Andreas Holzinger
The Need for Speed of AI Applications: Performance Comparison of Native vs. Browser-based Algorithm Implementations
Feb 11, 2018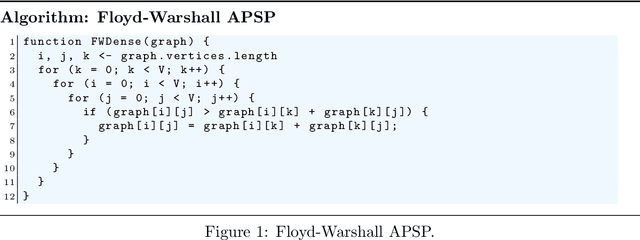



Abstract:AI applications pose increasing demands on performance, so it is not surprising that the era of client-side distributed software is becoming important. On top of many AI applications already using mobile hardware, and even browsers for computationally demanding AI applications, we are already witnessing the emergence of client-side (federated) machine learning algorithms, driven by the interests of large corporations and startups alike. Apart from mathematical and algorithmic concerns, this trend especially demands new levels of computational efficiency from client environments. Consequently, this paper deals with the question of state-of-the-art performance by presenting a comparison study between native code and different browser-based implementations: JavaScript, ASM.js as well as WebAssembly on a representative mix of algorithms. Our results show that current efforts in runtime optimization push the boundaries well towards (and even beyond) native binary performance. We analyze the results obtained and speculate on the reasons behind some surprises, rounding the paper off by outlining future possibilities as well as some of our own research efforts.
What do we need to build explainable AI systems for the medical domain?
Dec 28, 2017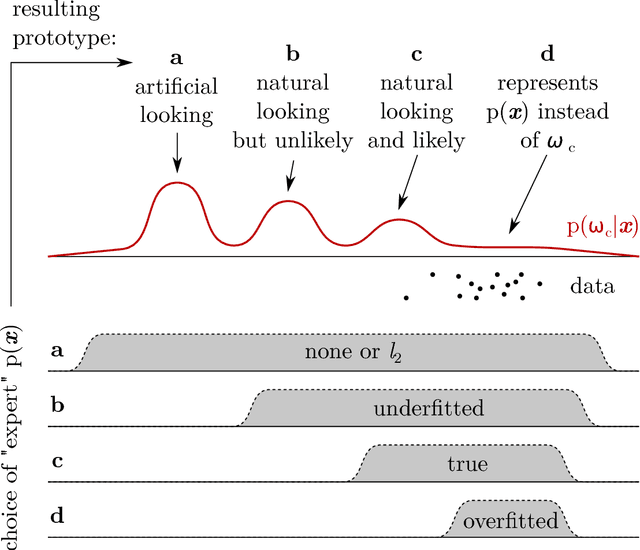
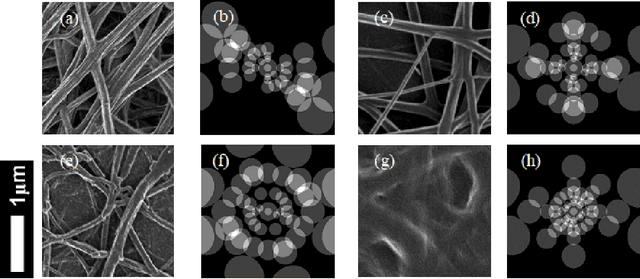
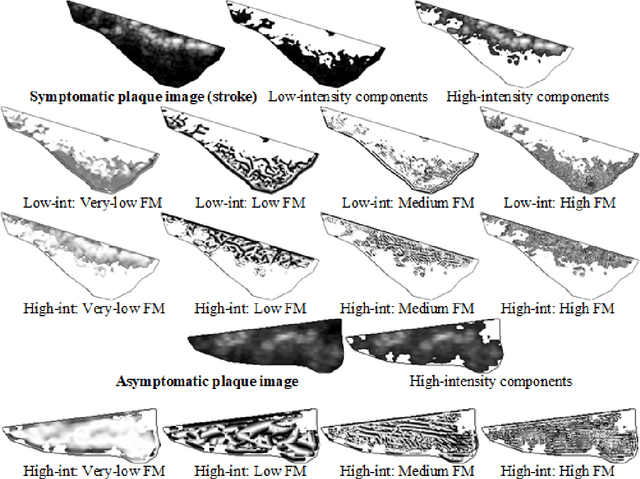

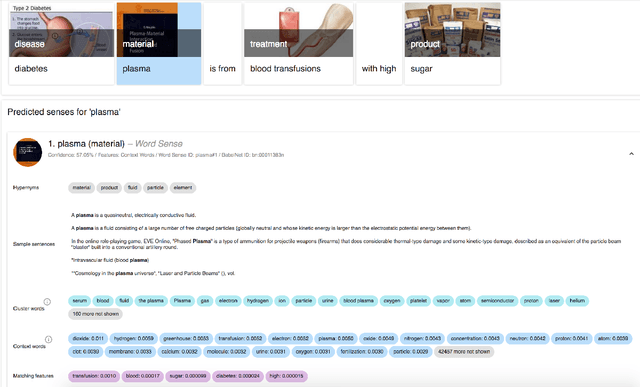
Abstract:Artificial intelligence (AI) generally and machine learning (ML) specifically demonstrate impressive practical success in many different application domains, e.g. in autonomous driving, speech recognition, or recommender systems. Deep learning approaches, trained on extremely large data sets or using reinforcement learning methods have even exceeded human performance in visual tasks, particularly on playing games such as Atari, or mastering the game of Go. Even in the medical domain there are remarkable results. The central problem of such models is that they are regarded as black-box models and even if we understand the underlying mathematical principles, they lack an explicit declarative knowledge representation, hence have difficulty in generating the underlying explanatory structures. This calls for systems enabling to make decisions transparent, understandable and explainable. A huge motivation for our approach are rising legal and privacy aspects. The new European General Data Protection Regulation entering into force on May 25th 2018, will make black-box approaches difficult to use in business. This does not imply a ban on automatic learning approaches or an obligation to explain everything all the time, however, there must be a possibility to make the results re-traceable on demand. In this paper we outline some of our research topics in the context of the relatively new area of explainable-AI with a focus on the application in medicine, which is a very special domain. This is due to the fact that medical professionals are working mostly with distributed heterogeneous and complex sources of data. In this paper we concentrate on three sources: images, *omics data and text. We argue that research in explainable-AI would generally help to facilitate the implementation of AI/ML in the medical domain, and specifically help to facilitate transparency and trust.
Towards the Augmented Pathologist: Challenges of Explainable-AI in Digital Pathology
Dec 18, 2017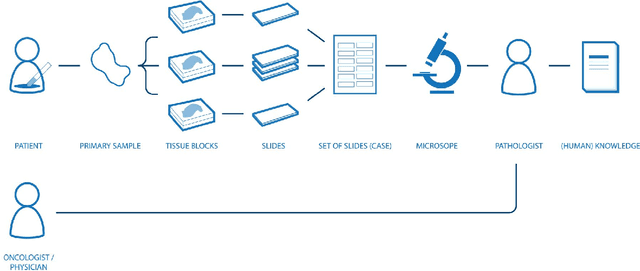
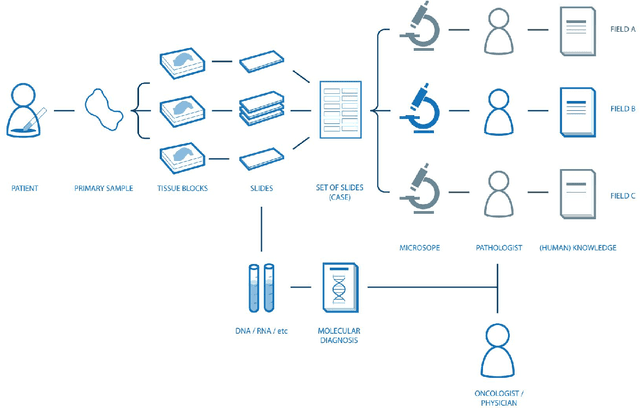
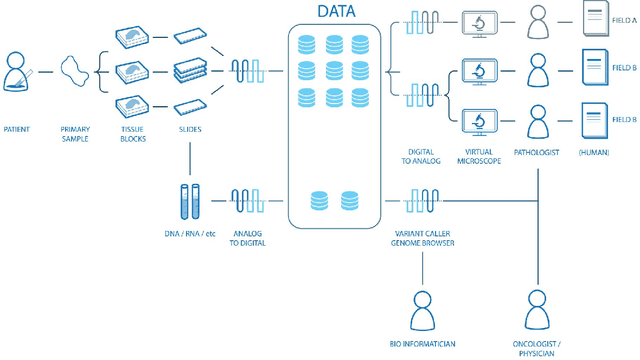
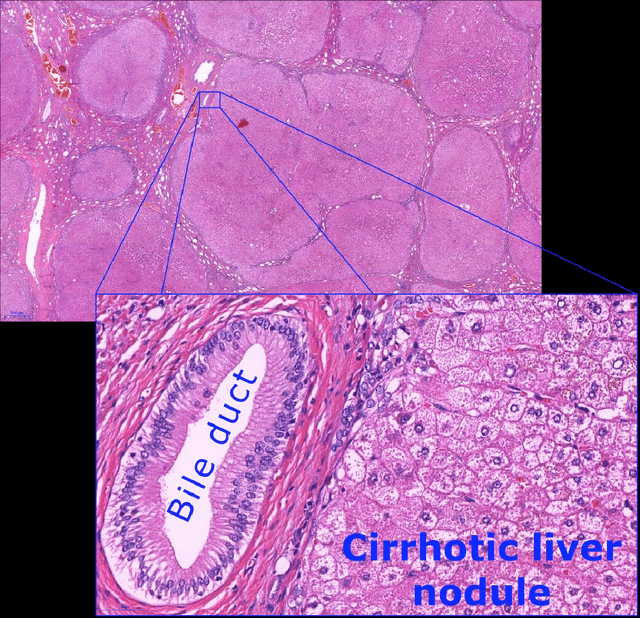
Abstract:Digital pathology is not only one of the most promising fields of diagnostic medicine, but at the same time a hot topic for fundamental research. Digital pathology is not just the transfer of histopathological slides into digital representations. The combination of different data sources (images, patient records, and *omics data) together with current advances in artificial intelligence/machine learning enable to make novel information accessible and quantifiable to a human expert, which is not yet available and not exploited in current medical settings. The grand goal is to reach a level of usable intelligence to understand the data in the context of an application task, thereby making machine decisions transparent, interpretable and explainable. The foundation of such an "augmented pathologist" needs an integrated approach: While machine learning algorithms require many thousands of training examples, a human expert is often confronted with only a few data points. Interestingly, humans can learn from such few examples and are able to instantly interpret complex patterns. Consequently, the grand goal is to combine the possibilities of artificial intelligence with human intelligence and to find a well-suited balance between them to enable what neither of them could do on their own. This can raise the quality of education, diagnosis, prognosis and prediction of cancer and other diseases. In this paper we describe some (incomplete) research issues which we believe should be addressed in an integrated and concerted effort for paving the way towards the augmented pathologist.
Augmentor: An Image Augmentation Library for Machine Learning
Aug 11, 2017
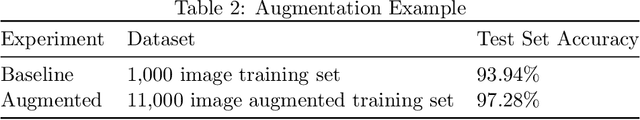

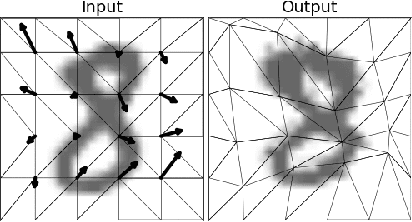

Abstract:The generation of artificial data based on existing observations, known as data augmentation, is a technique used in machine learning to improve model accuracy, generalisation, and to control overfitting. Augmentor is a software package, available in both Python and Julia versions, that provides a high level API for the expansion of image data using a stochastic, pipeline-based approach which effectively allows for images to be sampled from a distribution of augmented images at runtime. Augmentor provides methods for most standard augmentation practices as well as several advanced features such as label-preserving, randomised elastic distortions, and provides many helper functions for typical augmentation tasks used in machine learning.
A glass-box interactive machine learning approach for solving NP-hard problems with the human-in-the-loop
Aug 03, 2017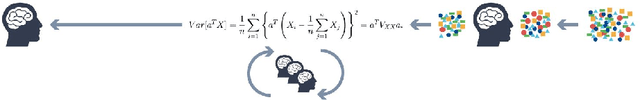

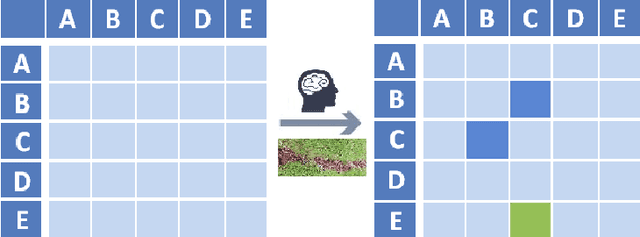
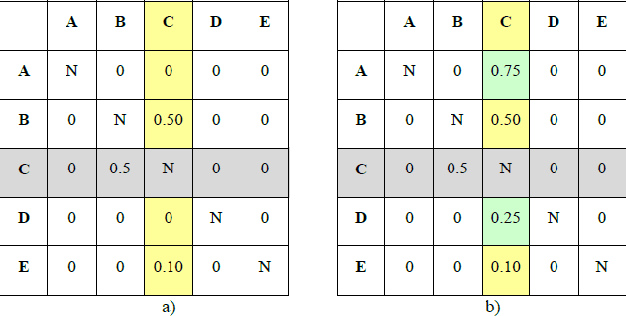
Abstract:The goal of Machine Learning to automatically learn from data, extract knowledge and to make decisions without any human intervention. Such automatic (aML) approaches show impressive success. Recent results even demonstrate intriguingly that deep learning applied for automatic classification of skin lesions is on par with the performance of dermatologists, yet outperforms the average. As human perception is inherently limited, such approaches can discover patterns, e.g. that two objects are similar, in arbitrarily high-dimensional spaces what no human is able to do. Humans can deal only with limited amounts of data, whilst big data is beneficial for aML; however, in health informatics, we are often confronted with a small number of data sets, where aML suffer of insufficient training samples and many problems are computationally hard. Here, interactive machine learning (iML) may be of help, where a human-in-the-loop contributes to reduce the complexity of NP-hard problems. A further motivation for iML is that standard black-box approaches lack transparency, hence do not foster trust and acceptance of ML among end-users. Rising legal and privacy aspects, e.g. with the new European General Data Protection Regulations, make black-box approaches difficult to use, because they often are not able to explain why a decision has been made. In this paper, we present some experiments to demonstrate the effectiveness of the human-in-the-loop approach, particularly in opening the black-box to a glass-box and thus enabling a human directly to interact with an learning algorithm. We selected the Ant Colony Optimization framework, and applied it on the Traveling Salesman Problem, which is a good example, due to its relevance for health informatics, e.g. for the study of protein folding. From studies of how humans extract so much from so little data, fundamental ML-research also may benefit.
Review of Machine Learning Algorithms in Differential Expression Analysis
Jul 28, 2017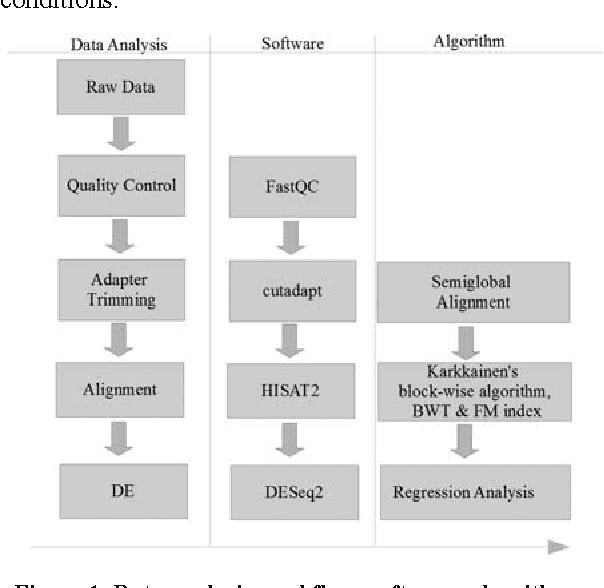
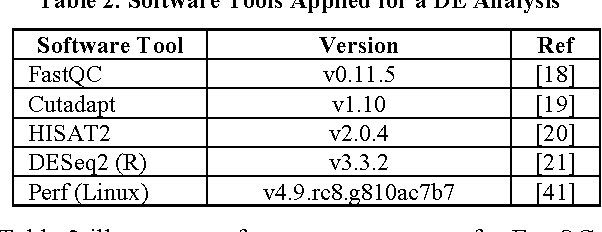
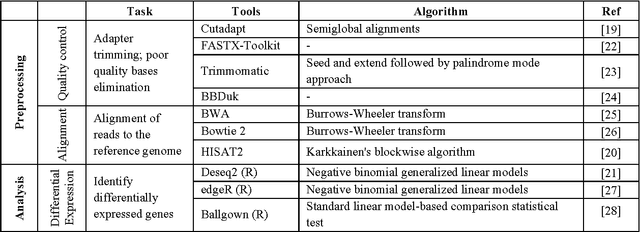
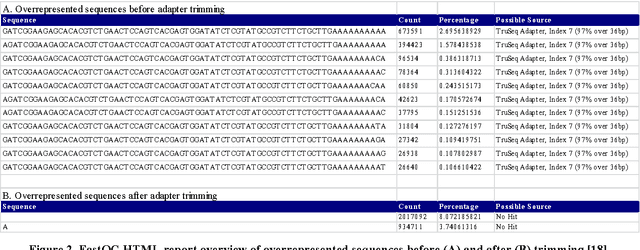

Abstract:In biological research machine learning algorithms are part of nearly every analytical process. They are used to identify new insights into biological phenomena, interpret data, provide molecular diagnosis for diseases and develop personalized medicine that will enable future treatments of diseases. In this paper we (1) illustrate the importance of machine learning in the analysis of large scale sequencing data, (2) present an illustrative standardized workflow of the analysis process, (3) perform a Differential Expression (DE) analysis of a publicly available RNA sequencing (RNASeq) data set to demonstrate the capabilities of various algorithms at each step of the workflow, and (4) show a machine learning solution in improving the computing time, storage requirements, and minimize utilization of computer memory in analyses of RNA-Seq datasets. The source code of the analysis pipeline and associated scripts are presented in the paper appendix to allow replication of experiments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge