Audrey G. Chung
Discovery Radiomics via StochasticNet Sequencers for Cancer Detection
Nov 11, 2015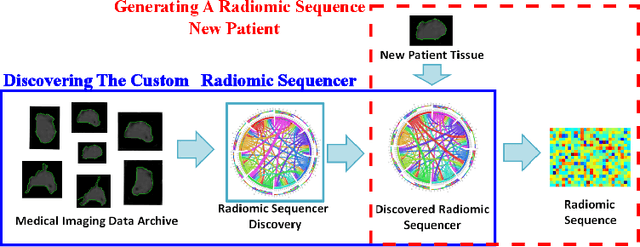
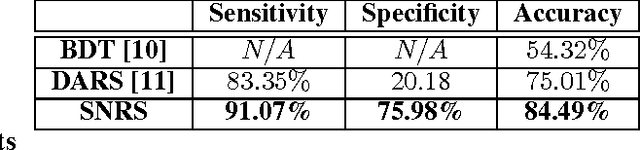
Abstract:Radiomics has proven to be a powerful prognostic tool for cancer detection, and has previously been applied in lung, breast, prostate, and head-and-neck cancer studies with great success. However, these radiomics-driven methods rely on pre-defined, hand-crafted radiomic feature sets that can limit their ability to characterize unique cancer traits. In this study, we introduce a novel discovery radiomics framework where we directly discover custom radiomic features from the wealth of available medical imaging data. In particular, we leverage novel StochasticNet radiomic sequencers for extracting custom radiomic features tailored for characterizing unique cancer tissue phenotype. Using StochasticNet radiomic sequencers discovered using a wealth of lung CT data, we perform binary classification on 42,340 lung lesions obtained from the CT scans of 93 patients in the LIDC-IDRI dataset. Preliminary results show significant improvement over previous state-of-the-art methods, indicating the potential of the proposed discovery radiomics framework for improving cancer screening and diagnosis.
Discovery Radiomics for Multi-Parametric MRI Prostate Cancer Detection
Oct 20, 2015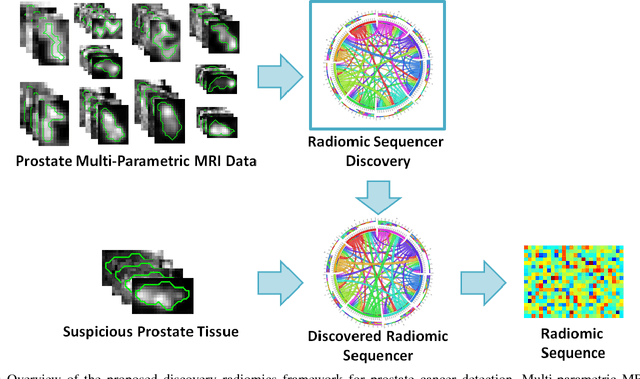
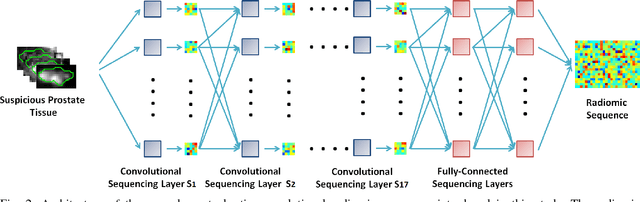

Abstract:Prostate cancer is the most diagnosed form of cancer in Canadian men, and is the third leading cause of cancer death. Despite these statistics, prognosis is relatively good with a sufficiently early diagnosis, making fast and reliable prostate cancer detection crucial. As imaging-based prostate cancer screening, such as magnetic resonance imaging (MRI), requires an experienced medical professional to extensively review the data and perform a diagnosis, radiomics-driven methods help streamline the process and has the potential to significantly improve diagnostic accuracy and efficiency, and thus improving patient survival rates. These radiomics-driven methods currently rely on hand-crafted sets of quantitative imaging-based features, which are selected manually and can limit their ability to fully characterize unique prostate cancer tumour phenotype. In this study, we propose a novel \textit{discovery radiomics} framework for generating custom radiomic sequences tailored for prostate cancer detection. Discovery radiomics aims to uncover abstract imaging-based features that capture highly unique tumour traits and characteristics beyond what can be captured using predefined feature models. In this paper, we discover new custom radiomic sequencers for generating new prostate radiomic sequences using multi-parametric MRI data. We evaluated the performance of the discovered radiomic sequencer against a state-of-the-art hand-crafted radiomic sequencer for computer-aided prostate cancer detection with a feedforward neural network using real clinical prostate multi-parametric MRI data. Results for the discovered radiomic sequencer demonstrate good performance in prostate cancer detection and clinical decision support relative to the hand-crafted radiomic sequencer. The use of discovery radiomics shows potential for more efficient and reliable automatic prostate cancer detection.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge