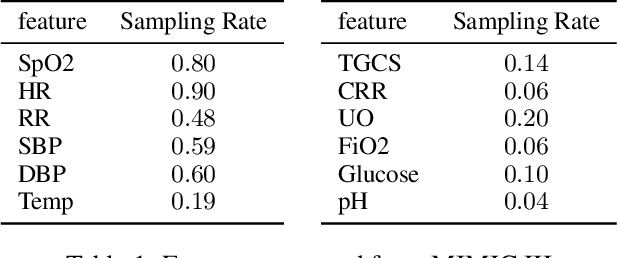
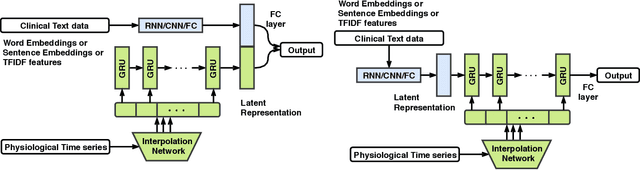
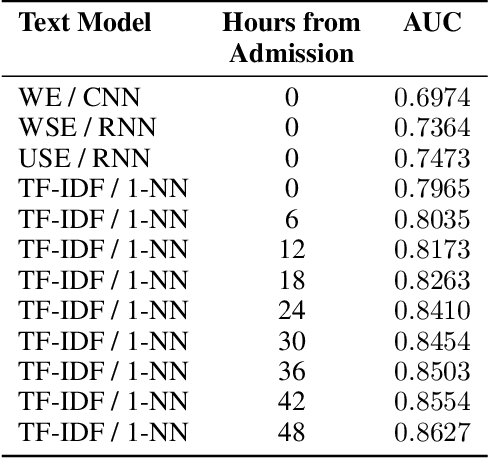
Abstract:Intensive Care Unit Electronic Health Records (ICU EHRs) store multimodal data about patients including clinical notes, sparse and irregularly sampled physiological time series, lab results, and more. To date, most methods designed to learn predictive models from ICU EHR data have focused on a single modality. In this paper, we leverage the recently proposed interpolation-prediction deep learning architecture(Shukla and Marlin 2019) as a basis for exploring how physiological time series data and clinical notes can be integrated into a unified mortality prediction model. We study both early and late fusion approaches and demonstrate how the relative predictive value of clinical text and physiological data change over time. Our results show that a late fusion approach can provide a statistically significant improvement in mortality prediction performance over using individual modalities in isolation.
Assessing the Adversarial Robustness of Monte Carlo and Distillation Methods for Deep Bayesian Neural Network Classification
Feb 07, 2020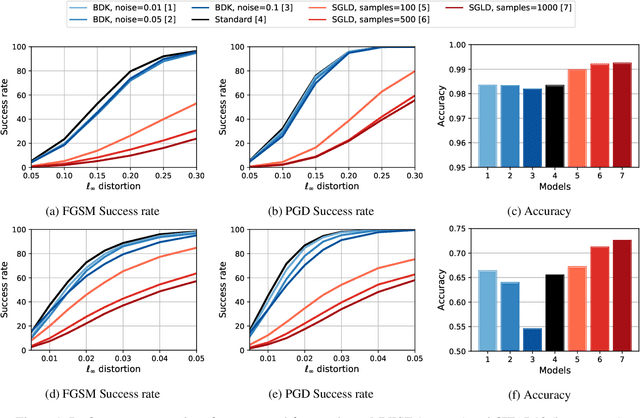
Abstract:In this paper, we consider the problem of assessing the adversarial robustness of deep neural network models under both Markov chain Monte Carlo (MCMC) and Bayesian Dark Knowledge (BDK) inference approximations. We characterize the robustness of each method to two types of adversarial attacks: the fast gradient sign method (FGSM) and projected gradient descent (PGD). We show that full MCMC-based inference has excellent robustness, significantly outperforming standard point estimation-based learning. On the other hand, BDK provides marginal improvements. As an additional contribution, we present a storage-efficient approach to computing adversarial examples for large Monte Carlo ensembles using both the FGSM and PGD attacks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge